

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0359.2
(453 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
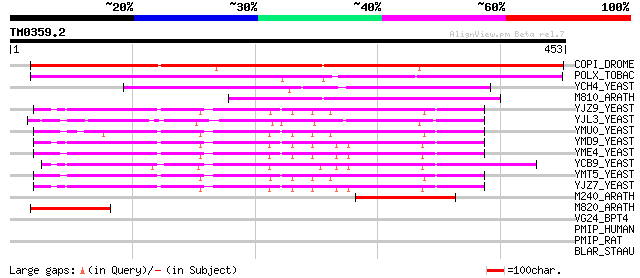
Score E
Sequences producing significant alignments: (bits) Value
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 332 1e-90
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 297 3e-80
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 161 4e-39
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 160 5e-39
YJZ9_YEAST (P47100) Transposon Ty1 protein B 97 7e-20
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 97 7e-20
YMU0_YEAST (Q04670) Transposon Ty1 protein B 96 2e-19
YMD9_YEAST (Q03434) Transposon Ty1 protein B 95 3e-19
YME4_YEAST (Q04711) Transposon Ty1 protein B 95 4e-19
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 95 4e-19
YMT5_YEAST (Q04214) Transposon Ty1 protein B 94 6e-19
YJZ7_YEAST (P47098) Transposon Ty1 protein B 93 1e-18
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 69 2e-11
M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820... 62 3e-09
VG24_BPT4 (P19896) Head vertex protein Gp24 34 0.88
PMIP_HUMAN (Q99797) Mitochondrial intermediate peptidase, mitoch... 31 5.7
PMIP_RAT (Q01992) Mitochondrial intermediate peptidase, mitochon... 30 9.8
BLAR_STAAU (P18357) Regulatory protein blaR1 30 9.8
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 332 bits (851), Expect = 1e-90
Identities = 177/442 (40%), Positives = 273/442 (61%), Gaps = 10/442 (2%)
Query: 18 LTKKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAI 77
+TK+PEN +++ ++WVF K NE G+ +R K RLVA+G++Q+ IDY ETFAPVAR+ +
Sbjct: 925 ITKRPENKNIVDSRWVFSVKYNELGNPIRYKARLVARGFTQKYQIDYEETFAPVARISSF 984
Query: 78 ILLISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLK 137
++S + +N+ +HQMDVK+AFLNG + +E+Y+ P G D+V KL K++YGLK
Sbjct: 985 RFILSLVIQYNLKVHQMDVKTAFLNGTLKEEIYMRLPQGISCNS--DNVCKLNKAIYGLK 1042
Query: 138 QAPRAWYERLSSFLLENEFVRGKVDTTLFC--KTYKDDILIVKIYVDDIIFGSANQSLCK 195
QA R W+E L E EFV VD ++ K ++ + V +YVDD++ + + +
Sbjct: 1043 QAARCWFEVFEQALKECEFVNSSVDRCIYILDKGNINENIYVLLYVDDVVIATGDMTRMN 1102
Query: 196 EFSKMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPM 255
F + + +F M+ + E+K+F+GI+++ + Y+ QS Y K++L KFNM TP+
Sbjct: 1103 NFKRYLMEKFRMTDLNEIKHFIGIRIEMQEDKIYLSQSAYVKKILSKFNMENCNAVSTPL 1162
Query: 256 HPTCILEKEDTSGKVCQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTAV 314
P+ I + S + C R +IG L+Y + +RPD+ +V++ +R+ S +
Sbjct: 1163 -PSKINYELLNSDEDCNTPCRSLIGCLMYIMLCTRPDLTTAVNILSRYSSKNNSELWQNL 1221
Query: 315 KRILRYLKGTINLGLMYKK--TSEYKLSGYCDADYAGDRTERKSTFGNC-QFLGSNLVSW 371
KR+LRYLKGTI++ L++KK E K+ GY D+D+AG +RKST G + NL+ W
Sbjct: 1222 KRVLRYLKGTIDMKLIFKKNLAFENKIIGYVDSDWAGSEIDRKSTTGYLFKMFDFNLICW 1281
Query: 372 ASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKN 430
+KRQ+++A S+ EAEY++ + LW+K L I LE+ I IY DN IS++ N
Sbjct: 1282 NTKRQNSVAASSTEAEYMALFEAVREALWLKFLLTSINIKLENPIKIYEDNQGCISIANN 1341
Query: 431 PILHSRAKHIEVKYHFIRGYVQ 452
P H RAKHI++KYHF R VQ
Sbjct: 1342 PSCHKRAKHIDIKYHFAREQVQ 1363
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 297 bits (761), Expect = 3e-80
Identities = 165/448 (36%), Positives = 263/448 (57%), Gaps = 19/448 (4%)
Query: 18 LTKKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAI 77
L + P+ + KWVF+ K + +VR K RLV +G+ Q++GID+ E F+PV ++ +I
Sbjct: 845 LVELPKGKRPLKCKWVFKLKKDGDCKLVRYKARLVVKGFEQKKGIDFDEIFSPVVKMTSI 904
Query: 78 ILLISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLK 137
++S + + ++ + Q+DVK+AFL+G + +E+Y+ QP GFE K V KL KSLYGLK
Sbjct: 905 RTILSLAASLDLEVEQLDVKTAFLHGDLEEEIYMEQPEGFEVAGKKHMVCKLNKSLYGLK 964
Query: 138 QAPRAWYERLSSFLLENEFVRGKVDTTLFCKTY-KDDILIVKIYVDDIIFGSANQSLCKE 196
QAPR WY + SF+ +++ D ++ K + +++ +I+ +YVDD++ ++ L +
Sbjct: 965 QAPRQWYMKFDSFMKSQTYLKTYSDPCVYFKRFSENNFIILLLYVDDMLIVGKDKGLIAK 1024
Query: 197 FSKMMQAEFEMSMMGELKYFLGIQV--DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTP 254
+ F+M +G + LG+++ ++T ++ Q KY + +L++FNM + TP
Sbjct: 1025 LKGDLSKSFDMKDLGPAQQILGMKIVRERTSRKLWLSQEKYIERVLERFNMKNAKPVSTP 1084
Query: 255 ----------MHPTCILEKEDTSGKVCQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQ 303
M PT + EK G + + Y +GSL+Y + +RPDI +V + +RF
Sbjct: 1085 LAGHLKLSKKMCPTTVEEK----GNMAKVPYSSAVGSLMYAMVCTRPDIAHAVGVVSRFL 1140
Query: 304 SDPRETHLTAVKRILRYLKGTINLGLMYKKTSEYKLSGYCDADYAGDRTERKSTFGNCQF 363
+P + H AVK ILRYL+GT L + S+ L GY DAD AGD RKS+ G
Sbjct: 1141 ENPGKEHWEAVKWILRYLRGTTGDCLCF-GGSDPILKGYTDADMAGDIDNRKSSTGYLFT 1199
Query: 364 LGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTA 423
+SW SK Q +ALST EAEYI+A +M+W+K L++ + + +YCD+ +
Sbjct: 1200 FSGGAISWQSKLQKCVALSTTEAEYIAATETGKEMIWLKRFLQELGLHQKEYVVYCDSQS 1259
Query: 424 AISLSKNPILHSRAKHIEVKYHFIRGYV 451
AI LSKN + H+R KHI+V+YH+IR V
Sbjct: 1260 AIDLSKNSMYHARTKHIDVRYHWIREMV 1287
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 161 bits (407), Expect = 4e-39
Identities = 96/309 (31%), Positives = 167/309 (53%), Gaps = 17/309 (5%)
Query: 94 MDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLE 153
MDV +AFLN + + +YV QPPGF +E+ PD+V++L +YGLKQAP W E +++ L +
Sbjct: 1 MDVDTAFLNSTMDEPIYVKQPPGFVNERNPDYVWELYGGMYGLKQAPLLWNEHINNTLKK 60
Query: 154 NEFVRGKVDTTLFCKTYKDDILIVKIYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGEL 213
F R + + L+ ++ D + + +YVDD++ + + + + + + M +G++
Sbjct: 61 IGFCRHEGEHGLYFRSTSDGPIYIGVYVDDLLVAAPSPKIYDRVKQELTKLYSMKDLGKV 120
Query: 214 KYFLGIQVDQTPEG-------TYIHQSKYTKELLKKFNMLESTVAKT-PMHPTCILEKED 265
FLG+ + Q+ G YI ++ E + F + ++ + + P+ T +D
Sbjct: 121 DKFLGLNIHQSTNGDITLSLQDYIAKAASESE-INTFKLTQTPLCNSKPLFETTSPHLKD 179
Query: 266 TSGKVCQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGT 324
+ Y+ ++G LL+ RPDI + V L +RF +PR HL + +R+LRYL T
Sbjct: 180 ITP------YQSIVGQLLFCANTGRPDISYPVSLLSRFLREPRAIHLESARRVLRYLYTT 233
Query: 325 INLGLMYKKTSEYKLSGYCDADYAGDRTERKSTFGNCQFLGSNLVSWASKR-QSTIALST 383
++ L Y+ S+ L+ YCDA + ST G L V+W+SK+ + I + +
Sbjct: 234 RSMCLKYRSGSQVALTVYCDASHGAIHDLPHSTGGYVTLLAGAPVTWSSKKLKGVIPVPS 293
Query: 384 AEAEYISAA 392
EAEYI+A+
Sbjct: 294 TEAEYITAS 302
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 160 bits (406), Expect = 5e-39
Identities = 78/222 (35%), Positives = 131/222 (58%), Gaps = 1/222 (0%)
Query: 179 IYVDDIIFGSANQSLCKEFSKMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKE 238
+YVDDI+ ++ +L + + F M +G + YFLGIQ+ P G ++ Q+KY ++
Sbjct: 5 LYVDDILLTGSSNTLLNMLIFQLSSTFSMKDLGPVHYFLGIQIKTHPSGLFLSQTKYAEQ 64
Query: 239 LLKKFNMLESTVAKTPMHPTCILEKEDTSGKVCQKLYRGMIGSLLYLTASRPDILFSVHL 298
+L ML+ TP+ P + T+ +R ++G+L YLT +RPDI ++V++
Sbjct: 65 ILNNAGMLDCKPMSTPL-PLKLNSSVSTAKYPDPSDFRSIVGALQYLTLTRPDISYAVNI 123
Query: 299 CARFQSDPRETHLTAVKRILRYLKGTINLGLMYKKTSEYKLSGYCDADYAGDRTERKSTF 358
+ +P +KR+LRY+KGTI GL K S+ + +CD+D+AG + R+ST
Sbjct: 124 VCQRMHEPTLADFDLLKRVLRYVKGTIFHGLYIHKNSKLNVQAFCDSDWAGCTSTRRSTT 183
Query: 359 GNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLW 400
G C FLG N++SW++KRQ T++ S+ E EY + A+ + ++ W
Sbjct: 184 GFCTFLGCNIISWSAKRQPTVSRSSTETEYRALALTAAELTW 225
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 97.4 bits (241), Expect = 7e-20
Identities = 99/393 (25%), Positives = 182/393 (46%), Gaps = 41/393 (10%)
Query: 20 KKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAIIL 79
K+ + VI + ++F N+K D +K R VA+G Q + + A++
Sbjct: 1276 KEIDPKRVINSMFIF----NKKRDGT-HKARFVARGDIQHPDTYDSGMQSNTVHHYALMT 1330
Query: 80 LISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQA 139
+S +L++N + Q+D+ SA+L I +E+Y+ PP D + +LKKSLYGLKQ+
Sbjct: 1331 SLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGMN---DKLIRLKKSLYGLKQS 1387
Query: 140 PRAWYERLSSFLLEN---EFVRGKVDTTLFCKTYKDDILIVKIYVDDIIFGSANQSLCKE 196
WYE + S+L++ E VRG + +K+ + + ++VDD++ S N + K
Sbjct: 1388 GANWYETIKSYLIQQCGMEEVRG------WSCVFKNSQVTICLFVDDMVLFSKNLNSNKR 1441
Query: 197 FSKMMQAEFEMSMMG------ELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM-- 245
+ ++ +++ ++ E++Y LG+++ + G Y + E + K N+
Sbjct: 1442 IIEKLKMQYDTKIINLGESDEEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPL 1500
Query: 246 -LESTVAKTPMHPTCI-----LEKEDTSGKVCQKLYRGMIGSLLYLTAS-RPDILFSVHL 298
+ P P LE E+ K+ + +IG Y+ R D+L+ ++
Sbjct: 1501 NPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINT 1560
Query: 299 CARFQSDPRETHLTAVKRILRYLKGTINLGLMYKKTSEY----KLSGYCDADYAGDRTER 354
A+ P + L +++++ T + L++ K+ KL DA Y G++
Sbjct: 1561 LAQHILFPSKQVLDMTYELIQFIWNTRDKQLIWHKSKPVKPTNKLVVISDASY-GNQPYY 1619
Query: 355 KSTFGNCQFLGSNLVSWASKRQSTIALSTAEAE 387
KS GN L ++ S + S ST EAE
Sbjct: 1620 KSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE 1652
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 97.4 bits (241), Expect = 7e-20
Identities = 98/400 (24%), Positives = 173/400 (42%), Gaps = 50/400 (12%)
Query: 15 KLDLTKKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARL 74
K ++ P+N+ ++ T +F K N K R+V +G +Q Y+
Sbjct: 1311 KYSRSEIPDNL-IVPTNTIFTKKRNGI-----YKARIVCRGDTQSPDT-YSVITTESLNH 1363
Query: 75 EAIILLISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLY 134
I + + + N N+ + +D+ AFL + +E+Y+ P D + V KL K+LY
Sbjct: 1364 NHIKIFLMIANNRNMFMKTLDINHAFLYAKLEEEIYIPHP---HDRRC---VVKLNKALY 1417
Query: 135 GLKQAPRAWYERLSSFL-----LENEFVRGKVDTTLFCKTYKDDILIVKIYVDDIIFGSA 189
GLKQ+P+ W + L +L +N + G T +D L++ +YVDD + ++
Sbjct: 1418 GLKQSPKEWNDHLRQYLNGIGLKDNSYTPGLYQT-------EDKNLMIAVYVDDCVIAAS 1470
Query: 190 NQSLCKEFSKMMQAEFEMSMMGEL------KYFLGIQ---------VDQTPEGTYIHQSK 234
N+ EF +++ FE+ + G L LG+ +D T + K
Sbjct: 1471 NEQRLDEFINKLKSNFELKITGTLIDDVLDTDILGMDLVYNKRLGTIDLTLKSFINRMDK 1530
Query: 235 YTKELLKKFNMLE----STVAKTPMHPTCILEKEDTSGKVCQKLYRGMIGSLLYLT-ASR 289
E LKK ST P + +E+ V + + ++G L Y+ R
Sbjct: 1531 KYNEELKKIRKSSIPHMSTYKIDPKKDVLQMSEEEFRQGVLK--LQQLLGELNYVRHKCR 1588
Query: 290 PDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTINLGLMYKK--TSEYKLSGYCDADY 347
DI F+V AR + P E + +I++YL ++G+ Y + + K+ DA
Sbjct: 1589 YDIEFAVKKVARLVNYPHERVFYMIYKIIQYLVRYKDIGIHYDRDCNKDKKVIAITDAS- 1647
Query: 348 AGDRTERKSTFGNCQFLGSNLVSWASKRQSTIALSTAEAE 387
G + +S G + G N+ + S + + +S+ EAE
Sbjct: 1648 VGSEYDAQSRIGVILWYGMNIFNVYSNKSTNRCVSSTEAE 1687
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 95.9 bits (237), Expect = 2e-19
Identities = 99/398 (24%), Positives = 184/398 (45%), Gaps = 51/398 (12%)
Query: 20 KKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLE---- 75
K+ + VI + ++F K + +K R VA+G I + +T+ P +
Sbjct: 849 KEIDPKRVINSMFIFNRKRDGT-----HKARFVARG-----DIQHPDTYDPGMQSNTVHH 898
Query: 76 -AIILLISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLY 134
A++ +S +L++N + Q+D+ SA+L I +E+Y+ PP D + +LKKSLY
Sbjct: 899 YALMTSLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGMN---DKLIRLKKSLY 955
Query: 135 GLKQAPRAWYERLSSFLLEN---EFVRGKVDTTLFCKTYKDDILIVKIYVDDIIFGSANQ 191
GLKQ+ WYE + S+L++ E VRG + +K+ + + ++VDD+I S +
Sbjct: 956 GLKQSGANWYETIKSYLIKQCGMEEVRG------WSCVFKNSQVTICLFVDDMILFSKDL 1009
Query: 192 SLCKEFSKMMQAEFEMSMM------GELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKK 242
+ K+ ++ +++ ++ E++Y LG+++ + G Y + E + K
Sbjct: 1010 NANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPK 1068
Query: 243 FNM---LESTVAKTPMHPTCI-----LEKEDTSGKVCQKLYRGMIGSLLYLTAS-RPDIL 293
N+ + P P LE E+ K+ + +IG Y+ R D+L
Sbjct: 1069 LNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKLIGLASYVGYKFRFDLL 1128
Query: 294 FSVHLCARFQSDPRETHLTAVKRILRYLKGTINLGLMYKKTSEY----KLSGYCDADYAG 349
+ ++ A+ P + L +++++ T + L++ K+ KL DA Y G
Sbjct: 1129 YYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQLIWHKSKPVKPTNKLVVISDASY-G 1187
Query: 350 DRTERKSTFGNCQFLGSNLVSWASKRQSTIALSTAEAE 387
++ KS GN L ++ S + S ST EAE
Sbjct: 1188 NQPYYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE 1225
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 95.1 bits (235), Expect = 3e-19
Identities = 96/393 (24%), Positives = 185/393 (46%), Gaps = 41/393 (10%)
Query: 20 KKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAIIL 79
K+ + VI + ++F N+K D +K R VA+G Q + + A++
Sbjct: 849 KEIDPKRVINSMFIF----NKKRDGT-HKARFVARGDIQHPDTYDSGMQSNTVHHYALMT 903
Query: 80 LISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQA 139
+S +L++N + Q+D+ SA+L I +E+Y+ PP D + +LKKSLYGLKQ+
Sbjct: 904 SLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGMN---DKLIRLKKSLYGLKQS 960
Query: 140 PRAWYERLSSFLLEN---EFVRGKVDTTLFCKTYKDDILIVKIYVDDIIFGSANQSLCKE 196
WYE + S+L++ E VRG + +K+ + + ++VDD+I S + + K+
Sbjct: 961 GANWYETIKSYLIKQCGMEEVRG------WSCVFKNSQVTICLFVDDMILFSKDLNANKK 1014
Query: 197 FSKMMQAEFEMSMM------GELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM-- 245
++ +++ ++ E++Y LG+++ + G Y + E + K N+
Sbjct: 1015 IITTLKKQYDTKIINLGESDNEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPL 1073
Query: 246 -LESTVAKTPMHPTCILEKED---TSGKVCQKLY--RGMIGSLLYLTAS-RPDILFSVHL 298
+ P P +++++ + +K++ + +IG Y+ R D+L+ ++
Sbjct: 1074 NPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINT 1133
Query: 299 CARFQSDPRETHLTAVKRILRYLKGTINLGLMYKKTS----EYKLSGYCDADYAGDRTER 354
A+ P L +++++ T + L++ K + KL DA Y G++
Sbjct: 1134 LAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPTEPDNKLVAISDASY-GNQPYY 1192
Query: 355 KSTFGNCQFLGSNLVSWASKRQSTIALSTAEAE 387
KS GN L ++ S + S ST EAE
Sbjct: 1193 KSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE 1225
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 94.7 bits (234), Expect = 4e-19
Identities = 94/393 (23%), Positives = 183/393 (45%), Gaps = 41/393 (10%)
Query: 20 KKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAIIL 79
K+ + VI + ++F K + +K R VA+G Q + + A++
Sbjct: 849 KEIDPKRVINSMFIFNRKRDGT-----HKARFVARGDIQHPDTYDSGMQSNTVHHYALMT 903
Query: 80 LISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQA 139
+S +L++N + Q+D+ SA+L I +E+Y+ PP D + +LKKSLYGLKQ+
Sbjct: 904 SLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGMN---DKLIRLKKSLYGLKQS 960
Query: 140 PRAWYERLSSFLLEN---EFVRGKVDTTLFCKTYKDDILIVKIYVDDIIFGSANQSLCKE 196
WYE + S+L++ E VRG + +K+ + + ++VDD+I S + + K+
Sbjct: 961 GANWYETIKSYLIKQCGMEEVRG------WSCVFKNSQVTICLFVDDMILFSKDLNANKK 1014
Query: 197 FSKMMQAEFEMSMM------GELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM-- 245
++ +++ ++ E++Y LG+++ + G Y + E + K N+
Sbjct: 1015 IITTLKKQYDTKIINLGESDNEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPL 1073
Query: 246 -LESTVAKTPMHPTCILEKED---TSGKVCQKLY--RGMIGSLLYLTAS-RPDILFSVHL 298
+ P P +++++ + +K++ + +IG Y+ R D+L+ ++
Sbjct: 1074 NPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINT 1133
Query: 299 CARFQSDPRETHLTAVKRILRYLKGTINLGLMYKKTS----EYKLSGYCDADYAGDRTER 354
A+ P L +++++ T + L++ K + KL DA Y G++
Sbjct: 1134 LAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPTEPDNKLVAISDASY-GNQPYY 1192
Query: 355 KSTFGNCQFLGSNLVSWASKRQSTIALSTAEAE 387
KS GN L ++ S + S ST EAE
Sbjct: 1193 KSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE 1225
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 94.7 bits (234), Expect = 4e-19
Identities = 100/430 (23%), Positives = 203/430 (46%), Gaps = 43/430 (10%)
Query: 27 VIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAIILLISFSLN 86
VI + ++F N+K D +K R VA+G Q ++ + A++ +S +L+
Sbjct: 1298 VINSMFIF----NKKRDGT-HKARFVARGDIQHPDTYDSDMQSNTVHHYALMTSLSIALD 1352
Query: 87 HNIILHQMDVKSAFLNGYISKEVYVHQPP--GFEDEKKPDHVFKLKKSLYGLKQAPRAWY 144
++ + Q+D+ SA+L I +E+Y+ PP G D+ + +L+KSLYGLKQ+ WY
Sbjct: 1353 NDYYITQLDISSAYLYADIKEELYIRPPPHLGLNDK-----LLRLRKSLYGLKQSGANWY 1407
Query: 145 ERLSSFLL---ENEFVRGKVDTTLFCKTYKDDILIVKIYVDDIIFGSANQSLCKEFSKMM 201
E + S+L+ + + VRG + +K+ + + ++VDD+I S + + K+ +
Sbjct: 1408 ETIKSYLINCCDMQEVRG------WSCVFKNSQVTICLFVDDMILFSKDLNANKKIITTL 1461
Query: 202 QAEFEMSMM------GELKY-FLGIQVD-QTPEGTYIHQSKYTKELLKKFNMLESTVAK- 252
+ +++ ++ E++Y LG+++ Q + + K E L K N+ + K
Sbjct: 1462 KKQYDTKIINLGESDNEIQYDILGLEIKYQRSKYMKLGMEKSLTEKLPKLNVPLNPKGKK 1521
Query: 253 --TPMHPTCILEKED---TSGKVCQKLY--RGMIGSLLYLTAS-RPDILFSVHLCARFQS 304
P P +++++ + +K++ + +IG Y+ R D+L+ ++ A+
Sbjct: 1522 LRAPGQPGHYIDQDELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHIL 1581
Query: 305 DPRETHLTAVKRILRYLKGTINLGLMYKKTS----EYKLSGYCDADYAGDRTERKSTFGN 360
P L +++++ T + L++ K + KL DA Y G++ KS GN
Sbjct: 1582 FPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPTKPDNKLVAISDASY-GNQPYYKSQIGN 1640
Query: 361 CQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCD 420
L ++ S + S ST EAE + + + + H +++ + D
Sbjct: 1641 IFLLNGKVIGGKSTKASLTCTSTTEAEIHAVSEAIPLLNNLSHLVQELNKKPIIKGLLTD 1700
Query: 421 NTAAISLSKN 430
+ + IS+ K+
Sbjct: 1701 SRSTISIIKS 1710
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 94.4 bits (233), Expect = 6e-19
Identities = 96/393 (24%), Positives = 179/393 (45%), Gaps = 41/393 (10%)
Query: 20 KKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAIIL 79
K+ + VI + ++F K + +K R VA+G Q + + A++
Sbjct: 849 KEIDPKRVINSMFIFNRKRDGT-----HKARFVARGDIQHPDTYDSGMQSNTVHHYALMT 903
Query: 80 LISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQA 139
+S +L++N + Q+D+ SA+L I +E+Y+ PP D + +LKKSLYGLKQ+
Sbjct: 904 SLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGMN---DKLIRLKKSLYGLKQS 960
Query: 140 PRAWYERLSSFLLEN---EFVRGKVDTTLFCKTYKDDILIVKIYVDDIIFGSANQSLCKE 196
WYE + S+L++ E VRG + +++ + + ++VDD++ S N + K
Sbjct: 961 GANWYETIKSYLIKQCGMEEVRG------WSCVFENSQVTICLFVDDMVLFSKNLNSNKR 1014
Query: 197 FSKMMQAEFEMSMMG------ELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM-- 245
++ +++ ++ E++Y LG+++ + G Y + E + K N+
Sbjct: 1015 IIDKLKMQYDTKIINLGESDEEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPL 1073
Query: 246 -LESTVAKTPMHPTCI-----LEKEDTSGKVCQKLYRGMIGSLLYLTAS-RPDILFSVHL 298
+ P P LE E+ K+ + +IG Y+ R D+L+ ++
Sbjct: 1074 NPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINT 1133
Query: 299 CARFQSDPRETHLTAVKRILRYLKGTINLGLMYKKTSEY----KLSGYCDADYAGDRTER 354
A+ P + L +++++ T + L++ K+ KL DA Y G++
Sbjct: 1134 LAQHILFPSKQVLDMTYELIQFIWNTRDKQLIWHKSKPVKPTNKLVVISDASY-GNQPYY 1192
Query: 355 KSTFGNCQFLGSNLVSWASKRQSTIALSTAEAE 387
KS GN L ++ S + S ST EAE
Sbjct: 1193 KSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAE 1225
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 93.2 bits (230), Expect = 1e-18
Identities = 96/393 (24%), Positives = 184/393 (46%), Gaps = 41/393 (10%)
Query: 20 KKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAIIL 79
K+ + VI + ++F N+K D +K R VA+G Q T + A++
Sbjct: 1276 KEIDPKRVINSMFIF----NKKRDGT-HKARFVARGDIQHPDTYDTGMQSNTVHHYALMT 1330
Query: 80 LISFSLNHNIILHQMDVKSAFLNGYISKEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQA 139
+S +L++N + Q+D+ SA+L I +E+Y+ PP D + +LKKS YGLKQ+
Sbjct: 1331 SLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGMN---DKLIRLKKSHYGLKQS 1387
Query: 140 PRAWYERLSSFLLEN---EFVRGKVDTTLFCKTYKDDILIVKIYVDDIIFGSANQSLCKE 196
WYE + S+L++ E VRG + +K+ + + ++VDD+I S + + K+
Sbjct: 1388 GANWYETIKSYLIKQCGMEEVRG------WSCVFKNSQVTICLFVDDMILFSKDLNANKK 1441
Query: 197 FSKMMQAEFEMSMM------GELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM-- 245
++ +++ ++ E++Y LG+++ + G Y + E + K N+
Sbjct: 1442 IITTLKKQYDTKIINLGESDNEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPL 1500
Query: 246 -LESTVAKTPMHPTCILEKED---TSGKVCQKLY--RGMIGSLLYLTAS-RPDILFSVHL 298
+ P P +++++ + +K++ + +IG Y+ R D+L+ ++
Sbjct: 1501 NPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINT 1560
Query: 299 CARFQSDPRETHLTAVKRILRYLKGTINLGLMYKKTS----EYKLSGYCDADYAGDRTER 354
A+ P L +++++ T + L++ K + KL DA Y G++
Sbjct: 1561 LAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPTEPDNKLVAISDASY-GNQPYY 1619
Query: 355 KSTFGNCQFLGSNLVSWASKRQSTIALSTAEAE 387
KS GN L ++ S + S ST EAE
Sbjct: 1620 KSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAE 1652
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 69.3 bits (168), Expect = 2e-11
Identities = 31/82 (37%), Positives = 50/82 (60%)
Query: 283 LYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTINLGLMYKKTSEYKLSGY 342
+YLT +RPD+ F+V+ ++F S R + AV ++L Y+KGT+ GL Y TS+ +L +
Sbjct: 1 MYLTITRPDLTFAVNRLSQFSSASRTAQMQAVYKVLHYVKGTVGQGLFYSATSDLQLKAF 60
Query: 343 CDADYAGDRTERKSTFGNCQFL 364
D+D+A R+S G C +
Sbjct: 61 ADSDWASCPDTRRSVTGFCSLV 82
>M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820
(ORF170)
Length = 170
Score = 62.0 bits (149), Expect = 3e-09
Identities = 28/65 (43%), Positives = 43/65 (66%)
Query: 18 LTKKPENVHVIGTKWVFRNKLNEKGDVVRNKERLVAQGYSQQEGIDYTETFAPVARLEAI 77
L P N +++G KWVF+ KL+ G + R K RLVA+G+ Q+EGI + ET++PV R I
Sbjct: 59 LVPPPVNQNILGCKWVFKTKLHSDGTLDRLKARLVAKGFHQEEGIYFVETYSPVVRTATI 118
Query: 78 ILLIS 82
+++
Sbjct: 119 RTILN 123
>VG24_BPT4 (P19896) Head vertex protein Gp24
Length = 427
Score = 33.9 bits (76), Expect = 0.88
Identities = 29/113 (25%), Positives = 50/113 (43%), Gaps = 15/113 (13%)
Query: 321 LKGTINLGLMYKKTSEYKLSGYCDADYAGDRTERKSTFGNCQFLGSNLVSWASKRQSTIA 380
L+ I + YK T SG+ D YA +S + + S++ +++ST
Sbjct: 215 LQSLITVSKRYKVTGITD-SGFIDLSYASAPEAGRSLYRMVCEMVSHI-----QKESTYT 268
Query: 381 LSTAEAEYISAAICSTQMLWMKHQLED--------YQILESNIPIYCDNTAAI 425
+ A +AAI + W+KH+ ED Y L + +P+YCD + +
Sbjct: 269 ATFCVASARAAAILAASG-WLKHKPEDDKYLSQNAYGFLANGLPLYCDTNSPL 320
>PMIP_HUMAN (Q99797) Mitochondrial intermediate peptidase,
mitochondrial precursor (EC 3.4.24.59) (MIP)
Length = 713
Score = 31.2 bits (69), Expect = 5.7
Identities = 18/66 (27%), Positives = 29/66 (43%), Gaps = 4/66 (6%)
Query: 207 MSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDT 266
+ + GE G+ PEG +I Q K L+K +L TP P +L ++
Sbjct: 56 LDLFGERARLFGVPELSAPEGFHIAQEK----ALRKTELLVDRACSTPPGPQTVLIFDEL 111
Query: 267 SGKVCQ 272
S +C+
Sbjct: 112 SDSLCR 117
>PMIP_RAT (Q01992) Mitochondrial intermediate peptidase,
mitochondrial precursor (EC 3.4.24.59) (MIP)
Length = 710
Score = 30.4 bits (67), Expect = 9.8
Identities = 18/64 (28%), Positives = 30/64 (46%), Gaps = 4/64 (6%)
Query: 209 MMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDTSG 268
++GE + G+ TPEG + Q +E L+K L TP P +L ++ S
Sbjct: 56 LLGERRGLFGVPELSTPEGFQVAQ----EEALRKTEWLVERACSTPPGPQTVLIFDELSD 111
Query: 269 KVCQ 272
+C+
Sbjct: 112 CLCR 115
>BLAR_STAAU (P18357) Regulatory protein blaR1
Length = 585
Score = 30.4 bits (67), Expect = 9.8
Identities = 21/52 (40%), Positives = 30/52 (57%), Gaps = 7/52 (13%)
Query: 127 FKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVK 178
FK K+L LK + ++ S +L ENE + K+DT LF YK +I+I K
Sbjct: 119 FKFLKALLYLK-----YLKKQSLYLNENE--KNKIDTILFNHQYKKNIVIRK 163
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.137 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,904,989
Number of Sequences: 164201
Number of extensions: 2122490
Number of successful extensions: 4439
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 4393
Number of HSP's gapped (non-prelim): 18
length of query: 453
length of database: 59,974,054
effective HSP length: 114
effective length of query: 339
effective length of database: 41,255,140
effective search space: 13985492460
effective search space used: 13985492460
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0359.2


