

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0349b.2
(400 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
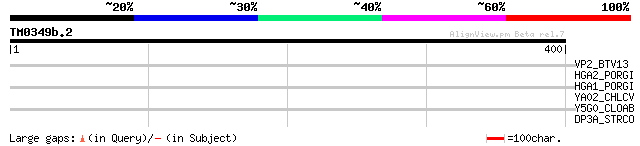
VP2_BTV13 (P12395) Outer capsid protein VP2 37 0.068
HGA2_PORGI (Q51845) Hemagglutinin A precursor 31 6.4
HGA1_PORGI (P59915) Hemagglutinin A precursor 31 6.4
YA02_CHLCV (P59707) Hypothetical UPF0242 protein CCA01002 30 8.4
Y5G0_CLOAB (P33747) Hypothetical protein CAP0160 precursor 30 8.4
DP3A_STRCO (Q9Z618) DNA polymerase III alpha subunit (EC 2.7.7.7) 30 8.4
>VP2_BTV13 (P12395) Outer capsid protein VP2
Length = 959
Score = 37.4 bits (85), Expect = 0.068
Identities = 21/85 (24%), Positives = 41/85 (47%), Gaps = 2/85 (2%)
Query: 5 KMDPPEFDLEAYLEKSEREDTYILNRFRDRRNLILEDSAPRGRKHLSRDHAGANQRLIDD 64
K+DP +F++E L + ++ +RF ++R + R HL R +GA + +
Sbjct: 654 KIDPSKFEVEGTLHDVSQVMVHLFDRFFEKRRFLRTVDESRWILHLIRSASGARRLEVLS 713
Query: 65 YFLNEPTYDDGIFRRRYRMQKHVFL 89
F P + DG+ R ++ + + L
Sbjct: 714 RFF--PAFSDGLRIREFKKVRDIML 736
>HGA2_PORGI (Q51845) Hemagglutinin A precursor
Length = 2628
Score = 30.8 bits (68), Expect = 6.4
Identities = 27/113 (23%), Positives = 45/113 (38%), Gaps = 7/113 (6%)
Query: 44 PRGRKHLSRDHAGANQRL---IDDYFL---NEPTYDDGIFRRRYRMQKHVFLRIVADLSS 97
P G K+++ H G +DD + N P+Y I+R ++ V D
Sbjct: 2469 PAGTKYVAFRHFGCTDFFWINLDDVVITSGNAPSYTYTIYRNNTQIASGVTETTYRDPDL 2528
Query: 98 SDNYFTQRVDAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLEC 150
+ ++T V G S + T + LA+ V A +G+T T+ C
Sbjct: 2529 ATGFYTYGVKVVYPNGESAIETATLNITSLAD-VTAQKPYTLTVVGKTITVTC 2580
>HGA1_PORGI (P59915) Hemagglutinin A precursor
Length = 2164
Score = 30.8 bits (68), Expect = 6.4
Identities = 27/113 (23%), Positives = 45/113 (38%), Gaps = 7/113 (6%)
Query: 44 PRGRKHLSRDHAGANQRL---IDDYFL---NEPTYDDGIFRRRYRMQKHVFLRIVADLSS 97
P G K+++ H G +DD + N P+Y I+R ++ V D
Sbjct: 2005 PAGTKYVAFRHFGCTDFFWINLDDVVITSGNAPSYTYTIYRNNTQIASGVTETTYRDPDL 2064
Query: 98 SDNYFTQRVDAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLEC 150
+ ++T V G S + T + LA+ V A +G+T T+ C
Sbjct: 2065 ATGFYTYGVKVVYPNGESAIETATLNITSLAD-VTAQKPYTLTVVGKTITVTC 2116
>YA02_CHLCV (P59707) Hypothetical UPF0242 protein CCA01002
Length = 401
Score = 30.4 bits (67), Expect = 8.4
Identities = 16/46 (34%), Positives = 27/46 (57%), Gaps = 1/46 (2%)
Query: 149 ECLRRFCKGII-RLYEQEYLRAPTQEDLQRILHVNEMWGVPKHDRE 193
E LR+ C+ + R YE + LR+ Q+ L ++LHV ++ K D +
Sbjct: 85 EGLRKICESVEERQYESQQLRSQNQKLLNQLLHVRSVFMKTKADTQ 130
>Y5G0_CLOAB (P33747) Hypothetical protein CAP0160 precursor
Length = 967
Score = 30.4 bits (67), Expect = 8.4
Identities = 14/51 (27%), Positives = 28/51 (54%)
Query: 103 TQRVDAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRR 153
T + AA ++G + + CT ++ LAN + + + + + +G+T L L R
Sbjct: 96 TVTITAATQDGSNLSSSCTVSITRLANSIVLNKMVDKLPVGQTDLLTALVR 146
>DP3A_STRCO (Q9Z618) DNA polymerase III alpha subunit (EC 2.7.7.7)
Length = 1179
Score = 30.4 bits (67), Expect = 8.4
Identities = 16/54 (29%), Positives = 27/54 (49%)
Query: 4 QKMDPPEFDLEAYLEKSEREDTYILNRFRDRRNLILEDSAPRGRKHLSRDHAGA 57
++ D P F L+ + E + TY+L + R+ L + D G +H+ D A A
Sbjct: 951 EQSDEPGFGLDVVFGEDEWDKTYLLAQEREMLGLYVSDHPLFGLEHVLSDKADA 1004
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.345 0.152 0.506
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,156,862
Number of Sequences: 164201
Number of extensions: 1574362
Number of successful extensions: 5945
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 5942
Number of HSP's gapped (non-prelim): 6
length of query: 400
length of database: 59,974,054
effective HSP length: 112
effective length of query: 288
effective length of database: 41,583,542
effective search space: 11976060096
effective search space used: 11976060096
T: 11
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0349b.2


