

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0344.6
(242 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
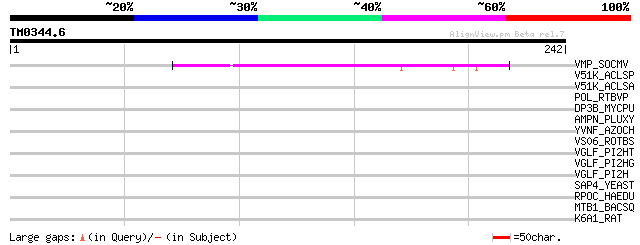
Score E
Sequences producing significant alignments: (bits) Value
VMP_SOCMV (P15631) Movement protein (Cell-TO-cell transport prot... 54 3e-07
V51K_ACLSP (P27739) 50.8 kDa protein (ORF2) 33 0.80
V51K_ACLSA (P54892) 50.4 kDa protein (ORF2) 32 1.1
POL_RTBVP (P27502) Polyprotein (P194 protein) [Contains: Coat pr... 32 1.4
DP3B_MYCPU (Q98RK6) DNA polymerase III, beta chain (EC 2.7.7.7) 32 1.4
AMPN_PLUXY (P91887) Aminopeptidase N precursor (EC 3.4.11.2) (Mi... 32 1.4
YVNF_AZOCH (P24423) Hypothetical protein in vnfD 5'region 31 3.1
VS06_ROTBS (Q00734) VP6 protein (Major capsid protein) (Major in... 30 4.0
VGLF_PI2HT (P26629) Fusion glycoprotein precursor [Contains: Fus... 30 4.0
VGLF_PI2HG (P27286) Fusion glycoprotein precursor [Contains: Fus... 30 4.0
VGLF_PI2H (P25467) Fusion glycoprotein precursor [Contains: Fusi... 30 4.0
SAP4_YEAST (P53036) SIT4-associating protein SAP4 30 4.0
RPOC_HAEDU (Q7VKL8) DNA-directed RNA polymerase beta' chain (EC ... 29 8.9
MTB1_BACSQ (Q9LAI2) Modification methylase BslI (EC 2.1.1.113) (... 29 8.9
K6A1_RAT (Q63531) Ribosomal protein S6 kinase alpha 1 (EC 2.7.1.... 29 8.9
>VMP_SOCMV (P15631) Movement protein (Cell-TO-cell transport
protein) (ORF IA)
Length = 303
Score = 53.9 bits (128), Expect = 3e-07
Identities = 43/152 (28%), Positives = 75/152 (49%), Gaps = 6/152 (3%)
Query: 72 SRIANQHVRKYGYLHIGLVQVAVKPLTRLRYCDLPILICLRDKRMLRFHESVVAVLESNI 131
S+I + R Y+HI +QV +K T L+ D P+ + LRD R+L ES +AV N+
Sbjct: 89 SKIEEKQRRLIRYVHISTLQVLIKS-TFLKGLDTPLELTLRDNRLLNLEESKIAVGHGNL 147
Query: 132 QNGPAYFNCFSSYPVCLRDPHVTTCVDLDIKIPNEMFI-DGIPPVSIMYRFCYKVTDNLF 190
+ G F+ + L+D + + L+ K F+ +G SI YR Y ++++
Sbjct: 148 KYGKMKFDVNLQLGLSLKDLDLDRSIILNYKFLRRNFMKEGNHAFSISYRINYALSNSHH 207
Query: 191 GI--KAVRDYTVNE--TEVTEINAENSSRITR 218
+ K ++E +EV E+ S++T+
Sbjct: 208 SVEFKQKEKIYIDELFSEVLELKHPVFSKLTK 239
>V51K_ACLSP (P27739) 50.8 kDa protein (ORF2)
Length = 460
Score = 32.7 bits (73), Expect = 0.80
Identities = 25/115 (21%), Positives = 51/115 (43%), Gaps = 4/115 (3%)
Query: 66 SISILDSRIANQHVRKY---GYLHIGLVQVAVKPLTRLRYCDLPILICLRDKRMLRFHES 122
SI ++ S +RK Y+H G + +++ L R + + + D R F ++
Sbjct: 65 SIPVIPSSEVQAVLRKRESTNYVHWGALSISIDALFR-KNAGVSGWCYVYDNRWETFEQA 123
Query: 123 VVAVLESNIQNGPAYFNCFSSYPVCLRDPHVTTCVDLDIKIPNEMFIDGIPPVSI 177
++ N+ +G A ++PV L DP ++ + + + N F P+S+
Sbjct: 124 MLQKFHFNLDSGSATLVTSPNFPVSLDDPGLSNSISVAVMFENLNFKFESYPISV 178
>V51K_ACLSA (P54892) 50.4 kDa protein (ORF2)
Length = 457
Score = 32.3 bits (72), Expect = 1.1
Identities = 25/115 (21%), Positives = 51/115 (43%), Gaps = 4/115 (3%)
Query: 66 SISILDSRIANQHVRKY---GYLHIGLVQVAVKPLTRLRYCDLPILICLRDKRMLRFHES 122
SI ++ S +RK Y+H G + +++ L R + + + D R F ++
Sbjct: 63 SIPVIPSSEVQAVLRKRESTNYVHWGALSISIDALFR-KNAGVSGWCYVYDNRWETFEQA 121
Query: 123 VVAVLESNIQNGPAYFNCFSSYPVCLRDPHVTTCVDLDIKIPNEMFIDGIPPVSI 177
++ N+ +G A ++PV L DP ++ + + + N F P+S+
Sbjct: 122 MLQKFRFNLDSGSATLVTSPNFPVSLDDPGLSNSISVAVMFENLNFKLESYPISV 176
>POL_RTBVP (P27502) Polyprotein (P194 protein) [Contains: Coat
protein; Protease (EC 3.4.23.-); Reverse transcriptase
(EC 2.7.7.49); Ribonuclease H (EC 3.1.26.4)]
Length = 1675
Score = 32.0 bits (71), Expect = 1.4
Identities = 33/148 (22%), Positives = 64/148 (42%), Gaps = 17/148 (11%)
Query: 14 EENDSNLEQKLQNWS-IPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPYQSISILDS 72
EE D N E + +W + I+ N++Y E N ++K ++S ++IS+ +
Sbjct: 36 EEVDVNQEVENFDWKKLSGIKPNKLY-----EKNWQEKVKLKQQSIVSAYKEEAISVTHN 90
Query: 73 RIAN--------QHVRKYG--YLHIGLVQVAVKPLTRLRYCDLPILICLRDKRMLRFHES 122
++V+ G Y HIG++ + VK L R R ++I D + E+
Sbjct: 91 AYTTTLFPQEVIKNVKNQGKLYYHIGMMAIGVKGLHR-RKIGTKVMIMFYDDSFGKAREA 149
Query: 123 VVAVLESNIQNGPAYFNCFSSYPVCLRD 150
+ +E ++ G F + ++D
Sbjct: 150 SIGSIEMDMNAGCGVFYSCPDFAKYIKD 177
>DP3B_MYCPU (Q98RK6) DNA polymerase III, beta chain (EC 2.7.7.7)
Length = 373
Score = 32.0 bits (71), Expect = 1.4
Identities = 20/62 (32%), Positives = 29/62 (46%), Gaps = 4/62 (6%)
Query: 111 LRDKRMLRFHESVVAVLESNIQNGPAYFNCFSSYPVCLRDPHVTTCVDLDIKIPNEMFID 170
L R LRF E+ ++ESN N + +Y +D + + DI I NE +D
Sbjct: 5 LEKSRFLRFLENTTKIIESNNSN----YELRGAYFQVRQDKIILISSNEDISIKNEYLLD 60
Query: 171 GI 172
GI
Sbjct: 61 GI 62
>AMPN_PLUXY (P91887) Aminopeptidase N precursor (EC 3.4.11.2)
(Microsomal aminopeptidase) (APN1)
Length = 946
Score = 32.0 bits (71), Expect = 1.4
Identities = 19/81 (23%), Positives = 36/81 (43%), Gaps = 8/81 (9%)
Query: 139 NCFSSYPVCLRDPHVTTCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDNLFGIKAVRDY 198
N FSSY + D H+ +KI +D + P+++ + + N+FG+ R Y
Sbjct: 103 NLFSSYTLATDDTHL-------LKIQFTRVLDALQPITVEISYSAQYAPNMFGVYVSR-Y 154
Query: 199 TVNETEVTEINAENSSRITRK 219
N V+ + ++ R+
Sbjct: 155 VENGATVSLVTSQLQPTFARR 175
>YVNF_AZOCH (P24423) Hypothetical protein in vnfD 5'region
Length = 516
Score = 30.8 bits (68), Expect = 3.1
Identities = 20/56 (35%), Positives = 29/56 (51%), Gaps = 4/56 (7%)
Query: 83 GYLHIGLVQ-VAVKPLTRLRYCDLPILICLRDKRMLRFHESVVAVLESNIQNGPAY 137
GYL IG VQ + +P RLRY +L + L + +LR+ +V+ Q G Y
Sbjct: 330 GYLTIGQVQRIGAQPRYRLRYPNLEVKASLNEALLLRYLPEPASVMR---QTGQLY 382
>VS06_ROTBS (Q00734) VP6 protein (Major capsid protein) (Major
internal structural protein)
Length = 395
Score = 30.4 bits (67), Expect = 4.0
Identities = 18/64 (28%), Positives = 31/64 (48%), Gaps = 7/64 (10%)
Query: 16 NDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPYQ-------SIS 68
N+ L ++N I +RT Y+ +I P + V++ F V GPY ++S
Sbjct: 265 NNGQLVDMVRNMGIATVRTFDSYRITIDMIRPAAMTQYVQQLFPVGGPYSHQAAYMLTLS 324
Query: 69 ILDS 72
+LD+
Sbjct: 325 VLDA 328
Score = 29.6 bits (65), Expect = 6.8
Identities = 14/46 (30%), Positives = 23/46 (49%), Gaps = 3/46 (6%)
Query: 16 NDSNLEQKLQNWSIPKIRTNQVYK---TSIFEFNPDYEIKTVEKSF 58
N +N Q ++NWS R N VY+ +FE+ Y + ++ F
Sbjct: 128 NFNNSSQSIKNWSAQSRRENPVYEYKNPMVFEYRNSYILHRADQQF 173
>VGLF_PI2HT (P26629) Fusion glycoprotein precursor [Contains: Fusion
glycoprotein F2; Fusion glycoprotein F1]
Length = 551
Score = 30.4 bits (67), Expect = 4.0
Identities = 28/102 (27%), Positives = 43/102 (41%), Gaps = 19/102 (18%)
Query: 133 NGPAYFNCFSSYPVCLRDPHVTTCVD------LDIKIPNEMFIDGIPPVSIMYRFCYKVT 186
NG Y NC S C PHV + D +DIK +EM +D F +++T
Sbjct: 377 NGVLYANCKSLLCRCADPPHVVSQDDTQGISIIDIKRCSEMMLD---------TFSFRIT 427
Query: 187 DNLFGIKAVRDYTV---NETEVTEINAENSSRITRKILKHDE 225
+ F V D+++ N ++ ++ N K LK E
Sbjct: 428 -STFNATYVTDFSMINANIVHLSPLDLSNQINSINKSLKSAE 468
>VGLF_PI2HG (P27286) Fusion glycoprotein precursor [Contains: Fusion
glycoprotein F2; Fusion glycoprotein F1]
Length = 551
Score = 30.4 bits (67), Expect = 4.0
Identities = 28/102 (27%), Positives = 43/102 (41%), Gaps = 19/102 (18%)
Query: 133 NGPAYFNCFSSYPVCLRDPHVTTCVD------LDIKIPNEMFIDGIPPVSIMYRFCYKVT 186
NG Y NC S C PHV + D +DIK +EM +D F +++T
Sbjct: 377 NGVLYANCKSLLCRCADPPHVVSQDDTQGISIIDIKRCSEMMLD---------TFSFRIT 427
Query: 187 DNLFGIKAVRDYTV---NETEVTEINAENSSRITRKILKHDE 225
+ F V D+++ N ++ ++ N K LK E
Sbjct: 428 -STFNATYVTDFSMINANIVHLSPLDLSNQINSINKSLKSAE 468
>VGLF_PI2H (P25467) Fusion glycoprotein precursor [Contains: Fusion
glycoprotein F2; Fusion glycoprotein F1]
Length = 551
Score = 30.4 bits (67), Expect = 4.0
Identities = 28/102 (27%), Positives = 43/102 (41%), Gaps = 19/102 (18%)
Query: 133 NGPAYFNCFSSYPVCLRDPHVTTCVD------LDIKIPNEMFIDGIPPVSIMYRFCYKVT 186
NG Y NC S C PHV + D +DIK +EM +D F +++T
Sbjct: 377 NGVLYANCKSLLCRCADPPHVVSQDDTQGISIIDIKRCSEMMLD---------TFSFRIT 427
Query: 187 DNLFGIKAVRDYTV---NETEVTEINAENSSRITRKILKHDE 225
+ F V D+++ N ++ ++ N K LK E
Sbjct: 428 -STFNATYVTDFSMINANIVHLSPLDLSNQINSINKSLKSAE 468
>SAP4_YEAST (P53036) SIT4-associating protein SAP4
Length = 818
Score = 30.4 bits (67), Expect = 4.0
Identities = 21/62 (33%), Positives = 30/62 (47%), Gaps = 4/62 (6%)
Query: 5 TLNQHANNSEENDSNLEQKLQNWSIPKIRTNQVYKTSIFEFNPDYEIKTVEKSFAVQGPY 64
TL NN++ NDSN QK + K N++Y T F+ + D + SF + PY
Sbjct: 502 TLEDKCNNNDSNDSNDNQKQKKNIKKKFHDNELYST--FDTSDDNIDDDDDMSFEI--PY 557
Query: 65 QS 66
S
Sbjct: 558 VS 559
>RPOC_HAEDU (Q7VKL8) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta' chain)
(RNA polymerase beta' subunit)
Length = 1420
Score = 29.3 bits (64), Expect = 8.9
Identities = 20/55 (36%), Positives = 30/55 (54%), Gaps = 8/55 (14%)
Query: 189 LFGIKAVRDYTVNETEV------TEINAENSSRITRKILKHDEIT--YPQEWLQG 235
L G+ AV DY VNE + +IN ++ I R++L+ IT Y E+L+G
Sbjct: 1223 LRGVHAVTDYIVNEVQEVYRLQGVKINDKHIEVIVRQMLRKAVITNAYDSEFLEG 1277
>MTB1_BACSQ (Q9LAI2) Modification methylase BslI (EC 2.1.1.113) (N-4
cytosine-specific methyltransferase BslI) (M.BslI)
Length = 912
Score = 29.3 bits (64), Expect = 8.9
Identities = 20/80 (25%), Positives = 33/80 (41%), Gaps = 1/80 (1%)
Query: 150 DPHVTTCVDLDIKIPNEMFIDGIPPVSIMYRFCYKVTDNLFGIKAVRDYTVNETEVTEIN 209
DP V + +D D K E F + + YK+ + + A+ T++ET E
Sbjct: 18 DPSVVSLIDEDAKKLEEQFPKALKHPVVDEEIVYKILCEKYNLNALNVKTISETLNKEYK 77
Query: 210 -AENSSRITRKILKHDEITY 228
NS +K L + + Y
Sbjct: 78 FGRNSKTALKKYLDYGKEEY 97
>K6A1_RAT (Q63531) Ribosomal protein S6 kinase alpha 1 (EC 2.7.1.37)
(S6K-alpha 1) (90 kDa ribosomal protein S6 kinase 1)
(p90-RSK 1) (Ribosomal S6 kinase 1) (RSK-1) (pp90RSK1)
(MAP kinase-activated protein kinase 1a) (MAPKAPK1A)
Length = 735
Score = 29.3 bits (64), Expect = 8.9
Identities = 37/154 (24%), Positives = 65/154 (42%), Gaps = 14/154 (9%)
Query: 55 EKSFAVQGPYQSISILDSRIANQHVRKYGYLHIGLVQVAVKPLTRLRYCDLPILICLRDK 114
+ F + P S I S A+Q R + ++ GL++ KP R L ++
Sbjct: 352 DTEFTSRTPRDSPGIPPSAGAHQLFRGFSFVATGLMEDDSKP--RATQAPLHSVVQQLHG 409
Query: 115 RMLRFHESVVAVLESNIQNGPAYFNCFSSYPVCLRDPHVTTCVDLDIKIPNEMFIDGIPP 174
+ L F + + ++ I G SY VC R H T ++ +K+ ++ D
Sbjct: 410 KNLVFSDGYI--VKETIGVG--------SYSVCKRCVHKATNMEYAVKVIDKSKRDPSEE 459
Query: 175 VSIMYRFCYKVTDNLFGIKAVRDYTVNETEVTEI 208
+ I+ R Y N+ +K V D + + VTE+
Sbjct: 460 IEILLR--YGQHPNIITLKDVYDDSKHVYLVTEL 491
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,980,189
Number of Sequences: 164201
Number of extensions: 1079703
Number of successful extensions: 2404
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 2400
Number of HSP's gapped (non-prelim): 16
length of query: 242
length of database: 59,974,054
effective HSP length: 107
effective length of query: 135
effective length of database: 42,404,547
effective search space: 5724613845
effective search space used: 5724613845
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0344.6


