

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0342b.6
(427 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
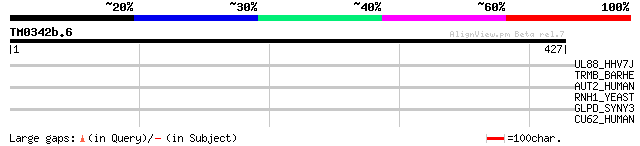
Score E
Sequences producing significant alignments: (bits) Value
UL88_HHV7J (P52364) Protein U59 34 0.82
TRMB_BARHE (Q6G4W3) tRNA (guanine-N(7)-)-methyltransferase (EC 2... 31 6.9
AUT2_HUMAN (Q8WXX7) Autism susceptibility gene 2 protein 31 6.9
RNH1_YEAST (Q04740) Ribonuclease H (EC 3.1.26.4) (RNase H) 30 9.1
GLPD_SYNY3 (P74257) Glycerol-3-phosphate dehydrogenase (EC 1.1.9... 30 9.1
CU62_HUMAN (Q9NYP8) Putative protein C21orf62 (B37) 30 9.1
>UL88_HHV7J (P52364) Protein U59
Length = 347
Score = 33.9 bits (76), Expect = 0.82
Identities = 15/46 (32%), Positives = 24/46 (51%)
Query: 322 WKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSC 367
WK C L ++N A + +ILN S LL+ EF + +++ C
Sbjct: 10 WKDCTLYVNNETATVHEILNSDLSELLQLKTEFVSMTDLCVYITGC 55
>TRMB_BARHE (Q6G4W3) tRNA (guanine-N(7)-)-methyltransferase (EC
2.1.1.33) (tRNA(m7G46)-methyltransferase)
Length = 233
Score = 30.8 bits (68), Expect = 6.9
Identities = 20/75 (26%), Positives = 31/75 (40%), Gaps = 9/75 (12%)
Query: 309 NILVPEGSDPSGHWKSCKLNIHNCNANLIDILNRFRSSLLKALNEFQQRKQIGMFVNSCF 368
++ P+ HWK +N+ N LNRF + +LK +F+ I +VN
Sbjct: 133 DLFYPDPWPKKKHWKRRFINMKN--------LNRF-ARVLKTGKKFRFASDIESYVNWTL 183
Query: 369 IHCQTWRGETWYSPN 383
HC W + N
Sbjct: 184 YHCSNHHSFEWEAKN 198
>AUT2_HUMAN (Q8WXX7) Autism susceptibility gene 2 protein
Length = 1259
Score = 30.8 bits (68), Expect = 6.9
Identities = 16/45 (35%), Positives = 21/45 (46%)
Query: 40 LSLAFFSQLHHHHHSHSLSNLIPFTPLPNAILRGAICLDGSAPGF 84
L F H H H+H ++ FTP P+AI AI + P F
Sbjct: 520 LRTEFHQHQHQHQHTHQHTHQHTFTPFPHAIPPTAIMPTPAPPMF 564
>RNH1_YEAST (Q04740) Ribonuclease H (EC 3.1.26.4) (RNase H)
Length = 348
Score = 30.4 bits (67), Expect = 9.1
Identities = 15/31 (48%), Positives = 18/31 (57%), Gaps = 1/31 (3%)
Query: 150 WNKVMIRYCDGASFA-GHPESEKGSGLFFRG 179
+NK M YCDG+SF G S G G +F G
Sbjct: 184 YNKSMNVYCDGSSFGNGTSSSRAGYGAYFEG 214
>GLPD_SYNY3 (P74257) Glycerol-3-phosphate dehydrogenase (EC
1.1.99.5)
Length = 553
Score = 30.4 bits (67), Expect = 9.1
Identities = 26/113 (23%), Positives = 47/113 (41%), Gaps = 17/113 (15%)
Query: 114 SRRKFTALGSSKYMETVVPFSGILSSDRSQNPDFFNWNKVMIRYCDGASFAGHPESEKGS 173
S R + + + + ++ F L S R +P F + + R A + KG
Sbjct: 103 SSRAYWEIQAGMILYDILSFDKTLPSHRMLSPQQF---QQLFR-------AAEKKGLKGG 152
Query: 174 GLFFRGQIIWEAIMDELLSIGMSKAKQALLSGCSAGG-------LATLVHCDD 219
+F GQ+ + +D +++ KA A+L+ + G L T +HC D
Sbjct: 153 AQYFDGQVEYAERLDLEVTLSAQKAGAAMLNYVAVKGLEKGENNLITAIHCQD 205
>CU62_HUMAN (Q9NYP8) Putative protein C21orf62 (B37)
Length = 247
Score = 30.4 bits (67), Expect = 9.1
Identities = 13/42 (30%), Positives = 22/42 (51%)
Query: 276 MEPSKCLFPSEILKNIKTPVFLVHSAYDFWQIRNILVPEGSD 317
+EPS+C +L ++ T + V S + WQ+ N L E +
Sbjct: 202 LEPSQCQATKAMLSHLFTSIITVSSCLELWQMMNNLQKESGN 243
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.138 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,672,229
Number of Sequences: 164201
Number of extensions: 2204601
Number of successful extensions: 7756
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 7748
Number of HSP's gapped (non-prelim): 10
length of query: 427
length of database: 59,974,054
effective HSP length: 113
effective length of query: 314
effective length of database: 41,419,341
effective search space: 13005673074
effective search space used: 13005673074
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0342b.6


