

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.9
(434 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
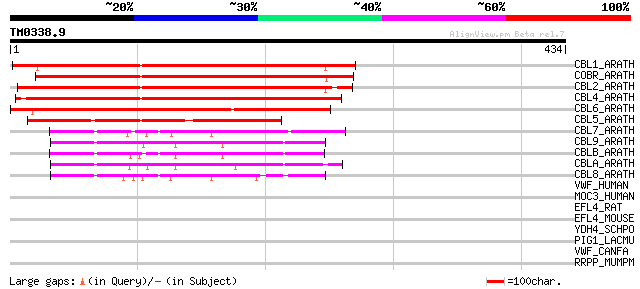
Score E
Sequences producing significant alignments: (bits) Value
CBL1_ARATH (Q9SRT7) COBRA-like protein 1 precursor 419 e-117
COBR_ARATH (Q94KT8) COBRA protein precursor (Cell expansion prot... 398 e-110
CBL2_ARATH (Q8L8Q7) COBRA-like protein 2 precursor 380 e-105
CBL4_ARATH (Q9LFW3) COBRA-like protein 4 precursor 373 e-103
CBL6_ARATH (O04500) COBRA-like protein 6 precursor 252 1e-66
CBL5_ARATH (Q9FME5) COBRA-like protein 5 precursor 202 1e-51
CBL7_ARATH (Q8GZ17) COBRA-like protein 7 precursor 105 2e-22
CBL9_ARATH (Q9FJ13) COBRA-like protein 9 precursor 102 3e-21
CBLB_ARATH (Q9T045) COBRA-like protein 11 precursor 101 4e-21
CBLA_ARATH (Q9LJU0) COBRA-like protein 10 precursor 96 2e-19
CBL8_ARATH (Q9LIB6) COBRA-like protein 8 precursor 90 1e-17
VWF_HUMAN (P04275) Von Willebrand factor precursor (vWF) [Contai... 32 4.2
MOC3_HUMAN (O95396) Molybdenum cofactor synthesis protein 3 (Mol... 32 4.2
EFL4_RAT (Q9QYP0) Multiple EGF-like-domain protein 4 (Multiple e... 31 5.4
EFL4_MOUSE (P60882) Multiple EGF-like-domain protein 4 31 5.4
YDH4_SCHPO (Q92349) Hypothetical protein C6G9.04 in chromosome I 31 7.1
PIG1_LACMU (P60591) Gamma-phospholipase A2 inhibitor LNF1 precursor 31 7.1
VWF_CANFA (Q28295) Von Willebrand factor precursor (vWF) 30 9.3
RRPP_MUMPM (P16595) RNA polymerase alpha subunit (EC 2.7.7.48) (... 30 9.3
>CBL1_ARATH (Q9SRT7) COBRA-like protein 1 precursor
Length = 452
Score = 419 bits (1076), Expect = e-117
Identities = 188/274 (68%), Positives = 225/274 (81%), Gaps = 7/274 (2%)
Query: 3 SLLLFRFVLPCTLLLLLP--SSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQ 60
S + F+F + L+ + SEAYDPLDP+GNIT++WDI++WTGDGYVA VT+ NFQQ
Sbjct: 9 SSIFFKFGISIIFLVSFSGLTPSEAYDPLDPSGNITVKWDIITWTGDGYVATVTVYNFQQ 68
Query: 61 YRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGA 120
YRHIQ PGW+LGW+WAK E+IW M GGQTTEQGDCSKFKG + PHCCKK P VDL+PG+
Sbjct: 69 YRHIQAPGWTLGWSWAKREVIWGMNGGQTTEQGDCSKFKGTI-PHCCKKTPSVVDLLPGS 127
Query: 121 PYSEQVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTC 180
PY++Q+ANCC+GGVL S QDPA AV++F++ VG+AGTT K+VR+PKNFT K+PGPGYTC
Sbjct: 128 PYNQQIANCCRGGVLNSWAQDPATAVSAFQLTVGQAGTTNKTVRVPKNFTLKAPGPGYTC 187
Query: 181 GPARIVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPT 240
PA+IVKPTRF+ DKRRVTQA+M+WNVTCTYSQFLA K PTCCVSLS+FYN TIVSCPT
Sbjct: 188 SPAKIVKPTRFIGTDKRRVTQALMTWNVTCTYSQFLAQKTPTCCVSLSSFYNKTIVSCPT 247
Query: 241 CACGC----QSGNCVDPNSPNLTSVISNPGNNGH 270
C+CGC Q GNCVDP P + SVI NPG N +
Sbjct: 248 CSCGCRNTSQPGNCVDPKGPRIASVIPNPGKNAY 281
>COBR_ARATH (Q94KT8) COBRA protein precursor (Cell expansion
protein)
Length = 456
Score = 398 bits (1023), Expect = e-110
Identities = 172/254 (67%), Positives = 210/254 (81%), Gaps = 6/254 (2%)
Query: 21 SSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEI 80
+S+EAYD LDP GNIT++WD++SWT DGYVA VT+ NFQ+YRHIQ PGW+LGW WAK E+
Sbjct: 32 TSTEAYDALDPEGNITMKWDVMSWTPDGYVAVVTMFNFQKYRHIQSPGWTLGWKWAKKEV 91
Query: 81 IWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVLTSLKQ 140
IW M+G QTTEQGDCSK+KG + PHCCKKDP VDL+PG PY++Q+ANCCKGGV+ S Q
Sbjct: 92 IWSMVGAQTTEQGDCSKYKGNI-PHCCKKDPTVVDLLPGTPYNQQIANCCKGGVMNSWVQ 150
Query: 141 DPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPTRFLRYDKRRVT 200
DPA A +SF++ VG AGTT K+VR+P+NFT PGPGYTCGPA+IV+PT+F+ D RR T
Sbjct: 151 DPATAASSFQISVGAAGTTNKTVRVPRNFTLMGPGPGYTCGPAKIVRPTKFVTTDTRRTT 210
Query: 201 QAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQ-----SGNCVDPNS 255
QAMM+WN+TCTYSQFLA + PTCCVSLS+FYN+TIV CPTCACGCQ SG C+DP++
Sbjct: 211 QAMMTWNITCTYSQFLAQRTPTCCVSLSSFYNETIVGCPTCACGCQNNRTESGACLDPDT 270
Query: 256 PNLTSVISNPGNNG 269
P+L SV+S P G
Sbjct: 271 PHLASVVSPPTKKG 284
>CBL2_ARATH (Q8L8Q7) COBRA-like protein 2 precursor
Length = 441
Score = 380 bits (976), Expect = e-105
Identities = 165/266 (62%), Positives = 210/266 (78%), Gaps = 8/266 (3%)
Query: 7 FRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQE 66
F + C+ +++EAYD LDP GNITI+WDI+SWTGDGYVA VT+ NFQQYRHI+
Sbjct: 10 FLLLFLCSWTSFTFTTTEAYDALDPYGNITIKWDIMSWTGDGYVAVVTIFNFQQYRHIEA 69
Query: 67 PGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQV 126
PGW LGW+W K E+IW M+GGQ TEQGDCSKFKG + PHCCKK P VDL+PG PY++Q+
Sbjct: 70 PGWQLGWSWMKKEVIWSMVGGQATEQGDCSKFKGNI-PHCCKKTPAIVDLLPGTPYNQQI 128
Query: 127 ANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIV 186
+NCC+GGV+++ QDPA A++SF++ VG++GTT +VR P+N T K+PGPGYTCGPA++V
Sbjct: 129 SNCCRGGVISAWAQDPATAISSFQISVGQSGTTNTTVRAPRNITLKAPGPGYTCGPAKLV 188
Query: 187 KPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGC- 245
KP+RF+ DKRR TQ++++WN+TCTYSQFLA K PTCCVSLS FYN+TIV CPTC+CGC
Sbjct: 189 KPSRFISADKRRKTQSLLTWNITCTYSQFLARKTPTCCVSLSAFYNETIVPCPTCSCGCQ 248
Query: 246 ---QSGNCVDPNSPNLTSVISNPGNN 268
Q+G CVD P + SV+ G N
Sbjct: 249 NSSQAGTCVD---PKIASVVPALGKN 271
>CBL4_ARATH (Q9LFW3) COBRA-like protein 4 precursor
Length = 431
Score = 373 bits (958), Expect = e-103
Identities = 170/256 (66%), Positives = 202/256 (78%), Gaps = 5/256 (1%)
Query: 5 LLFRFVLPCTLLLLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAVTLNNFQQYRHI 64
LLF F C ++ ++ AYDPLDP+GNITI+WDI+SWT DGYVA VT+NNFQ YRHI
Sbjct: 3 LLFSF---CFFFFMIIFTATAYDPLDPSGNITIKWDIMSWTADGYVATVTMNNFQIYRHI 59
Query: 65 QEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSE 124
Q PGW+LGWTWAK E+IW M+G QTTEQGDCSKFKG V PHCCKK P VDL+PG PY++
Sbjct: 60 QNPGWTLGWTWAKKEVIWSMVGAQTTEQGDCSKFKGNV-PHCCKKTPTVVDLLPGVPYNQ 118
Query: 125 QVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPAR 184
Q +NCCKGGV+ + QDP+ AV+ F+V G AGTT K+V+LPKNFT PGPGYTCGPA+
Sbjct: 119 QFSNCCKGGVIGAWGQDPSAAVSQFQVSAGLAGTTNKTVKLPKNFTLLGPGPGYTCGPAK 178
Query: 185 IVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACG 244
IV T FL DKRR TQA+M+WNVTCTYSQFLA K P+CCVS S+FYNDTI CP+CACG
Sbjct: 179 IVPSTVFLTTDKRRKTQALMTWNVTCTYSQFLARKHPSCCVSFSSFYNDTITPCPSCACG 238
Query: 245 CQS-GNCVDPNSPNLT 259
C++ +CV +S LT
Sbjct: 239 CENKKSCVKADSKILT 254
>CBL6_ARATH (O04500) COBRA-like protein 6 precursor
Length = 454
Score = 252 bits (644), Expect = 1e-66
Identities = 121/259 (46%), Positives = 161/259 (61%), Gaps = 10/259 (3%)
Query: 1 MDSLLLFRFVLPCTLL-------LLLPSSSEAYDPLDPNGNITIRWDILSWTGDGYVAAV 53
M +LLL V+ C++L ++ ++ YDPLDP G I I+WD+L + + V
Sbjct: 4 MLNLLLVVTVILCSILSPTRFMIMIDKMVADGYDPLDPFGKIIIKWDLLLSSPGQHHVQV 63
Query: 54 TLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQGDCSKFKGQVS-PHCCKKDPV 112
TL N Q+YRH+++PGW L W W E+IW M G +TTEQG+CS F + PHCC + P
Sbjct: 64 TLENMQEYRHVEKPGWKLSWHWLNQEVIWDMKGAETTEQGNCSAFASSGNLPHCCLERPT 123
Query: 113 AVDLIPGAPYSEQVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFK 172
VDL+PGA + QVANCC+GGVLTS+ QD AN V++F + VG + + +P NF
Sbjct: 124 IVDLLPGASLNVQVANCCRGGVLTSMSQDHANHVSAFHMTVGSSPDGPEEFNMPSNFDIG 183
Query: 173 SPGPGYTCGPARIVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYN 232
PGY+C A V PT+F RR TQA+ +W C YSQF ++ +P CCVSLS FY
Sbjct: 184 V--PGYSCDNATSVSPTKFSTDKGRRKTQALATWEAVCVYSQFRSSPSPKCCVSLSAFYY 241
Query: 233 DTIVSCPTCACGCQSGNCV 251
IV CPTC+CGC S +CV
Sbjct: 242 QNIVPCPTCSCGCSSSHCV 260
>CBL5_ARATH (Q9FME5) COBRA-like protein 5 precursor
Length = 204
Score = 202 bits (515), Expect = 1e-51
Identities = 95/199 (47%), Positives = 132/199 (65%), Gaps = 10/199 (5%)
Query: 15 LLLLLPSSSEAYDPLDPN-GNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGW 73
LLL+ S + + L N GNIT++WD+L+WT DGYVA VT N+Q+ R I PGW + W
Sbjct: 11 LLLVSFSCLISTEALTSNYGNITVKWDLLNWTPDGYVAVVTAYNYQKQRSI--PGWKMSW 68
Query: 74 TWAKNEIIWQMIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGG 133
K E+IW M+G +TT QG CS FKG + P C + P VDL+PG P+++Q+ANCCK G
Sbjct: 69 RGTKKEVIWNMLGAKTTGQGGCSMFKGNI-PQSCVRKPTVVDLLPGTPFNQQIANCCKSG 127
Query: 134 VLTSLKQDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPGPGYTCGPARIVKPTRFLR 193
VL + ++F++ VG AG + K+ R+P NF F +P Y CGP++ V+PTRF
Sbjct: 128 VLKP------GSESAFQLSVGSAGNSVKTARMPANFMFTAPKQQYICGPSKNVRPTRFTT 181
Query: 194 YDKRRVTQAMMSWNVTCTY 212
DKRR+T A+M+WN+TC +
Sbjct: 182 ADKRRITAALMTWNITCVF 200
>CBL7_ARATH (Q8GZ17) COBRA-like protein 7 precursor
Length = 661
Score = 105 bits (262), Expect = 2e-22
Identities = 70/251 (27%), Positives = 114/251 (44%), Gaps = 26/251 (10%)
Query: 32 NGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTT- 90
+G++TI +D++ Y+A VT+ N + W L + W ++E I+ M G +
Sbjct: 216 DGDLTIMYDVVRSYSSNYMAQVTMENHNPLGRLDN--WKLSFDWMRDEFIYTMKGAYPSI 273
Query: 91 -EQGDCSKFKGQVSPH----------CCKKDPVAVDLIPGAPYSEQ----VANCCKGGVL 135
+ DC G + H C + P +DL P Y++ + CC+ G +
Sbjct: 274 VDSSDC--VDGPQAKHYQDLDFSNVLSCARRPTVIDL-PPTKYNDSTFGLIPFCCRNGTI 330
Query: 136 TSLKQDPANAVASFRVVVGRA--GTTTKSVRLPKNFTFKSP-GPGYTCGPARIVKPTRFL 192
DP+ + + F++ V + ++ P+N+ P Y CGP V P++F+
Sbjct: 331 LPRSMDPSKSSSVFQMQVYKMPPDLNISALSPPQNWRINGTLNPDYKCGPPVRVSPSQFV 390
Query: 193 RYDKRRVTQ-AMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQSGNCV 251
+ A SW V C +Q A +P CCVS S ++ND+IV C TCACGC S
Sbjct: 391 DPSGLPSNRTAFASWQVVCNITQPKDA-SPRCCVSFSAYFNDSIVPCKTCACGCSSNKAA 449
Query: 252 DPNSPNLTSVI 262
S S++
Sbjct: 450 RACSATAPSLL 460
>CBL9_ARATH (Q9FJ13) COBRA-like protein 9 precursor
Length = 663
Score = 102 bits (253), Expect = 3e-21
Identities = 66/232 (28%), Positives = 102/232 (43%), Gaps = 20/232 (8%)
Query: 33 GNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQ 92
G++TI +D++ Y+ VT+ N + W L + W ++E I +M G T
Sbjct: 216 GDLTIMYDVIRAYDQNYLTEVTMENHNPLGRLDH--WELSFDWMRDEFIQKMQGAYPTVV 273
Query: 93 GDCSKFKGQVS----------PHCCKKDPVAVDLIPGAPYSEQVAN---CCKGGVLTSLK 139
G S C++ P+ +DL P + N CC+ G +
Sbjct: 274 DATKCIFGPQSLIYTGLDFADVLTCERRPIIIDLPPTKKDDSTLGNIPSCCRNGTILPRI 333
Query: 140 QDPANAVASFRVVVGRAGTTTKSVRL--PKNFTFKSP-GPGYTCGPARIVKPTRFLRYDK 196
DP+ +V+ F + V + L P+N+ K P Y+CGP V PT +
Sbjct: 334 MDPSKSVSVFTMQVAKMPPDFNRSALFPPQNWRIKGTLNPDYSCGPPVRVTPTFYPDPSG 393
Query: 197 RRVTQAMM-SWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQS 247
++ SW + C +Q + P CCVS S F+ND+I+ C TCACGC S
Sbjct: 394 MPTNKSSFASWQIVCNITQ-AKTEIPKCCVSFSAFFNDSIIPCNTCACGCVS 444
>CBLB_ARATH (Q9T045) COBRA-like protein 11 precursor
Length = 668
Score = 101 bits (251), Expect = 4e-21
Identities = 72/235 (30%), Positives = 104/235 (43%), Gaps = 26/235 (11%)
Query: 32 NGNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTE 91
+G++ I +D+L Y+A VT++N + W+L W W + E I M G E
Sbjct: 218 HGDLNIIYDVLQAYASSYMAQVTIDNTSPLGRLDH--WNLTWEWMRGEFIHSMRGAYAAE 275
Query: 92 QG--DCSKFKG----------QVSPHCCKKDPVAVDLIPGAPYSEQVAN----CCKGGVL 135
+ +C K QV+ C+K P+ DL P + V CCK G L
Sbjct: 276 KNTLECLSSKAGQFYGDLDFSQVAN--CQKKPIIKDL-PAERKDDNVTGKLPFCCKNGTL 332
Query: 136 TSLKQDPANAVASFRVVVGRAGTTTKSVRL--PKNFTFKS-PGPGYTCGPARIVKPTRFL 192
DP+ + A F++ V + P+N+ P Y CGP V T F
Sbjct: 333 LPTHMDPSKSKAIFQLQVYKVPPDQNRTAFFPPRNWKIDGIVNPTYKCGPPIRVDATPFP 392
Query: 193 RYDK-RRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQ 246
+ T A+ SW V C ++ +A CCVS S FYND+ + C TCACGC+
Sbjct: 393 DPSGLQATTYAIASWQVICNITK-PKPQAARCCVSFSAFYNDSAIPCNTCACGCK 446
>CBLA_ARATH (Q9LJU0) COBRA-like protein 10 precursor
Length = 672
Score = 95.9 bits (237), Expect = 2e-19
Identities = 75/245 (30%), Positives = 109/245 (43%), Gaps = 23/245 (9%)
Query: 33 GNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGGQTTEQ 92
G++ I +D+L Y+A VT++N + W+L + W + E I M G T ++
Sbjct: 229 GDLNIVYDVLQSFDSNYLAQVTIDNDNPLGRLDR--WNLTFEWMRGEFINTMRGAYTHKK 286
Query: 93 --GDCSKFK-GQVSPHC-------CKKDPVAVDLIPGAPYSEQVAN---CCKGGVLTSLK 139
+C K GQ C++ P DL P CCK G L
Sbjct: 287 DPSECLYSKAGQYYKDLDFSQVMNCQRKPAISDLPPEKKEDNMTGKLPFCCKNGTLLPPI 346
Query: 140 QDPANAVASFRVVVGRAGTTTKSVRLPKNFTFKSPG---PGYTCGPARIVKPTRFLRYDK 196
DP+ + + F++ V + L +K G P Y CGP V P++F
Sbjct: 347 MDPSKSRSMFQLQVFKLPPDLNRTALYPPQHWKIDGVLNPQYKCGPPVRVDPSQFPDPSG 406
Query: 197 R-RVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCACGCQSGNCVDPNS 255
VT A+ SW V C ++ A+A CCVS S FYN++ V C TCACGC N +D ++
Sbjct: 407 LLAVTYAISSWQVVCNITK-PKAQASRCCVSFSAFYNNSAVPCNTCACGC---NDIDTDT 462
Query: 256 PNLTS 260
N S
Sbjct: 463 CNANS 467
>CBL8_ARATH (Q9LIB6) COBRA-like protein 8 precursor
Length = 653
Score = 89.7 bits (221), Expect = 1e-17
Identities = 65/238 (27%), Positives = 107/238 (44%), Gaps = 33/238 (13%)
Query: 33 GNITIRWDILSWTGDGYVAAVTLNNFQQYRHIQEPGWSLGWTWAKNEIIWQMIGG--QTT 90
G++TI +D+ Y A VT+ N + W L + W K+E ++ G
Sbjct: 210 GDLTIMYDVTRAYQSSYSAQVTIENHNLLGRLDN--WDLSFMWMKDEFLFSTKGAYPSVV 267
Query: 91 EQGDC-----SKFKGQV---SPHCCKKDPVAVDLIPGAPYSE----QVANCCKGGVLTSL 138
+ DC +K+ + + C + P +DL P Y++ ++ CC+ G +
Sbjct: 268 DSSDCITGPQAKYYKDLDFSNVMSCARRPHIIDL-PLTKYNDTNVGRIPYCCRNGTILPR 326
Query: 139 KQDPANAVASFRVVVGRA--GTTTKSVRLPKNFTFKSP-GPGYTCGPARIVKPTRF---- 191
DP + + F++ V + S+ P+++ K P Y CGP V ++F
Sbjct: 327 SMDPEKSKSVFQIEVYKMPPDLNISSITPPQSWQIKGNLNPDYKCGPPLRVSSSQFPDTS 386
Query: 192 -LRYDKRRVTQAMMSWNVTCTYSQFLAAKAPTCCVSLSTFYNDTIVSCPTCAC-GCQS 247
L +K A SW V C +Q P CCVS S+++ND+++ C TCAC GC S
Sbjct: 387 GLPSNK----SAFASWQVVCNITQ---PTPPKCCVSFSSYFNDSVIPCKTCACGGCSS 437
>VWF_HUMAN (P04275) Von Willebrand factor precursor (vWF) [Contains:
Von Willebrand antigen II]
Length = 2813
Score = 31.6 bits (70), Expect = 4.2
Identities = 20/64 (31%), Positives = 27/64 (41%), Gaps = 5/64 (7%)
Query: 208 VTCTYSQFLAAKAPTC-CVSLSTFYNDTIVSCPTCACGCQSGNCVDPNSPN----LTSVI 262
V CT AKAPTC ++ + CP C C +C P P+ L +
Sbjct: 2289 VNCTTQPCPTAKAPTCGLCEVARLRQNADQCCPEYECVCDPVSCDLPPVPHCERGLQPTL 2348
Query: 263 SNPG 266
+NPG
Sbjct: 2349 TNPG 2352
>MOC3_HUMAN (O95396) Molybdenum cofactor synthesis protein 3
(Molybdopterin synthase sulfurylase) (MPT synthase
sulfurylase)
Length = 460
Score = 31.6 bits (70), Expect = 4.2
Identities = 15/52 (28%), Positives = 25/52 (47%)
Query: 84 MIGGQTTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGVL 135
++ G+ +F+GQ++ + P + P P +E V NC GGVL
Sbjct: 194 VLAGRPLVSASALRFEGQITVYHYDGGPCYRCIFPQPPPAETVTNCADGGVL 245
>EFL4_RAT (Q9QYP0) Multiple EGF-like-domain protein 4 (Multiple
epidermal growth factor-like domains 8) (Fragment)
Length = 874
Score = 31.2 bits (69), Expect = 5.4
Identities = 14/31 (45%), Positives = 19/31 (61%)
Query: 224 CVSLSTFYNDTIVSCPTCACGCQSGNCVDPN 254
C + T N T V P CA GC +G+CV+P+
Sbjct: 235 CKTGYTMDNVTGVCRPVCAQGCVNGSCVEPD 265
>EFL4_MOUSE (P60882) Multiple EGF-like-domain protein 4
Length = 2330
Score = 31.2 bits (69), Expect = 5.4
Identities = 14/31 (45%), Positives = 19/31 (61%)
Query: 224 CVSLSTFYNDTIVSCPTCACGCQSGNCVDPN 254
C + T N T V P CA GC +G+CV+P+
Sbjct: 1691 CKTGYTMDNVTGVCRPVCAQGCVNGSCVEPD 1721
>YDH4_SCHPO (Q92349) Hypothetical protein C6G9.04 in chromosome I
Length = 1318
Score = 30.8 bits (68), Expect = 7.1
Identities = 38/154 (24%), Positives = 65/154 (41%), Gaps = 14/154 (9%)
Query: 124 EQVANCCKGGVLTSLKQDPANAVASFRVVVGRAGTTTKSVRL--PKNFTFKSPGPGYTCG 181
+ V+N KG +L+++P N ++R + +++ L KN + + P P + G
Sbjct: 271 DYVSNLSKGHTTNALEENPFNLGPNYRASTRKCNRIKEAINLFAAKNGSMEVP-PKVSVG 329
Query: 182 PARIVKPTRFLRYDKRRVTQAMMSWNVTCTYSQFLAA------KAPTCCVSLSTFYNDTI 235
+ + +FL+ + + T +M+ + Q LAA KAPT S
Sbjct: 330 DSVLTTSQKFLQVIREK-TALLMNQDSNSVQPQALAAAESPTTKAPTTKAPTSEAPPKGH 388
Query: 236 VSCPTCACG----CQSGNCVDPNSPNLTSVISNP 265
V G QS N V+P S + S + NP
Sbjct: 389 VKQLAKQLGNIYMPQSINNVEPTSHSSISKVVNP 422
>PIG1_LACMU (P60591) Gamma-phospholipase A2 inhibitor LNF1 precursor
Length = 200
Score = 30.8 bits (68), Expect = 7.1
Identities = 14/42 (33%), Positives = 22/42 (52%), Gaps = 2/42 (4%)
Query: 93 GDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQVANCCKGGV 134
G S +G+++ CC+K+P L PG P S+ C G +
Sbjct: 83 GHHSYIRGRIN--CCEKEPCEDQLFPGLPLSKPNGYYCPGAI 122
>VWF_CANFA (Q28295) Von Willebrand factor precursor (vWF)
Length = 2813
Score = 30.4 bits (67), Expect = 9.3
Identities = 19/64 (29%), Positives = 26/64 (39%), Gaps = 5/64 (7%)
Query: 208 VTCTYSQFLAAKAPTC-CVSLSTFYNDTIVSCPTCACGCQSGNC----VDPNSPNLTSVI 262
V CT A+APTC ++ + CP C C +C V P L +
Sbjct: 2289 VNCTLQPCPTARAPTCGPCEVARLRQNAEQCCPEYECVCDLVSCDLPPVPPCEDGLQMTL 2348
Query: 263 SNPG 266
+NPG
Sbjct: 2349 TNPG 2352
>RRPP_MUMPM (P16595) RNA polymerase alpha subunit (EC 2.7.7.48)
(Nucleocapsid phosphoprotein)
Length = 391
Score = 30.4 bits (67), Expect = 9.3
Identities = 30/103 (29%), Positives = 44/103 (42%), Gaps = 20/103 (19%)
Query: 89 TTEQGDCSKFKGQVSPHCCKKDPVAVDLIPGAPYSEQV-----ANCCKGGVLTSLKQDPA 143
T+ QG SK +G + P+ V G QV A +GG++T++ QDP
Sbjct: 62 TSHQGSKSKGRGSGAR------PIIVSSSEGGTGGTQVPEPLFAQTGQGGIVTTVYQDPT 115
Query: 144 -NAVASFRVV----VGRAGTTTKSVRLPKNFT----FKSPGPG 177
S+R V +G+ + V P+ T FK GPG
Sbjct: 116 IQPTGSYRSVELAKIGKERMINRFVEKPRTSTPVTEFKRGGPG 158
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.334 0.145 0.504
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,759,577
Number of Sequences: 164201
Number of extensions: 2131089
Number of successful extensions: 6436
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 6381
Number of HSP's gapped (non-prelim): 31
length of query: 434
length of database: 59,974,054
effective HSP length: 113
effective length of query: 321
effective length of database: 41,419,341
effective search space: 13295608461
effective search space used: 13295608461
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0338.9


