

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0338.4
(197 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
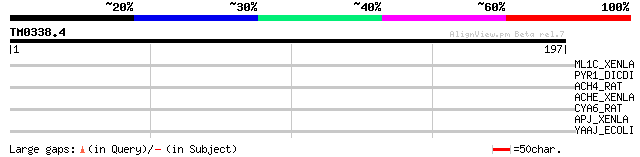
Score E
Sequences producing significant alignments: (bits) Value
ML1C_XENLA (P49219) Melatonin receptor type 1C (MEL-1C-R) 36 0.050
PYR1_DICDI (P20054) Protein PYR1-3 [Includes: Glutamine-dependen... 34 0.25
ACH4_RAT (P09483) Neuronal acetylcholine receptor protein, alpha... 32 1.2
ACHE_XENLA (P49580) Acetylcholine receptor protein, epsilon chai... 31 2.1
CYA6_RAT (Q03343) Adenylate cyclase, type VI (EC 4.6.1.1) (ATP p... 30 3.6
APJ_XENLA (P79960) G protein-coupled receptor APJ homolog (Angio... 29 6.1
YAAJ_ECOLI (P30143) Putative transporter yaaJ 29 8.0
>ML1C_XENLA (P49219) Melatonin receptor type 1C (MEL-1C-R)
Length = 420
Score = 36.2 bits (82), Expect = 0.050
Identities = 24/82 (29%), Positives = 41/82 (49%), Gaps = 7/82 (8%)
Query: 19 SCTTRSKLDLTVSSLPVVVVVVDLLVPCVLISNFTCIRCYGIT---KHFGRYAFKSSLTD 75
SCT + SS + VVVV +VP +++ F +R + + KH R FK LT
Sbjct: 181 SCTFAQTVS---SSYTITVVVVHFIVPLSVVT-FCYLRIWVLVIQVKHRVRQDFKQKLTQ 236
Query: 76 IPLVSLMRSLIILCVYSICDAP 97
L + + ++ ++++C AP
Sbjct: 237 TDLRNFLTMFVVFVLFAVCWAP 258
>PYR1_DICDI (P20054) Protein PYR1-3 [Includes: Glutamine-dependent
carbamoyl-phosphate synthase (EC 6.3.5.5); Aspartate
carbamoyltransferase (EC 2.1.3.2); Dihydroorotase (EC
3.5.2.3)]
Length = 2185
Score = 33.9 bits (76), Expect = 0.25
Identities = 22/82 (26%), Positives = 44/82 (52%), Gaps = 10/82 (12%)
Query: 104 YLGTVTLCSI-LSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHLKKSWGMPVLFLSSVV 162
+LG + C+I S+ S + C+ VN+R+ ++ + K+ G P+ F+S+ V
Sbjct: 630 HLGVIGECNIQYSLNPYSEEYCIIEVNARLSRSSALA--------SKATGYPLAFISAKV 681
Query: 163 FSLGHIVVACRTSCRARRSSCF 184
+LG+ + A R + + ++CF
Sbjct: 682 -ALGYDLAALRNTITKKTTACF 702
>ACH4_RAT (P09483) Neuronal acetylcholine receptor protein, alpha-4
chain precursor
Length = 630
Score = 31.6 bits (70), Expect = 1.2
Identities = 27/90 (30%), Positives = 47/90 (52%), Gaps = 7/90 (7%)
Query: 40 VDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLIILCVYSICDAPTL 99
++L++PC+LIS T + Y + G K +L L+SL ++ +L + I +PT
Sbjct: 250 INLIIPCLLISCLTVLVFY-LPSECGE---KVTLCISVLLSL--TVFLLLITEIIPSPTS 303
Query: 100 SHGPYLGTVTLCSILSIVLLSVKTCVFTVN 129
P +G L +++ V LS+ VF +N
Sbjct: 304 LVIPLIGEYLLFTMI-FVTLSIVITVFVLN 332
>ACHE_XENLA (P49580) Acetylcholine receptor protein, epsilon chain
precursor
Length = 504
Score = 30.8 bits (68), Expect = 2.1
Identities = 25/109 (22%), Positives = 54/109 (48%), Gaps = 7/109 (6%)
Query: 39 VVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLIILCVYSICDAPT 98
+++++VPCVLIS F + Y + G S++ V L +++ + + + +
Sbjct: 245 IINIIVPCVLIS-FLVVLVYFLPAKAGGQKCTVSIS----VLLAQTVFLFLIAQMVPETS 299
Query: 99 LSHGPYLGTVTLCSILSIVLLSVKTCVFTVNSRIEAEASASLKRQRLHL 147
LS P +G L ++ + L V +CV +N + + ++ +L + H+
Sbjct: 300 LS-VPLIGKY-LMFVMFVSTLIVLSCVIVLNVSLRSPSTHNLSTKVKHM 346
>CYA6_RAT (Q03343) Adenylate cyclase, type VI (EC 4.6.1.1) (ATP
pyrophosphate-lyase) (Ca(2+)-inhibitable adenylyl
cyclase)
Length = 1166
Score = 30.0 bits (66), Expect = 3.6
Identities = 15/58 (25%), Positives = 31/58 (52%)
Query: 64 FGRYAFKSSLTDIPLVSLMRSLIILCVYSICDAPTLSHGPYLGTVTLCSILSIVLLSV 121
+ RY F+ + + + L+ + L++ + + AP L Y+ +T S+L +VL+ V
Sbjct: 138 YQRYFFQMNQSSLTLLMAVLVLLMAVLLTFHAAPALPQPAYVALLTCASVLFVVLMVV 195
>APJ_XENLA (P79960) G protein-coupled receptor APJ homolog
(Angiotensin receptor related protein)
(Mesenchyme-associated serpentine receptor)
Length = 353
Score = 29.3 bits (64), Expect = 6.1
Identities = 29/109 (26%), Positives = 51/109 (46%), Gaps = 11/109 (10%)
Query: 30 VSSLPVVVVVVDLLVPCVLISNFTCIRCYGITKHFGRYAFKSSLTDIPLVSLMRSLIILC 89
+ L ++ V L+P +L++ F C +T HF K L+ ++ +L++
Sbjct: 202 IGGLSILTTVPGFLLPLLLMTIFYCFIGGKVTMHFQNLK-KEEQKKKRLLKIIITLVV-- 258
Query: 90 VYSICDAP-----TLSHGPYLGTVTL-CSILSIV--LLSVKTCVFTVNS 130
V++IC P T+ +G + L CS +I+ L TC+ VNS
Sbjct: 259 VFAICWLPFHILKTIHFLDLMGFLELSCSTQNIIVSLHPYATCLAYVNS 307
>YAAJ_ECOLI (P30143) Putative transporter yaaJ
Length = 476
Score = 28.9 bits (63), Expect = 8.0
Identities = 12/36 (33%), Positives = 23/36 (63%)
Query: 20 CTTRSKLDLTVSSLPVVVVVVDLLVPCVLISNFTCI 55
CT + + T+ SLP++ + D+++ C+ I+N T I
Sbjct: 392 CTFATVIGGTLLSLPLMWQLADIIMACMAITNLTAI 427
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.343 0.149 0.469
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,483,673
Number of Sequences: 164201
Number of extensions: 634926
Number of successful extensions: 3334
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 3331
Number of HSP's gapped (non-prelim): 7
length of query: 197
length of database: 59,974,054
effective HSP length: 105
effective length of query: 92
effective length of database: 42,732,949
effective search space: 3931431308
effective search space used: 3931431308
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0338.4


