

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0336.12
(267 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
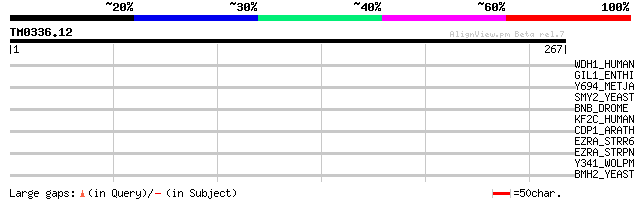
Score E
Sequences producing significant alignments: (bits) Value
WDH1_HUMAN (O75717) WD repeat and HMG-box DNA binding protein 1 ... 33 0.55
GIL1_ENTHI (P32022) Galactose-inhibitable lectin 170 kDa subunit 32 1.6
Y694_METJA (Q58105) Hypothetical protein MJ0694 32 2.1
SMY2_YEAST (P32909) SMY2 protein 31 2.7
BNB_DROME (P29746) Bangles and beads protein 31 2.7
KF2C_HUMAN (Q99661) Kinesin-like protein KIF2C (Mitotic centrome... 30 4.7
CDP1_ARATH (Q06850) Calcium-dependent protein kinase, isoform AK... 30 4.7
EZRA_STRR6 (Q8DQE5) Septation ring formation regulator ezrA 30 6.1
EZRA_STRPN (Q97RK0) Septation ring formation regulator ezrA 30 6.1
Y341_WOLPM (Q73I41) Hypothetical UPF0085 protein WD0341 30 8.0
BMH2_YEAST (P34730) BMH2 protein 30 8.0
>WDH1_HUMAN (O75717) WD repeat and HMG-box DNA binding protein 1
(Acidic nucleoplasmic DNA-binding protein 1) (And-1)
Length = 1129
Score = 33.5 bits (75), Expect = 0.55
Identities = 17/58 (29%), Positives = 28/58 (47%)
Query: 11 LSSSTMNEREPHLRNVCPKPPLGLKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFAPA 68
LS +T NE+ P ++ + PKP S ++R K ++ K++N L PA
Sbjct: 949 LSRTTNNEKSPIIKPLIPKPKPKQASAASYFQKRNSQTNKTEEVKEENLKNVLSETPA 1006
>GIL1_ENTHI (P32022) Galactose-inhibitable lectin 170 kDa subunit
Length = 1276
Score = 32.0 bits (71), Expect = 1.6
Identities = 23/107 (21%), Positives = 51/107 (47%), Gaps = 24/107 (22%)
Query: 87 VNQLVVKCQFARRKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTEILEIE----- 141
+N+ +K F +++ + I K EV WD + ++E + E ++I
Sbjct: 78 INKTNIKQDFCQKEYAYPIE--------KYEVDWDNVP-----VDEQRIESVDINGKTCF 124
Query: 142 --LDEQPIFHKEINSQSTHTT----WEISDHDFTGGHAMTYRIHHLE 182
++P+ + +N++ T+ T +++ DF GG ++T+R + E
Sbjct: 125 KYAAKRPLAYVYLNTKMTYATKTEAYDVCRMDFIGGRSITFRSFNTE 171
>Y694_METJA (Q58105) Hypothetical protein MJ0694
Length = 414
Score = 31.6 bits (70), Expect = 2.1
Identities = 15/49 (30%), Positives = 25/49 (50%)
Query: 16 MNEREPHLRNVCPKPPLGLKRTTSAMKEREKMWRKMQKSKKQNSSTTLK 64
+N+R +R PKPP K+ K++EK +K + KK++ K
Sbjct: 351 LNKRVEEIRRKYPKPPKKKKKEKPKAKKKEKKGKKEKSKKKKDKKKDKK 399
>SMY2_YEAST (P32909) SMY2 protein
Length = 790
Score = 31.2 bits (69), Expect = 2.7
Identities = 18/75 (24%), Positives = 38/75 (50%), Gaps = 8/75 (10%)
Query: 13 SSTMNEREPHLRNVCPKPPLGLKRTT--------SAMKEREKMWRKMQKSKKQNSSTTLK 64
SS + + +++ K P G+K ++ + ++E++K+W K+Q S KQ ST+
Sbjct: 570 SSLPSNQTIDIKSQFQKSPKGMKESSPLKELEDPNFIEEQKKLWEKVQSSSKQVKSTSSA 629
Query: 65 FAPATFYAKLLNVGK 79
+ + + + GK
Sbjct: 630 STTTSSWTTVTSKGK 644
>BNB_DROME (P29746) Bangles and beads protein
Length = 442
Score = 31.2 bits (69), Expect = 2.7
Identities = 15/41 (36%), Positives = 22/41 (53%)
Query: 6 PFGLELSSSTMNEREPHLRNVCPKPPLGLKRTTSAMKEREK 46
P + S+ T+ E P L+ P P G + T A+KE+EK
Sbjct: 296 PVDQQKSTETVAESAPVLKTNAPLAPAGATKVTEAVKEQEK 336
>KF2C_HUMAN (Q99661) Kinesin-like protein KIF2C (Mitotic
centromere-associated kinesin) (MCAK) (Kinesin-like
protein 6)
Length = 725
Score = 30.4 bits (67), Expect = 4.7
Identities = 19/59 (32%), Positives = 30/59 (50%), Gaps = 1/59 (1%)
Query: 9 LELSSSTMNEREPHLRNVCPKPPLG-LKRTTSAMKEREKMWRKMQKSKKQNSSTTLKFA 66
L + S M E+ +R P+ ++R + +KE EKM K ++ K QNS +K A
Sbjct: 162 LRMVSEEMEEQVHSIRGSSSANPVNSVRRKSCLVKEVEKMKNKREEKKAQNSEMRMKRA 220
>CDP1_ARATH (Q06850) Calcium-dependent protein kinase, isoform AK1
(EC 2.7.1.-) (CDPK)
Length = 610
Score = 30.4 bits (67), Expect = 4.7
Identities = 30/102 (29%), Positives = 43/102 (41%), Gaps = 7/102 (6%)
Query: 1 MNSKPPFGLELSSSTMNEREPHLRNVCPKPPLGLKRTTSAMKEREKMWRKMQKSKKQNSS 60
+ SKP E S T E +P PK P +KR +SA E + ++ ++ K+ S
Sbjct: 93 LESKPETKQETKSETKPESKPD-PPAKPKKPKHMKRVSSAGLRTESVLQRKTENFKEFYS 151
Query: 61 TTLKFAPATFYAKLLNVGKYNFIPTTVNQLVVKCQFARRKLI 102
K F L V K TT + K A+RKL+
Sbjct: 152 LGRKLGQGQFGTTFLCVEK-----TTGKEFACK-SIAKRKLL 187
>EZRA_STRR6 (Q8DQE5) Septation ring formation regulator ezrA
Length = 575
Score = 30.0 bits (66), Expect = 6.1
Identities = 22/76 (28%), Positives = 32/76 (41%), Gaps = 5/76 (6%)
Query: 129 FIEENKTEILEIELDEQPIFHKEINSQSTHTTWEISDHDFTGGHAMTYRIHHLEFASGDL 188
F N TE L +EL++ K IN +S EI+ +D TY I + L
Sbjct: 455 FTASNNTEDLMVELEQ-----KMINIESVTRVLEIATNDMEALETETYNIVQYATLTEQL 509
Query: 189 GQHYENMLKCDERLMK 204
Q+ DER+ +
Sbjct: 510 LQYSNRYRSFDERIQE 525
>EZRA_STRPN (Q97RK0) Septation ring formation regulator ezrA
Length = 575
Score = 30.0 bits (66), Expect = 6.1
Identities = 22/76 (28%), Positives = 32/76 (41%), Gaps = 5/76 (6%)
Query: 129 FIEENKTEILEIELDEQPIFHKEINSQSTHTTWEISDHDFTGGHAMTYRIHHLEFASGDL 188
F N TE L +EL++ K IN +S EI+ +D TY I + L
Sbjct: 455 FTASNNTEDLMVELEQ-----KMINIESVTRVLEIATNDMEALETETYNIVQYATLTEQL 509
Query: 189 GQHYENMLKCDERLMK 204
Q+ DER+ +
Sbjct: 510 LQYSNRYRSFDERIQE 525
>Y341_WOLPM (Q73I41) Hypothetical UPF0085 protein WD0341
Length = 276
Score = 29.6 bits (65), Expect = 8.0
Identities = 28/98 (28%), Positives = 46/98 (46%), Gaps = 20/98 (20%)
Query: 110 KNKKFKAEVPWDRISAIRAFIEENKTEILEIELDEQPIFHKEINSQSTHTTWEISDHDFT 169
K+ K ++P+ AI + I + LEIE DE+ H EIN++ F
Sbjct: 80 KDNCIKLKIPY---RAILSHIIREISSYLEIEKDEKLDLHTEINNEY-----------FQ 125
Query: 170 GGHAMTYRIHHLEFASGDLGQHYENMLKCDERLMKLSQ 207
A+ Y I+H D GQ+ +++ K D L+ +S+
Sbjct: 126 RIEAINYTINH------DDGQNIQDIDKADIILVGVSR 157
>BMH2_YEAST (P34730) BMH2 protein
Length = 272
Score = 29.6 bits (65), Expect = 8.0
Identities = 21/96 (21%), Positives = 42/96 (42%), Gaps = 4/96 (4%)
Query: 99 RKLIWEILDSSKNKKFKAEVPWDRISAIRAFIEENKTEILEIELDEQPIFHKEINSQSTH 158
R+ W I+ S + K+ E ++ IR++ + +TE+ +I D + + +T
Sbjct: 56 RRASWRIVSSIEQKEESKEKSEHQVELIRSYRSKIETELTKISDDILSVLDSHLIPSATT 115
Query: 159 TTWEISDHDFTGGHAMTYRIHHLEFASGDLGQHYEN 194
++ + G Y + EF+SGD + N
Sbjct: 116 GESKVFYYKMKG----DYHRYLAEFSSGDAREKATN 147
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.133 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,308,036
Number of Sequences: 164201
Number of extensions: 1265415
Number of successful extensions: 3872
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 3868
Number of HSP's gapped (non-prelim): 12
length of query: 267
length of database: 59,974,054
effective HSP length: 108
effective length of query: 159
effective length of database: 42,240,346
effective search space: 6716215014
effective search space used: 6716215014
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0336.12


