

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
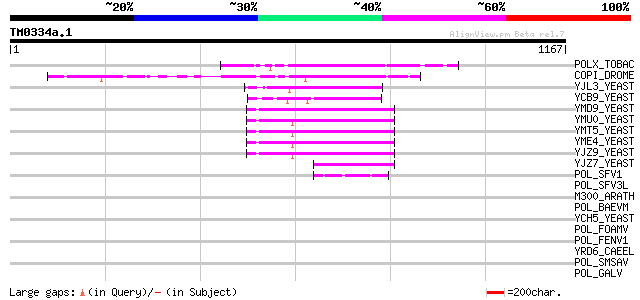
Query= TM0334a.1
(1167 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 174 1e-42
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 147 2e-34
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 86 4e-16
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 76 6e-13
YMD9_YEAST (Q03434) Transposon Ty1 protein B 69 1e-10
YMU0_YEAST (Q04670) Transposon Ty1 protein B 68 1e-10
YMT5_YEAST (Q04214) Transposon Ty1 protein B 68 1e-10
YME4_YEAST (Q04711) Transposon Ty1 protein B 68 1e-10
YJZ9_YEAST (P47100) Transposon Ty1 protein B 68 1e-10
YJZ7_YEAST (P47098) Transposon Ty1 protein B 67 4e-10
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 45 9e-04
POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.2... 44 0.003
M300_ARATH (P93293) Hypothetical mitochondrial protein AtMg00300... 44 0.003
POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.2... 42 0.013
YCH5_YEAST (P25601) Transposon Ty5-1 16.0 kDa hypothetical protein 41 0.017
POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse transcript... 41 0.017
POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse transcript... 41 0.017
YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II 41 0.022
POL_SMSAV (P03359) Pol polyprotein [Contains: Reverse transcript... 40 0.037
POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC 3.4.23... 39 0.11
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 174 bits (442), Expect = 1e-42
Identities = 149/519 (28%), Positives = 235/519 (44%), Gaps = 54/519 (10%)
Query: 443 DWIIDSGASDHICFNKLCFDTLNRIKPVSVRLPNGISIQTCYAGVVSITKQILLTHVL-- 500
+W++D+ AS H + F +V++ N + G + I + T VL
Sbjct: 293 EWVVDTAASHHATPVRDLFCRYVAGDFGTVKMGNTSYSKIAGIGDICIKTNVGCTLVLKD 352
Query: 501 --YVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSL------KEDDLY 552
+VP+ NLIS L R+ E F + K R+ GSL LY
Sbjct: 353 VRHVPDLRMNLISGIAL-DRDGYESYFA---------NQKWRLTKGSLVIAKGVARGTLY 402
Query: 553 HLVVTSPSSTCFAIPNVVERISDSDQLIPPGALWHFRLGHLSHD--RILALNALYPSIDV 610
+ + C N + D LWH R+GH+S +ILA +L
Sbjct: 403 R----TNAEICQGELNAAQDEISVD-------LWHKRMGHMSEKGLQILAKKSLISYAKG 451
Query: 611 SKHFVCDICHLAKLKRKMFPDSLHNAKCNFDLLHMDIWGPISVSSVHSHRYFLTVLDDHS 670
+ CD C K R F S DL++ D+ GP+ + S+ ++YF+T +DD S
Sbjct: 452 TTVKPCDYCLFGKQHRVSFQTSSERKLNILDLVYSDVCGPMEIESMGGNKYFVTFIDDAS 511
Query: 671 RFVWVILLKRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEFMLHDF---YSAHGILH 727
R +WV +LK K +V Q + F ALV+ + +K +RSDNG E+ +F S+HGI H
Sbjct: 512 RKLWVYILKTKDQVFQVFQKFHALVERETGRKLKRLRSDNGGEYTSREFEEYCSSHGIRH 571
Query: 728 QRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRIQSKVLEN 787
+++ TPQ NG ER ++ I+ R++L + LPK WG A+ A +L++R S L
Sbjct: 572 EKTVPGTPQHNGVAERMNRTIVEKVRSMLRMAKLPKSFWGEAVQTACYLINRSPSVPLAF 631
Query: 788 KSPYEILFGGKKIDLQELRVFGSLCFASTSSSSRSKLDSRARKCVFLGFKQGVKGFVLLD 847
+ P E ++ K++ L+VFG FA R+KLD ++ C+F+G+ G+ L D
Sbjct: 632 EIP-ERVWTNKEVSYSHLKVFGCRAFAHVPKEQRTKLDDKSIPCIFIGYGDEEFGYRLWD 690
Query: 848 LNSHEIFVSRDVTFHEQILPYKSATSTAWTCLDLEVKDQSS---SSLNIPETTSAAQELI 904
++ SRDV F E S T D+ K ++ + + IP T++
Sbjct: 691 PVKKKVIRSRDVVFRE---------SEVRTAADMSEKVKNGIIPNFVTIPSTSNNPTSAE 741
Query: 905 SDNENFSNT*LPCHTQSETIIEEENTQSETLIEEEEDDT 943
S + S + + E+ Q + +EE E T
Sbjct: 742 STTDEVSE-----QGEQPGEVIEQGEQLDEGVEEVEHPT 775
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 147 bits (370), Expect = 2e-34
Identities = 182/816 (22%), Positives = 334/816 (40%), Gaps = 151/816 (18%)
Query: 79 PLNGRNYNAWA*SMKRVLVAKNKFKFINGEIPIAVPGDANYEAWDRCNNLIHSWILNSVT 138
P +G Y W ++ +L ++ K ++G +P V ++W + S I+ ++
Sbjct: 10 PFDGEKYAIWKFRIRALLAEQDVLKVVDGLMPNEVD-----DSWKKAERCAKSTIIEYLS 64
Query: 139 SSIANSIVFVENACDAWRDLRDRFSQGVLVRIAELQNEIGNLKQNT-LSVNDYY------ 191
S N A +L + + L L+ + +LK ++ +S+ ++
Sbjct: 65 DSFLNFATSDITARQILENLDAVYERKSLASQLALRKRLLSLKLSSEMSLLSHFHIFDEL 124
Query: 192 -TEIKTLWEELEQYRPIPQCRCVVP-CRCEAIEHAKMFREQDNAIRFLLGLNETFSVVNS 249
+E+ ++E+ I +P C I + E++ + F+ +++
Sbjct: 125 ISELLAAGAKIEEMDKISHLLITLPSCYDGIITAIETLSEENLTLAFVKN-----RLLDQ 179
Query: 250 QILMSNPLPPIAKVVSLAMQHERQSETGENEESKSLVKMAEGKKTYGKGKAPGSSSSGSG 309
+I + N +K V A+ H + T +N K+ ++ + KK +
Sbjct: 180 EIKIKNDHNDTSKKVMNAIVHNNNN-TYKNNLFKN--RVTKPKKIF-------------- 222
Query: 310 YKSTGKY---CTHCKKPGHTVDVCYRLHGYPTTSKSKPAFNNVSHINNIT*GYDSEDDQE 366
K KY C HC + GH C+ H I + E++++
Sbjct: 223 -KGNSKYKVKCHHCGREGHIKKDCF-------------------HYKRILNNKNKENEKQ 262
Query: 367 ESSKSQRGNGDLFTADQYKTIMAMIQQVTATSASQKHTESGKAFANLVTKSAVCGANTAG 426
+ + G I M+++V TS CG
Sbjct: 263 VQTATSHG------------IAFMVKEVNNTSVMDN-----------------CG----- 288
Query: 427 NGKHSTLSYSSQYDSRDWIIDSGASDHICFNKLCF-DTLNRIKPVSVRLPN-GISIQTCY 484
+++DSGASDH+ ++ + D++ + P+ + + G I
Sbjct: 289 -----------------FVLDSGASDHLINDESLYTDSVEVVPPLKIAVAKQGEFIYATK 331
Query: 485 AGVVSITK--QILLTHVLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIG 542
G+V + +I L VL+ E NL+SV +L + G F + T + G
Sbjct: 332 RGIVRLRNDHEITLEDVLFCKEAAGNLMSVKRLQEA-------GMSIEFDKSGVTISKNG 384
Query: 543 LGSLKEDDLYHLVVTSPSSTCFAIPNVVERISDSDQLIPPGALWHFRLGHLSHDRILALN 602
L +K + + V F ++ + ++ +L WH R GH+S ++L +
Sbjct: 385 LMVVKNSGMLNNVPVIN----FQAYSINAKHKNNFRL------WHERFGHISDGKLLEIK 434
Query: 603 ALYPSIDVSK----HFVCDICH------LAKLKRKMFPDSLHNAKCNFDLLHMDIWGPIS 652
D S C+IC A+L K D H + F ++H D+ GPI+
Sbjct: 435 RKNMFSDQSLLNNLELSCEICEPCLNGKQARLPFKQLKDKTHIKRPLF-VVHSDVCGPIT 493
Query: 653 VSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGP 712
++ YF+ +D + + L+K K +V ++FVA + F+ V + DNG
Sbjct: 494 PVTLDDKNYFVIFVDQFTHYCVTYLIKYKSDVFSMFQDFVAKSEAHFNLKVVYLYIDNGR 553
Query: 713 EFM---LHDFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYA 769
E++ + F GI + + +TPQ NG ER + I AR ++ + L K WG A
Sbjct: 554 EYLSNEMRQFCVKKGISYHLTVPHTPQLNGVSERMIRTITEKARTMVSGAKLDKSFWGEA 613
Query: 770 ILHAVFLMSRIQSKVL--ENKSPYEILFGGKKIDLQELRVFGSLCFASTSSSSRSKLDSR 827
+L A +L++RI S+ L +K+PYE ++ KK L+ LRVFG+ + + + K D +
Sbjct: 614 VLTATYLINRIPSRALVDSSKTPYE-MWHNKKPYLKHLRVFGATVYVHI-KNKQGKFDDK 671
Query: 828 ARKCVFLGFKQGVKGFVLLDLNSHEIFVSRDVTFHE 863
+ K +F+G++ GF L D + + V+RDV E
Sbjct: 672 SFKSIFVGYEP--NGFKLWDAVNEKFIVARDVVVDE 705
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 86.3 bits (212), Expect = 4e-16
Identities = 84/315 (26%), Positives = 143/315 (44%), Gaps = 39/315 (12%)
Query: 495 LLTHVLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSLK-EDDLYH 553
LLT+ YVPE +IS + LAK+ + + K+T R+G +K + + +
Sbjct: 472 LLTY--YVPEEESTIISCYDLAKKTKM---------VLSRKYT--RLGNKIIKIKTKIVN 518
Query: 554 LVVTSPSSTCFAIPNVVERISDSDQLIPPGALW----------HFRLGHLSHDRI---LA 600
V+ + P+ +I+ PG H R+GH +I +
Sbjct: 519 GVIHVKMNELIERPSDDSKINAIKPTSSPGFKLNKRSITLEDAHKRMGHTGIQQIENSIK 578
Query: 601 LNALYPSIDVSKH---FVCDICHLAKL-KRKMFPDSLHNAKCNFD---LLHMDIWGPISV 653
N S+D+ K F C C ++K KR + S++N + + MDI+GP+S
Sbjct: 579 HNHYEESLDLIKEPNEFWCQTCKISKATKRNHYTGSMNNHSTDHEPGSSWCMDIFGPVSS 638
Query: 654 SSVHSHRYFLTVLDDHSRFVWVI--LLKRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNG 711
S+ + RY L ++D+++R+ K + ++ + V+TQF V+ I SD G
Sbjct: 639 SNADTKRYMLIMVDNNTRYCMTSTHFNKNAETILAQVRKNIQYVETQFDRKVREINSDRG 698
Query: 712 PEF---MLHDFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGY 768
EF + +++ + GI H + NGR ER + I+ A LL QS+L K W Y
Sbjct: 699 TEFTNDQIEEYFISKGIHHILTSTQDHAANGRAERYIRTIITDATTLLRQSNLRVKFWEY 758
Query: 769 AILHAVFLMSRIQSK 783
A+ A + + ++ K
Sbjct: 759 AVTSATNIRNYLEHK 773
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 75.9 bits (185), Expect = 6e-13
Identities = 78/313 (24%), Positives = 135/313 (42%), Gaps = 47/313 (15%)
Query: 500 LYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQE--KHTKKRIGLGSLKEDDLYHLVVT 557
L+ P Y+L+S+ +LA +N CF + + + + +K D Y L
Sbjct: 513 LHTPNIAYDLLSLSELANQNIT-------ACFTRNTLERSDGTVLAPIVKHGDFYWL--- 562
Query: 558 SPSSTCFAIPNVVERISDSDQLIP------PGALWHFRLGHLSHDRI---LALNAL--YP 606
S + IP+ + +++ ++ P L H LGH + I L NA+
Sbjct: 563 ---SKKYLIPSHISKLTINNVNKSKSVNKYPYPLIHRMLGHANFRSIQKSLKKNAVTYLK 619
Query: 607 SIDV----SKHFVCDICHLAKL--------KRKMFPDSLHNAKCNFDLLHMDIWGPISVS 654
D+ + + C C + K R + +S F LH DI+GP+
Sbjct: 620 ESDIEWSNASTYQCPDCLIGKSTKHRHVKGSRLKYQESYEP----FQYLHTDIFGPVHHL 675
Query: 655 SVHSHRYFLTVLDDHSRFVWVILL--KRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGP 712
+ YF++ D+ +RF WV L +R+ + + +A +K QF+ V VI+ D G
Sbjct: 676 PKSAPSYFISFTDEKTRFQWVYPLHDRREESILNVFTSILAFIKNQFNARVLVIQMDRGS 735
Query: 713 EF---MLHDFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYA 769
E+ LH F++ GI + + +G ER ++ +LN R LL S LP +W A
Sbjct: 736 EYTNKTLHKFFTNRGITACYTTTADSRAHGVAERLNRTLLNDCRTLLHCSGLPNHLWFSA 795
Query: 770 ILHAVFLMSRIQS 782
+ + + + + S
Sbjct: 796 VEFSTIIRNSLVS 808
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 68.6 bits (166), Expect = 1e-10
Identities = 79/329 (24%), Positives = 137/329 (41%), Gaps = 25/329 (7%)
Query: 499 VLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSLKEDDLYHLVVTS 558
VL+ P Y+L+S+++LA + + FT V E+ + + +K D Y +
Sbjct: 89 VLHTPNIAYDLLSLNELAAVD----ITACFTKNVLER-SDGTVLAPIVKYGDFYWVSKKY 143
Query: 559 PSSTCFAIPNVVERISDSDQLIPPGALWHFRLGHLSHDRI---LALNAL--YPSIDVSKH 613
+ ++P + + P H L H + I L N + + DV +
Sbjct: 144 LLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDRS 203
Query: 614 ----FVCDICHLAK-LKRKMFPDS---LHNAKCNFDLLHMDIWGPISVSSVHSHRYFLTV 665
+ C C + K K + S N+ F LH DI+GP+ + YF++
Sbjct: 204 SAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISF 263
Query: 666 LDDHSRFVWVILL--KRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEF---MLHDFY 720
D+ ++F WV L +R+ + +A +K QF V VI+ D G E+ LH F
Sbjct: 264 TDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFL 323
Query: 721 SAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRI 780
+GI + + +G ER ++ +L+ R L S LP +W AI + + + +
Sbjct: 324 EKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSL 383
Query: 781 QSKVLENKSPYEILFGGKKIDLQELRVFG 809
S + + G +D+ L FG
Sbjct: 384 ASPKSKKSARQHAGLAG--LDISTLLPFG 410
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 68.2 bits (165), Expect = 1e-10
Identities = 77/329 (23%), Positives = 134/329 (40%), Gaps = 25/329 (7%)
Query: 499 VLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSLKEDDLYHLVVTS 558
VL+ P Y+L+S+++LA + + FT V E+ + + +K D Y +
Sbjct: 89 VLHTPNIAYDLLSLNELAAVD----ITACFTKNVLER-SDGTVLAPIVKYGDFYWVSKKY 143
Query: 559 PSSTCFAIPNVVERISDSDQLIPPGALWHFRLGH---------LSHDRILALNALYPSID 609
+ ++P + + P H L H L ++ I N
Sbjct: 144 LLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDWS 203
Query: 610 VSKHFVCDICHLAK-LKRKMFPDS---LHNAKCNFDLLHMDIWGPISVSSVHSHRYFLTV 665
+ + C C + K K + S N+ F LH DI+GP+ + YF++
Sbjct: 204 SAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISF 263
Query: 666 LDDHSRFVWVILL--KRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEF---MLHDFY 720
D+ ++F WV L +R+ + +A +K QF V VI+ D G E+ LH F
Sbjct: 264 TDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFL 323
Query: 721 SAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRI 780
+GI + + +G ER ++ +L+ R L S LP +W AI + + + +
Sbjct: 324 EKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSL 383
Query: 781 QSKVLENKSPYEILFGGKKIDLQELRVFG 809
S + + G +D+ L FG
Sbjct: 384 ASPKSKKSARQHAGLAG--LDISTLLPFG 410
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 68.2 bits (165), Expect = 1e-10
Identities = 77/329 (23%), Positives = 134/329 (40%), Gaps = 25/329 (7%)
Query: 499 VLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSLKEDDLYHLVVTS 558
VL+ P Y+L+S+++LA + + FT V E+ + + +K D Y +
Sbjct: 89 VLHTPNIAYDLLSLNELAAVD----ITACFTKNVLER-SDGTVLAPIVKYGDFYWVSKKY 143
Query: 559 PSSTCFAIPNVVERISDSDQLIPPGALWHFRLGH---------LSHDRILALNALYPSID 609
+ ++P + + P H L H L ++ I N
Sbjct: 144 LLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDWS 203
Query: 610 VSKHFVCDICHLAK-LKRKMFPDS---LHNAKCNFDLLHMDIWGPISVSSVHSHRYFLTV 665
+ + C C + K K + S N+ F LH DI+GP+ + YF++
Sbjct: 204 SAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISF 263
Query: 666 LDDHSRFVWVILL--KRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEF---MLHDFY 720
D+ ++F WV L +R+ + +A +K QF V VI+ D G E+ LH F
Sbjct: 264 TDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFL 323
Query: 721 SAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRI 780
+GI + + +G ER ++ +L+ R L S LP +W AI + + + +
Sbjct: 324 EKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSL 383
Query: 781 QSKVLENKSPYEILFGGKKIDLQELRVFG 809
S + + G +D+ L FG
Sbjct: 384 ASPKSKKSARQHAGLAG--LDISTLLPFG 410
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 68.2 bits (165), Expect = 1e-10
Identities = 77/329 (23%), Positives = 134/329 (40%), Gaps = 25/329 (7%)
Query: 499 VLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSLKEDDLYHLVVTS 558
VL+ P Y+L+S+++LA + + FT V E+ + + +K D Y +
Sbjct: 89 VLHTPNIAYDLLSLNELAAVD----ITACFTKNVLER-SDGTVLAPIVKYGDFYWVSKKY 143
Query: 559 PSSTCFAIPNVVERISDSDQLIPPGALWHFRLGH---------LSHDRILALNALYPSID 609
+ ++P + + P H L H L ++ I N
Sbjct: 144 LLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDWS 203
Query: 610 VSKHFVCDICHLAK-LKRKMFPDS---LHNAKCNFDLLHMDIWGPISVSSVHSHRYFLTV 665
+ + C C + K K + S N+ F LH DI+GP+ + YF++
Sbjct: 204 SAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISF 263
Query: 666 LDDHSRFVWVILL--KRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEF---MLHDFY 720
D+ ++F WV L +R+ + +A +K QF V VI+ D G E+ LH F
Sbjct: 264 TDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFL 323
Query: 721 SAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRI 780
+GI + + +G ER ++ +L+ R L S LP +W AI + + + +
Sbjct: 324 EKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSL 383
Query: 781 QSKVLENKSPYEILFGGKKIDLQELRVFG 809
S + + G +D+ L FG
Sbjct: 384 ASPKSKKSARQHAGLAG--LDISTLLPFG 410
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 68.2 bits (165), Expect = 1e-10
Identities = 77/329 (23%), Positives = 134/329 (40%), Gaps = 25/329 (7%)
Query: 499 VLYVPEFTYNLISVHKLAKRNRCEVVFGEFTCFVQEKHTKKRIGLGSLKEDDLYHLVVTS 558
VL+ P Y+L+S+++LA + + FT V E+ + + +K D Y +
Sbjct: 516 VLHTPNIAYDLLSLNELAAVD----ITACFTKNVLER-SDGTVLAPIVKYGDFYWVSKKY 570
Query: 559 PSSTCFAIPNVVERISDSDQLIPPGALWHFRLGH---------LSHDRILALNALYPSID 609
+ ++P + + P H L H L ++ I N
Sbjct: 571 LLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDWS 630
Query: 610 VSKHFVCDICHLAK-LKRKMFPDS---LHNAKCNFDLLHMDIWGPISVSSVHSHRYFLTV 665
+ + C C + K K + S N+ F LH DI+GP+ + YF++
Sbjct: 631 SAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISF 690
Query: 666 LDDHSRFVWVILL--KRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEF---MLHDFY 720
D+ ++F WV L +R+ + +A +K QF V VI+ D G E+ LH F
Sbjct: 691 TDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFL 750
Query: 721 SAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHAVFLMSRI 780
+GI + + +G ER ++ +L+ R L S LP +W AI + + + +
Sbjct: 751 EKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSL 810
Query: 781 QSKVLENKSPYEILFGGKKIDLQELRVFG 809
S + + G +D+ L FG
Sbjct: 811 ASPKSKKSARQHAGLAG--LDISTLLPFG 837
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 66.6 bits (161), Expect = 4e-10
Identities = 48/175 (27%), Positives = 80/175 (45%), Gaps = 7/175 (4%)
Query: 640 FDLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILL--KRKGEVQQ*IKNFVALVKT 697
F LH DI+GP+ + YF++ D+ ++F WV L +R+ + +A +K
Sbjct: 665 FQYLHTDIFGPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKN 724
Query: 698 QFSHHVKVIRSDNGPEF---MLHDFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARA 754
QF V VI+ D G E+ LH F +GI + + +G ER ++ +L+ R
Sbjct: 725 QFQASVLVIQMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRT 784
Query: 755 LLFQSHLPKKMWGYAILHAVFLMSRIQSKVLENKSPYEILFGGKKIDLQELRVFG 809
L S LP +W AI + + + + S + + G +D+ L FG
Sbjct: 785 QLQCSGLPNHLWFSAIEFSTIVRNSLASPKSKKSARQHAGLAG--LDISTLLPFG 837
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 45.4 bits (106), Expect = 9e-04
Identities = 47/160 (29%), Positives = 68/160 (42%), Gaps = 13/160 (8%)
Query: 640 FDLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQF 699
FD ++D GP+ S+ + H L V+D + FVW+ K +K L
Sbjct: 884 FDKFYIDYIGPLPPSNGYLH--VLVVVDSMTGFVWLYPTKAPSTSAT-VKALNMLTSIAI 940
Query: 700 SHHVKVIRSDNGPEFM---LHDFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALL 756
KV+ SD G F D+ GI + S PQ +G+VERK+ I + LL
Sbjct: 941 P---KVLHSDQGAAFTSSTFADWAKEKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTKLL 997
Query: 757 FQSHLPKKMWGYAILHAVFLMSRIQSKVLENKSPYEILFG 796
P K W Y +L V L +P+++LFG
Sbjct: 998 IGR--PAK-W-YDLLPVVQLALNNSYSPSSKYTPHQLLFG 1033
>POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1157
Score = 43.9 bits (102), Expect = 0.003
Identities = 48/160 (30%), Positives = 67/160 (41%), Gaps = 13/160 (8%)
Query: 640 FDLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQF 699
FD +D GP+ S+ + H L V+D + FVW+ K +K L
Sbjct: 886 FDKFFIDYIGPLPPSNGYLH--VLVVVDSMTGFVWLYPTKAPSTSAT-VKALNMLTSIAV 942
Query: 700 SHHVKVIRSDNGPEFM---LHDFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALL 756
KVI SD G F D+ GI + S PQ +G+VERK+ I + LL
Sbjct: 943 P---KVIHSDQGAAFTSATFADWAKNKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTKLL 999
Query: 757 FQSHLPKKMWGYAILHAVFLMSRIQSKVLENKSPYEILFG 796
P K W Y +L V L +P+++LFG
Sbjct: 1000 VGR--PAK-W-YDLLPVVQLALNNSYSPSSKYTPHQLLFG 1035
>M300_ARATH (P93293) Hypothetical mitochondrial protein AtMg00300
(ORF145a) (ORF1451)
Length = 145
Score = 43.9 bits (102), Expect = 0.003
Identities = 25/71 (35%), Positives = 30/71 (42%), Gaps = 2/71 (2%)
Query: 585 LWHFRLGHLSHD--RILALNALYPSIDVSKHFVCDICHLAKLKRKMFPDSLHNAKCNFDL 642
LWH RL H+S +L S VS C+ C K R F H K D
Sbjct: 71 LWHSRLAHMSQRGMELLVKKGFLDSSKVSSLKFCEDCIYGKTHRVNFSTGQHTTKNPLDY 130
Query: 643 LHMDIWGPISV 653
+H D+WG SV
Sbjct: 131 VHSDLWGAPSV 141
>POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1189
Score = 41.6 bits (96), Expect = 0.013
Identities = 41/131 (31%), Positives = 57/131 (43%), Gaps = 20/131 (15%)
Query: 704 KVIRSDNGPEFMLH---DFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSH 760
KVI SDNGP F+ GI + C PQ +G+VER ++ I L ++
Sbjct: 965 KVIGSDNGPAFVSQVSQGLARILGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTLETG 1024
Query: 761 LP--KKMWGYAILHAVFLMSRIQSKVLENKSPYEILFGGKKIDLQELRVFGSLCFASTSS 818
L +++ A+L A +R +PYEIL+GG L F S
Sbjct: 1025 LKDWRRLLSLALLRARNTPNRF------GLTPYEILYGGPPPLSTLLNSF---------S 1069
Query: 819 SSRSKLDSRAR 829
S SK D +AR
Sbjct: 1070 PSNSKTDLQAR 1080
>YCH5_YEAST (P25601) Transposon Ty5-1 16.0 kDa hypothetical protein
Length = 146
Score = 41.2 bits (95), Expect = 0.017
Identities = 25/72 (34%), Positives = 36/72 (49%), Gaps = 1/72 (1%)
Query: 441 SRDWIIDSGASDHICFNKLCFDTLNRIKPVSVRLPNGISIQTCYAGVVSITKQILLTHVL 500
S +WI D+G + H+C ++ F + R G SI +G V+I + L V
Sbjct: 76 SSEWIFDTGCTSHMCHDRSIFSSFTRSSRKDFVRGVGGSIPIMGSGTVNI-GTVQLHDVS 134
Query: 501 YVPEFTYNLISV 512
YVP+ NLISV
Sbjct: 135 YVPDLPVNLISV 146
>POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 886
Score = 41.2 bits (95), Expect = 0.017
Identities = 49/161 (30%), Positives = 69/161 (42%), Gaps = 15/161 (9%)
Query: 640 FDLLHMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQF 699
FD +D GP+ S + Y L V+D + F W+ K +K+ L
Sbjct: 676 FDKFFIDYIGPLPPSQ--GYLYVLVVVDGMTGFTWLYPTKAPSTSAT-VKSLNVLTSIAI 732
Query: 700 SHHVKVIRSDNGPEFMLHDFYS---AHGILHQRSCVNTPQQNGRVERKHQHILNIARALL 756
KVI SD G F F GI + S PQ +VERK+ I + LL
Sbjct: 733 P---KVIHSDQGAAFTSSTFAEWAKERGIHLEFSTPYHPQSGSKVERKNSDIKRLLTKLL 789
Query: 757 FQSHLPKKMWGYAILHAVFL-MSRIQSKVLENKSPYEILFG 796
P K W Y +L V L ++ S VL+ +P+++LFG
Sbjct: 790 VGR--PTK-W-YDLLPVVQLALNNTYSPVLK-YTPHQLLFG 825
>POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 1046
Score = 41.2 bits (95), Expect = 0.017
Identities = 32/99 (32%), Positives = 47/99 (47%), Gaps = 11/99 (11%)
Query: 704 KVIRSDNGPEFMLH---DFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSH 760
KVI SDNGP F+ GI + C PQ +G+VER ++ I L ++
Sbjct: 822 KVIGSDNGPAFVSQVSQGLARTLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTLETG 881
Query: 761 LP--KKMWGYAILHAVFLMSRIQSKVLENKSPYEILFGG 797
L +++ A+L A +R +PYEIL+GG
Sbjct: 882 LKDWRRLLSLALLRARNTPNRF------GLTPYEILYGG 914
>YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II
Length = 1268
Score = 40.8 bits (94), Expect = 0.022
Identities = 33/106 (31%), Positives = 50/106 (47%), Gaps = 16/106 (15%)
Query: 643 LHMDIWGPISVSSVHSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFSHH 702
+H+D GP++ Y L V+D +++ V L + V + L++ FS H
Sbjct: 852 IHIDFAGPLNGC------YLLVVVDAKTKYAEVKLTRSISAVTT-----IDLLEEIFSIH 900
Query: 703 --VKVIRSDNGPEFMLHDFYS---AHGILHQRSCVNTPQQNGRVER 743
+ I SDNG + H F +HGI H+ S V P+ NG ER
Sbjct: 901 GYPETIISDNGTQLTSHLFAQMCQSHGIEHKTSAVYYPRSNGAAER 946
>POL_SMSAV (P03359) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 294
Score = 40.0 bits (92), Expect = 0.037
Identities = 39/144 (27%), Positives = 61/144 (42%), Gaps = 13/144 (9%)
Query: 657 HSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*IKNFVALVKTQFSHHVKVIRSDNGPEFML 716
+ +RY L +D S +V K + + +N L KV+ SDNGP F+
Sbjct: 25 YGNRYLLVFIDTFSGWVEAFPTKTETALTVCKRNSTPL------RIPKVLGSDNGPAFVA 78
Query: 717 H---DFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILHA 773
+ GI + C PQ +G+VER ++ I L ++ K W + A
Sbjct: 79 QVSQGLATQLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLALET--GGKDWVALLPLA 136
Query: 774 VFLMSRIQSKVLENKSPYEILFGG 797
+ S+ +PYEIL+GG
Sbjct: 137 LLRAKNTPSRF--GLTPYEILYGG 158
>POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1165
Score = 38.5 bits (88), Expect = 0.11
Identities = 41/145 (28%), Positives = 64/145 (43%), Gaps = 11/145 (7%)
Query: 657 HSHRYFLTVLDDHSRFVWVILLKRKGEVQQ*I-KNFVALVKTQFSHHVKVIRSDNGPEFM 715
+ ++Y L +D S WV K E + K + + +F KV+ SDNGP F+
Sbjct: 892 YGNKYLLVFIDTFSG--WVEAFPTKTETALIVCKKILEEILPRFGIP-KVLGSDNGPAFV 948
Query: 716 LH---DFYSAHGILHQRSCVNTPQQNGRVERKHQHILNIARALLFQSHLPKKMWGYAILH 772
+ GI + C PQ +G+VER ++ I L ++ K W +L
Sbjct: 949 AQVSQGLATQLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLALET--GGKDW-VTLLP 1005
Query: 773 AVFLMSRIQSKVLENKSPYEILFGG 797
L +R + +PYEIL+GG
Sbjct: 1006 LALLRAR-NTPGRFGLTPYEILYGG 1029
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.331 0.141 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 127,697,538
Number of Sequences: 164201
Number of extensions: 5175112
Number of successful extensions: 15925
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 35
Number of HSP's that attempted gapping in prelim test: 15850
Number of HSP's gapped (non-prelim): 77
length of query: 1167
length of database: 59,974,054
effective HSP length: 121
effective length of query: 1046
effective length of database: 40,105,733
effective search space: 41950596718
effective search space used: 41950596718
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0334a.1


