

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
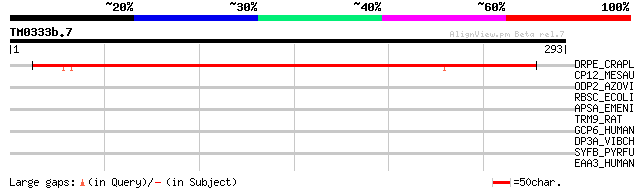
Query= TM0333b.7
(293 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
DRPE_CRAPL (P22242) Desiccation-related protein PCC13-62 precursor 265 8e-71
CP12_MESAU (P24453) Cytochrome P450 1A2 (EC 1.14.14.1) (CYPIA2) ... 33 0.83
ODP2_AZOVI (P10802) Dihydrolipoyllysine-residue acetyltransferas... 32 2.4
RBSC_ECOLI (P04984) Ribose transport system permease protein rbsC 31 3.2
APSA_EMENI (Q00083) Anucleate primary sterigmata protein A 31 4.1
TRM9_RAT (Q91ZY8) Tripartite motif-containing protein 9 (SNAP-25... 30 7.1
GCP6_HUMAN (Q96RT7) Gamma-tubulin complex component 6 (GCP-6) 30 7.1
DP3A_VIBCH (P52022) DNA polymerase III alpha subunit (EC 2.7.7.7) 30 7.1
SYFB_PYRFU (Q8U260) Phenylalanyl-tRNA synthetase beta chain (EC ... 30 9.2
EAA3_HUMAN (P43005) Excitatory amino acid transporter 3 (Sodium-... 30 9.2
>DRPE_CRAPL (P22242) Desiccation-related protein PCC13-62 precursor
Length = 313
Score = 265 bits (678), Expect = 8e-71
Identities = 141/290 (48%), Positives = 188/290 (64%), Gaps = 24/290 (8%)
Query: 13 SSLVLSLFIPIFCSF---------EDVV--DVDLVQFHLNLEFLEAEFYFYGTLGWGLDH 61
S+ ++S F+ + CS +D+ DV L++F LNLE LEAEF+ + G G+D
Sbjct: 9 SAALVSFFLALICSCSYAAWHHEKDDIPKSDVSLLEFPLNLELLEAEFFAWAAFGKGIDE 68
Query: 62 ADPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISKEA 121
+P+LA+GGP P+G Q A L P +RDI +QFA+Q+ GH+RAI V+GFPRPLL++S ++
Sbjct: 69 LEPELAKGGPSPIGVQKANLSPFIRDIIAQFAYQEFGHVRAIQSSVEGFPRPLLDLSAKS 128
Query: 122 FAEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNLVSGL 181
FA V++SAFGK L PPFDPYAN +NYLLA Y++PYVGLTGYVG NPKL+ +R LV+GL
Sbjct: 129 FATVMDSAFGKTLKPPFDPYANDINYLLACYVVPYVGLTGYVGANPKLESPVSRKLVAGL 188
Query: 182 LGAKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLLRNDLGKK------------- 228
L ++GQDA+IR LLYER +V+PY TV E T+++S LRN LG K
Sbjct: 189 LAVEAGQDAIIRALLYERATDKVEPYGITVAEFTNKISELRNKLGDKGVKDLGLIVEPEL 248
Query: 229 QAEGKVVGQVFGLNMYSLTYPRSPEEILRIVYGSGDEHVPGGFFPKGGNG 278
AEGK+ G V + SL +PR+PE L + P F PK G
Sbjct: 249 GAEGKISGNVLAGDKNSLAFPRTPERCLGSCTAAAMRPSPAAFIPKAPTG 298
>CP12_MESAU (P24453) Cytochrome P450 1A2 (EC 1.14.14.1) (CYPIA2)
(P450-MC4) (Hepatic cytochrome P-450MC1)
Length = 513
Score = 33.1 bits (74), Expect = 0.83
Identities = 26/75 (34%), Positives = 37/75 (48%), Gaps = 12/75 (16%)
Query: 211 VYEVTHRLSLLRN---DLGKKQAEGKVV---GQVFGLNMYSLTYPRSPEEILRIVYGSGD 264
+ E H +S L+ ++G + +VV V G + +PR EE+LRIV GS D
Sbjct: 165 IKEANHLVSKLQKLTAEVGHFEPVNQVVESVANVIGAMCFGKNFPRKSEEMLRIVKGSSD 224
Query: 265 --EHVPGG----FFP 273
E+V G FFP
Sbjct: 225 FVENVSSGNAVDFFP 239
>ODP2_AZOVI (P10802) Dihydrolipoyllysine-residue acetyltransferase
component of pyruvate dehydrogenase complex (EC
2.3.1.12) (E2) (Dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase complex)
Length = 637
Score = 31.6 bits (70), Expect = 2.4
Identities = 23/84 (27%), Positives = 36/84 (42%), Gaps = 6/84 (7%)
Query: 62 ADPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAG------HLRAITEEVKGFPRPLL 115
A P A G P G + P +R + +F + A R + E+V+ + + ++
Sbjct: 319 AAPAPAPVGAPSRNGAKVHAGPAVRQLAREFGVELAAINSTGPRGRILKEDVQAYVKAMM 378
Query: 116 NISKEAFAEVINSAFGKPLNPPFD 139
+KEA A S G P PP D
Sbjct: 379 QKAKEAPAAGAASGAGIPPIPPVD 402
>RBSC_ECOLI (P04984) Ribose transport system permease protein rbsC
Length = 321
Score = 31.2 bits (69), Expect = 3.2
Identities = 27/116 (23%), Positives = 51/116 (43%), Gaps = 7/116 (6%)
Query: 131 GKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYV---GTN---PKLQGVRTRNLVSGLLGA 184
G+PL P + + +L A YML + L Y+ G N +L G+ N + ++ +
Sbjct: 167 GRPLGVPTPVWIMGIVFLAAWYMLHHTRLGRYIYALGGNEAATRLSGINV-NKIKIIVYS 225
Query: 185 KSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLLRNDLGKKQAEGKVVGQVFG 240
G A + ++ R +P T YE+ +++ +G++VG + G
Sbjct: 226 LCGLLASLAGIIEVARLSSAQPTAGTGYELDAIAAVVLGGTSLAGGKGRIVGTLIG 281
>APSA_EMENI (Q00083) Anucleate primary sterigmata protein A
Length = 1676
Score = 30.8 bits (68), Expect = 4.1
Identities = 28/104 (26%), Positives = 39/104 (36%), Gaps = 27/104 (25%)
Query: 118 SKEAFAEVINSAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPKLQGVRTRNL 177
S +FA + F NPPF P S GT+P++ T+ +
Sbjct: 1354 SVSSFASELEERFNMQPNPPFAPQGYS------------------TGTDPRMIQAITQTM 1395
Query: 178 VSGLL-----GAKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTH 216
+ L A SG+ + R RRY V PY T+Y H
Sbjct: 1396 IGEFLWKYTRRAVSGEISNTR----HRRYFWVHPYTRTLYWSEH 1435
>TRM9_RAT (Q91ZY8) Tripartite motif-containing protein 9
(SNAP-25-interacting RING finger protein)
Length = 710
Score = 30.0 bits (66), Expect = 7.1
Identities = 23/89 (25%), Positives = 38/89 (41%), Gaps = 9/89 (10%)
Query: 209 WTVYEVTHRLSLLRNDLGKKQAEGKV-----VGQVFGLNMYSLTYPRSPEEILRIVYGSG 263
W +Y +R + N+ + EG + +G + LN +LT+ + E+ I +
Sbjct: 616 WAMYVDNNRSWFMHNNSHTNRTEGGITKGATIGVLLDLNRKTLTFFVNNEQQGPIAF--- 672
Query: 264 DEHVPGGFFPKGGNGNIAKSYLHPNAPAP 292
E+V G FFP + LH P P
Sbjct: 673 -ENVEGLFFPAVSLNRNVQVTLHTGLPVP 700
>GCP6_HUMAN (Q96RT7) Gamma-tubulin complex component 6 (GCP-6)
Length = 1819
Score = 30.0 bits (66), Expect = 7.1
Identities = 19/62 (30%), Positives = 27/62 (42%)
Query: 70 GPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISKEAFAEVINSA 129
G PP + YL RD F +F G L+ + V P+P+L E +V+N
Sbjct: 301 GCPPGHREEPYLTEAGRDAFDKFCRLHQGELQLLAGGVLQAPQPVLVKECELVKDVLNVL 360
Query: 130 FG 131
G
Sbjct: 361 IG 362
>DP3A_VIBCH (P52022) DNA polymerase III alpha subunit (EC 2.7.7.7)
Length = 1159
Score = 30.0 bits (66), Expect = 7.1
Identities = 30/123 (24%), Positives = 50/123 (40%), Gaps = 13/123 (10%)
Query: 11 LLSSLVLSLFIPIFCSFEDVVDVDLVQFHLN----LEFLEAEFYFYGTL---GWGLDHAD 63
++S ++ F PI+C E + + QF N ++ +F TL W L +
Sbjct: 516 VISPTAITDFAPIYCDAEG--NFPVTQFDKNDVETAGLVKFDFLGLRTLTIIDWALGLVN 573
Query: 64 PKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPLLNISKEAFA 123
P+L + G PPV + LD D S Q A E +G + + + F
Sbjct: 574 PRLKKAGKPPVRIEAIPLD----DARSFRNLQDAKTTAVFQLESRGMKELIKRLQPDCFE 629
Query: 124 EVI 126
++I
Sbjct: 630 DII 632
>SYFB_PYRFU (Q8U260) Phenylalanyl-tRNA synthetase beta chain (EC
6.1.1.20) (Phenylalanine--tRNA ligase beta chain)
(PheRS)
Length = 556
Score = 29.6 bits (65), Expect = 9.2
Identities = 42/195 (21%), Positives = 78/195 (39%), Gaps = 27/195 (13%)
Query: 55 LGWGLDHADPKLAQGGPPPVGGQYAYLDPILRDIFSQFAFQQAGHLRAITEEVKGFPRPL 114
+ +G ++ DP+ + G + + +RD+ F Q+ ++EV+ F +
Sbjct: 345 IAYGYNNIDPEEPKLAVQGRGDPFKDFEDAIRDLMVGFGLQEVMTFNLTSKEVQ-FDK-- 401
Query: 115 LNISKEAFAEVIN------SAFGKPLNPPFDPYANSLNYLLAAYMLPYVGLTGYVGTNPK 168
+NI +E E+ N SA K L P + ++ + + VGL + + +
Sbjct: 402 MNIPEEEIVEIANPISSRWSALRKWLLPSLMEFLSNNTHEEYPQRIFEVGLATLIDESRE 461
Query: 169 LQGVRTRNLVSGLLG-------AKSGQDAMIRTLLYERRYLQVKPYQWTVYEVTHRLSLL 221
+ V L L G AK D+++R L E + + E H S +
Sbjct: 462 TKTVSEPKLAVALAGSGYTFTNAKEILDSLMRHLGIE----------YDIEETVHG-SFI 510
Query: 222 RNDLGKKQAEGKVVG 236
+GK +GK +G
Sbjct: 511 PGRVGKILVDGKEIG 525
>EAA3_HUMAN (P43005) Excitatory amino acid transporter 3
(Sodium-dependent glutamate/aspartate transporter 3)
(Excitatory amino-acid carrier 1) (Neuronal and
epithelial glutamate transporter)
Length = 524
Score = 29.6 bits (65), Expect = 9.2
Identities = 31/104 (29%), Positives = 50/104 (47%), Gaps = 12/104 (11%)
Query: 69 GGPPPVGGQYAYLDPILRDIFS----QFAFQQAGHLRAITEEVKGFPRPLLNISKEAFAE 124
G P V A LD ++R++F Q FQQ R EEVK P +N+++E+F
Sbjct: 131 GSTPEVSTVDAMLD-LIRNMFPENLVQACFQQYKTKR---EEVKPPSDPEMNMTEESFTA 186
Query: 125 VINSAFGKPLNPPF---DPYANSLNYL-LAAYMLPYVGLTGYVG 164
V+ +A K + Y++ +N L L + L + + G +G
Sbjct: 187 VMTTAISKNKTKEYKIVGMYSDGINVLGLIVFCLVFGLVIGKMG 230
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,093,220
Number of Sequences: 164201
Number of extensions: 1757114
Number of successful extensions: 3412
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 3407
Number of HSP's gapped (non-prelim): 10
length of query: 293
length of database: 59,974,054
effective HSP length: 109
effective length of query: 184
effective length of database: 42,076,145
effective search space: 7742010680
effective search space used: 7742010680
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0333b.7


