

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0328.16
(738 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
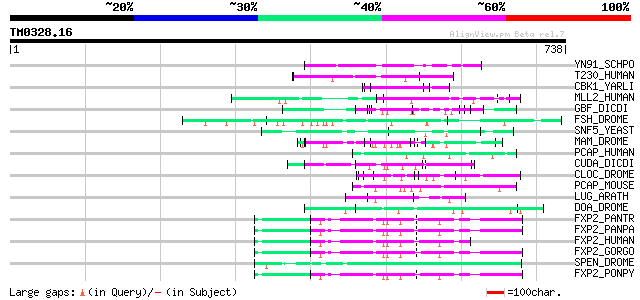
Score E
Sequences producing significant alignments: (bits) Value
YN91_SCHPO (Q9Y808) Hypothetical protein C146.01 in chromosome II 87 2e-16
T230_HUMAN (Q93074) Thyroid hormone receptor-associated protein ... 86 5e-16
CBK1_YARLI (Q6CFS5) Serine/threonine-protein kinase CBK1 (EC 2.7... 84 2e-15
MLL2_HUMAN (O14686) Myeloid/lymphoid or mixed-lineage leukemia p... 82 5e-15
GBF_DICDI (P36417) G-box binding factor (GBF) 82 5e-15
FSH_DROME (P13709) Female sterile homeotic protein (Fragile-chor... 82 7e-15
SNF5_YEAST (P18480) Transcription regulatory protein SNF5 (SWI/S... 80 2e-14
MAM_DROME (P21519) Neurogenic protein mastermind 80 2e-14
PCAP_HUMAN (Q96RN5) Positive cofactor 2 glutamine/Q-rich-associa... 79 3e-14
CUDA_DICDI (O00841) Putative transcriptional regulator cudA 79 3e-14
CLOC_DROME (O61735) Circadian locomoter output cycles Kaput prot... 78 1e-13
PCAP_MOUSE (Q924H2) Positive cofactor 2 glutamine/Q-rich-associa... 77 2e-13
LUG_ARATH (Q9FUY2) Transcriptional corepressor LEUNIG 77 2e-13
DOA_DROME (P49762) Serine/threonine-protein kinase Doa (EC 2.7.1... 77 2e-13
FXP2_PANTR (Q8MJA0) Forkhead box protein P2 76 4e-13
FXP2_PANPA (Q8HZ00) Forkhead box protein P2 76 4e-13
FXP2_HUMAN (O15409) Forkhead box protein P2 (CAG repeat protein ... 75 5e-13
FXP2_GORGO (Q8MJ99) Forkhead box protein P2 75 6e-13
SPEN_DROME (Q8SX83) Split ends protein 75 8e-13
FXP2_PONPY (Q8MJ98) Forkhead box protein P2 74 1e-12
>YN91_SCHPO (Q9Y808) Hypothetical protein C146.01 in chromosome II
Length = 1063
Score = 87.0 bits (214), Expect = 2e-16
Identities = 78/239 (32%), Positives = 98/239 (40%), Gaps = 36/239 (15%)
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQV- 451
Q Q QQQ Q Q P Q QQ + P+ ++ S + R P + Q+Q+
Sbjct: 151 QSQIQQQRQQQQQQSQQPQQTQQPQASSPTAPNTEANQQRSG--SVPGRIVPALTQQQLN 208
Query: 452 -LIQSGTTRVPTHSNPVDP--KLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFD 508
L T + + NP P KL P I + Q+ Q +F QH+ QQQ
Sbjct: 209 NLCNQITALLARNGNPPIPMQKLQSMPPARLISIYQNQIQ------KFRSLQHMQQQQQQ 262
Query: 509 QQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTH 568
QQQ QQQQ QQQQ QQQQ QQQQQQQQ+Q P NA +P+ Q
Sbjct: 263 QQQQQQQQQQQQQQQQQQQQQQQQQQQKQA-----------PQNAF------FPNPQGNV 305
Query: 569 PQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRNAPLPK 627
+ P Q S QQ + P P NP N M N P+PK
Sbjct: 306 GAQSLQSMSPQDQP-----STQQQQPQRTAAP--PNNPNVNATNNNRINIEMLNIPVPK 357
Score = 43.9 bits (102), Expect = 0.002
Identities = 55/214 (25%), Positives = 75/214 (34%), Gaps = 27/214 (12%)
Query: 469 PKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQ 528
P ++P+ ++Q + QQF + L QQ Q Q QQ+Q QQQQ Q
Sbjct: 107 PNRLANNPRVMPRLQNQNVPMQAGMQQFARNMKLTPQQRQFLLQQSQIQQQRQQQQQQSQ 166
Query: 529 QQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQV--YYVPA-RQAQAY 585
Q QQ QQ Q A P P+ + + V VPA Q Q
Sbjct: 167 QPQQTQQPQ------------------ASSPTAPNTEANQQRSGSVPGRIVPALTQQQLN 208
Query: 586 NLSMQ-QANMGESGKPQTPPNPTTLVPQT---AAYNNPMRNAPLPKTEMTAAGYRTATGG 641
NL Q A + +G P P +P + Y N ++ + +
Sbjct: 209 NLCNQITALLARNGNPPIPMQKLQSMPPARLISIYQNQIQK--FRSLQHMQQQQQQQQQQ 266
Query: 642 SPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPN 675
Q Q Q Q QQQ Q P + PN
Sbjct: 267 QQQQQQQQQQQQQQQQQQQQQQQKQAPQNAFFPN 300
Score = 41.6 bits (96), Expect = 0.008
Identities = 79/338 (23%), Positives = 119/338 (34%), Gaps = 65/338 (19%)
Query: 434 SLANAMSRQKPVIYQEQVLIQSGTTRVPTH--SNP-VDPKLNVSDPQSRIQVQQHHEQGY 490
+L +Q P +Y + + T P +NP V P+L + + +QQ
Sbjct: 80 TLQATQMKQMPSVYNNAGNVGALPTAGPNRLANNPRVMPRLQNQNVPMQAGMQQFARNMK 139
Query: 491 LL--QQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQ-----QQQQQQQQQQYIHG- 542
L Q+QF QQ +QQQ +QQ QQQ Q QQ QQ Q + QQ+ + G
Sbjct: 140 LTPQQRQFLLQQSQIQQQ--RQQQQQQSQQPQQTQQPQASSPTAPNTEANQQRSGSVPGR 197
Query: 543 -----TQFIHHNPANAIPA----------------------YYPVYPSQQQTHPQHAQVY 575
TQ +N N I A +Y +Q Q +
Sbjct: 198 IVPALTQQQLNNLCNQITALLARNGNPPIPMQKLQSMPPARLISIYQNQIQKFRSLQHMQ 257
Query: 576 YVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGY 635
+Q Q QQ + + Q PQ A + NP N G
Sbjct: 258 QQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQKQAPQNAFFPNPQGN----------VGA 307
Query: 636 RTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPNSAPPANYA-YDYVDPAHAQI 694
++ SPQ +Q QQQ P ++ AP + P N + ++ I
Sbjct: 308 QSLQSMSPQ------DQPSTQQQ--------QPQRTAAPPNNPNVNATNNNRINIEMLNI 353
Query: 695 YYSQPLAPTMPSQYQTMTAAAMVLPEGSSAQLPSDGMK 732
+ L +P+ + VL G S +LP + +K
Sbjct: 354 PVPKQLFDQIPNLPPNVKIWRDVLELGQSQRLPPEQLK 391
>T230_HUMAN (Q93074) Thyroid hormone receptor-associated protein
complex 230 kDa component (Trap230) (Activator-recruited
cofactor 240 KDa component) (ARC240) (CAG repeat protein
45) (OPA-containing protein) (Trinucleotide repeat
containing 11)
Length = 2212
Score = 85.5 bits (210), Expect = 5e-16
Identities = 71/218 (32%), Positives = 91/218 (41%), Gaps = 10/218 (4%)
Query: 378 HGVPVGYRKPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPD---SVSSDSS 434
H P G PP+ + Q T QQR SG +P ++S
Sbjct: 1991 HTGPAGTMVPPSYSSQPYQSTHPSTNPTLVDPTRHLQQRPSGYVHQQAPTYGHGLTSTQR 2050
Query: 435 LANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQ 494
++ +Q P+I + G + S + P+ Q + Q QQ +Q QQ
Sbjct: 2051 FSHQTLQQTPMISTMTPMSAQGV-QAGVRSTAILPEQQQQQQQQQQQQQQQQQQ----QQ 2105
Query: 495 QFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAI 554
Q QQQ+ ++QQ QQ +QQQ QQQQ QQQQ QQQQQQQQQQQ Q P
Sbjct: 2106 QQQQQQYHIRQQQQQQILRQQQQQQQQQQQQQQQQQQQQQQQQQQHQQQQQQQAAPPQPQ 2165
Query: 555 PAYYPVYPSQ--QQTHPQHAQVYYVPARQAQAYNLSMQ 590
P P + Q QQT Q V Q Q N Q
Sbjct: 2166 PQSQPQFQRQGLQQTQQQQQTAALVRQLQQQLSNTQPQ 2203
Score = 54.7 bits (130), Expect = 9e-07
Identities = 57/172 (33%), Positives = 63/172 (36%), Gaps = 62/172 (36%)
Query: 376 SDHGVPVGYRKPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSL 435
S GV G R QQQ QQQ Q Q Q Q QQ
Sbjct: 2069 SAQGVQAGVRSTAILPEQQQQQQQQQQQQQQQQQQQQQQQ-------------------- 2108
Query: 436 ANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQ 495
+Q+ I Q+Q Q +I QQ QQQ
Sbjct: 2109 -----QQQYHIRQQQ--------------------------QQQILRQQ--------QQQ 2129
Query: 496 FDQQQHLLQQQFDQQQHQQQQHQQQQHQQ---QQHQQQQQQQQQQQYIHGTQ 544
QQQ QQQ QQQ QQQQHQQQQ QQ Q Q Q Q Q Q+Q + TQ
Sbjct: 2130 QQQQQQQQQQQQQQQQQQQQQHQQQQQQQAAPPQPQPQSQPQFQRQGLQQTQ 2181
>CBK1_YARLI (Q6CFS5) Serine/threonine-protein kinase CBK1 (EC
2.7.1.37)
Length = 588
Score = 83.6 bits (205), Expect = 2e-15
Identities = 51/115 (44%), Positives = 60/115 (51%), Gaps = 23/115 (20%)
Query: 472 NVSDPQSRIQVQQHHEQGYLL--QQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQ 529
N+ P S + Q ++ L QQQF+QQQ+ QQQ+ QQQH Q Q QQQQHQQQQ QQ
Sbjct: 21 NLMPPPSGLYQQNSNQSNTSLDQQQQFNQQQYNSQQQYQQQQHSQYQQQQQQHQQQQQQQ 80
Query: 530 QQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQA 584
QQQQQQQQQ QQQ H QH Q P +Q+ A
Sbjct: 81 QQQQQQQQQQ---------------------QQQQQQQHQQHQQQQQQPQQQSVA 114
Score = 72.0 bits (175), Expect = 5e-12
Identities = 42/79 (53%), Positives = 47/79 (59%), Gaps = 1/79 (1%)
Query: 481 QVQQHHEQGYLLQQQFDQQQHL-LQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQY 539
Q QQ ++Q Y QQQ+ QQQH QQQ Q Q QQQQ QQQQ QQQQ QQQQQQQ QQ
Sbjct: 43 QQQQFNQQQYNSQQQYQQQQHSQYQQQQQQHQQQQQQQQQQQQQQQQQQQQQQQQHQQHQ 102
Query: 540 IHGTQFIHHNPANAIPAYY 558
Q + A P+ Y
Sbjct: 103 QQQQQPQQQSVAQQQPSDY 121
Score = 70.1 bits (170), Expect = 2e-11
Identities = 38/81 (46%), Positives = 44/81 (53%)
Query: 470 KLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQ 529
+ N S+ Q QQ Y QQQ QQQ QQQ QQQ QQQQ QQQQHQQ Q QQ
Sbjct: 46 QFNQQQYNSQQQYQQQQHSQYQQQQQQHQQQQQQQQQQQQQQQQQQQQQQQQHQQHQQQQ 105
Query: 530 QQQQQQQQQYIHGTQFIHHNP 550
QQ QQQ + +++ P
Sbjct: 106 QQPQQQSVAQQQPSDYVNFTP 126
Score = 43.5 bits (101), Expect = 0.002
Identities = 33/116 (28%), Positives = 42/116 (35%), Gaps = 21/116 (18%)
Query: 503 LQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYP 562
L QQ Q + QQQ +QQQ + QQQ QQQQ + Y
Sbjct: 29 LYQQNSNQSNTSLDQQQQFNQQQYNSQQQYQQQQ---------------------HSQYQ 67
Query: 563 SQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNN 618
QQQ H Q Q +Q Q QQ + + Q P + Q + Y N
Sbjct: 68 QQQQQHQQQQQQQQQQQQQQQQQQQQQQQQHQQHQQQQQQPQQQSVAQQQPSDYVN 123
Score = 35.8 bits (81), Expect = 0.42
Identities = 26/94 (27%), Positives = 37/94 (38%)
Query: 423 LPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQV 482
+P P + +S + S + + +Q + HS + Q + Q
Sbjct: 23 MPPPSGLYQQNSNQSNTSLDQQQQFNQQQYNSQQQYQQQQHSQYQQQQQQHQQQQQQQQQ 82
Query: 483 QQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQ 516
QQ +Q QQQ QQH QQQ QQQ QQ
Sbjct: 83 QQQQQQQQQQQQQQQHQQHQQQQQQPQQQSVAQQ 116
>MLL2_HUMAN (O14686) Myeloid/lymphoid or mixed-lineage leukemia
protein 2 (ALL1-related protein)
Length = 5262
Score = 82.0 bits (201), Expect = 5e-15
Identities = 99/372 (26%), Positives = 139/372 (36%), Gaps = 60/372 (16%)
Query: 295 GGVRVQQQDQKVLGIEEQFAQLGVAAGVGVGQKQDEGFVVITSPPPPPVPTTLSAPAMGV 354
G V +QQQ Q+ G++ A G+ V SP PP P + PA
Sbjct: 3474 GQVAIQQQQQQGPGVQTNQALGPKPQGLMPPSSHQGLLVQQLSPQPPQGPQGMLGPAQVA 3533
Query: 355 PIN-----SAVVSGDY------QNRVFSDDERSDHGVPV-GYRKPPTPTPQQQAQQQLHT 402
+ + G + Q+RV S + + G + G+R T QQQ QQQ H
Sbjct: 3534 VLQQQHPGALGPQGPHRQVLMTQSRVLSSPQLAQQGQGLMGHR---LVTAQQQQQQQQHQ 3590
Query: 403 QTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSGTTRVPT 462
Q S + + QQ ++ SG ++
Sbjct: 3591 QQGSMAGLSHLQQ----------------------------------SLMSHSGQPKLSA 3616
Query: 463 HSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQH 522
P ++ Q + Q+QQ + QQQ QQQ L QQQ QQQ QQQ QQQQ
Sbjct: 3617 Q-----PMGSLQQLQQQQQLQQQQQLQQQQQQQLQQQQQLQQQQLQQQQQQQQLQQQQQQ 3671
Query: 523 QQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQA 582
Q QQ QQQ QQQQQQQ Q + + QQQ Q A +PA+
Sbjct: 3672 QLQQQQQQLQQQQQQQQQQFQQQQQQQQMGLLNQSRTLLSPQQQQQQQVALGPGMPAKPL 3731
Query: 583 QAYNL-SMQQANMGESGKPQTPPNPTTLVPQTAA----YNNPMRNAPLPKTEMTAAG-YR 636
Q ++ + +GK Q +P T P+ P++ T G +
Sbjct: 3732 QHFSSPGALGPTLLLTGKEQNTVDPAVSSEATEGPSTHQGGPLAIGTTPESMATEPGEVK 3791
Query: 637 TATGGSPQLVQV 648
+ G QL+ V
Sbjct: 3792 PSLSGDSQLLLV 3803
Score = 50.4 bits (119), Expect = 2e-05
Identities = 61/201 (30%), Positives = 83/201 (40%), Gaps = 25/201 (12%)
Query: 489 GYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQ--------QQQQQQ-- 538
G L + LLQ++ Q Q Q+ Q Q+ QQQQ QQQQQ QQQQQQ
Sbjct: 3427 GNLALRSLGPDSRLLQERQLQLQQQRMQLAQKLQQQQQQQQQQQHLLGQVAIQQQQQQGP 3486
Query: 539 YIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESG 598
+ Q + P +P QQ PQ Q AQ +QQ + G G
Sbjct: 3487 GVQTNQALGPKPQGLMPPSSHQGLLVQQLSPQPPQGPQGMLGPAQV--AVLQQQHPGALG 3544
Query: 599 KPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQ 658
PQ P+ L+ Q+ ++P L + G+R T Q Q QHQQQ
Sbjct: 3545 -PQ-GPHRQVLMTQSRVLSSPQ----LAQQGQGLMGHRLVTAQQQQ------QQQQHQQQ 3592
Query: 659 YVTYSQIHHPSQSIAPNSAPP 679
+ + + H QS+ +S P
Sbjct: 3593 -GSMAGLSHLQQSLMSHSGQP 3612
Score = 47.8 bits (112), Expect = 1e-04
Identities = 45/167 (26%), Positives = 67/167 (39%), Gaps = 3/167 (1%)
Query: 498 QQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAY 557
+QQ +Q+Q DQ + QQ++H + + QQQQQQQQQQQ + + +P+ + P
Sbjct: 3294 EQQSKIQKQLDQVRKQQKEHTNLMAEYRNKQQQQQQQQQQQQQQHSAVLALSPSQS-PRL 3352
Query: 558 YPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYN 617
P Q P H QA L + M G+P P T L Q +
Sbjct: 3353 LTKLPG--QLLPGHGLQPPQGPPGGQAGGLRLTPGGMALPGQPGGPFLNTALAQQQQQQH 3410
Query: 618 NPMRNAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQ 664
+ + + G P + QLQ QQQ + +Q
Sbjct: 3411 SGGAGSLAGPSGGFFPGNLALRSLGPDSRLLQERQLQLQQQRMQLAQ 3457
Score = 37.4 bits (85), Expect = 0.14
Identities = 23/55 (41%), Positives = 29/55 (51%), Gaps = 8/55 (14%)
Query: 473 VSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQH 527
V++ QS+IQ Q L Q QQ+ + + QQQQ QQQQ QQQQH
Sbjct: 3292 VTEQQSKIQKQ--------LDQVRKQQKEHTNLMAEYRNKQQQQQQQQQQQQQQH 3338
Score = 33.1 bits (74), Expect = 2.7
Identities = 38/165 (23%), Positives = 60/165 (36%), Gaps = 36/165 (21%)
Query: 502 LLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHG-----TQFIHHN------- 549
L++ + ++ + Q+ QQ Q Q QQQQQQQQQ + + H
Sbjct: 2973 LIEDLLEHEKKELQKKQQLSAQLQPAQQQQQQQQQHSLLPAPGPAQAMSLPHEGSSPSLA 3032
Query: 550 ------------------PANAIPAYYPVYPSQQQTHPQHAQV----YYVPARQAQAYNL 587
P +P P + QQ+ P A V + + + L
Sbjct: 3033 GSQQQLSLGLAVARQPGLPQPLMPTQPPAHALQQRLAPSMAMVSNQGHMLSGQHGGQAGL 3092
Query: 588 SMQQANMGE-SGKPQ-TPPNPTTLVPQTAAYNNPMRNAPLPKTEM 630
QQ++ S KP T P + PQ A + N+ P T++
Sbjct: 3093 VPQQSSQPVLSQKPMGTMPPSMCMKPQQLAMQQQLANSFFPDTDL 3137
Score = 32.3 bits (72), Expect = 4.6
Identities = 22/58 (37%), Positives = 29/58 (49%), Gaps = 5/58 (8%)
Query: 3 APPQPPT-PSAAPPLSSVAPPPQPPPHLNYADSVDSS---PRSRNTDSWDDPFPPTSK 56
+P PPT P P +A PP P P L A S +S+ PR+R + +D PP K
Sbjct: 4649 SPASPPTEPLVELPTEPLAEPPVPSP-LPLASSPESARPKPRARPPEEGEDTRPPRLK 4705
Score = 31.6 bits (70), Expect = 7.9
Identities = 37/147 (25%), Positives = 50/147 (33%), Gaps = 17/147 (11%)
Query: 390 PTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQE 449
P QQQ QQQ H+ + PAQ G+ PS SL A++RQ
Sbjct: 2997 PAQQQQQQQQQHSLLPAPG-PAQAMSLPHEGSS-PSLAGSQQQLSLGLAVARQ------- 3047
Query: 450 QVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQ 509
+P P P + + + QG++L Q Q L+ QQ Q
Sbjct: 3048 --------PGLPQPLMPTQPPAHALQQRLAPSMAMVSNQGHMLSGQHGGQAGLVPQQSSQ 3099
Query: 510 QQHQQQQHQQQQHQQQQHQQQQQQQQQ 536
Q+ QQ QQQ
Sbjct: 3100 PVLSQKPMGTMPPSMCMKPQQLAMQQQ 3126
>GBF_DICDI (P36417) G-box binding factor (GBF)
Length = 708
Score = 82.0 bits (201), Expect = 5e-15
Identities = 59/162 (36%), Positives = 72/162 (44%), Gaps = 26/162 (16%)
Query: 479 RIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQ--- 535
++Q QQHH+Q + Q Q QH QQQ QQQHQQQ HQQQQ QQQQH QQQQ Q
Sbjct: 171 QMQQQQHHQQ--MQHHQLQQHQHQHQQQQQQQQHQQQHHQQQQQQQQQHHQQQQHHQHSQ 228
Query: 536 -QQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVP------ARQAQAYNLS 588
QQQ+ H Q H + +QQQ Q Q+ VP + N +
Sbjct: 229 PQQQHQHNQQQQHQH-------------NQQQHQQQQNQIQMVPQQPQSLSNSGNNNNNN 275
Query: 589 MQQANMGESGKPQTPPNPTTLVPQTAAYNNPM-RNAPLPKTE 629
N + N L T + NN N P P T+
Sbjct: 276 NNNNNSNNNNNNNNNNNSHQLNNLTLSQNNTSGSNTPSPSTK 317
Score = 78.6 bits (192), Expect = 6e-14
Identities = 51/127 (40%), Positives = 60/127 (47%), Gaps = 16/127 (12%)
Query: 479 RIQVQQHHEQGYLLQQ---QFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQ 535
++Q QQHH+Q QQ Q QQQH Q Q Q Q Q QHQQQQ QQQ QQ QQQQ
Sbjct: 152 QMQQQQHHQQMQQQQQHHQQMQQQQHHQQMQHHQLQQHQHQHQQQQQQQQHQQQHHQQQQ 211
Query: 536 QQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYN----LSMQQ 591
QQQ H Q HH + P QQ H Q Q + + Q N + Q
Sbjct: 212 QQQQQHHQQQQHHQHSQ---------PQQQHQHNQQQQHQHNQQQHQQQQNQIQMVPQQP 262
Query: 592 ANMGESG 598
++ SG
Sbjct: 263 QSLSNSG 269
Score = 78.2 bits (191), Expect = 7e-14
Identities = 55/136 (40%), Positives = 65/136 (47%), Gaps = 16/136 (11%)
Query: 477 QSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQ 536
Q + Q QHH+Q +QQQ QQ QQQ QQ QQQ HQQ QH Q Q Q Q QQQQ
Sbjct: 141 QQQQQQPQHHQQ---MQQQQHHQQMQQQQQHHQQMQQQQHHQQMQHHQLQQHQHQHQQQQ 197
Query: 537 QQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGE 596
QQ H Q HH + QQQ H QH+Q P +Q Q + QQ +
Sbjct: 198 QQQQHQQQ--HHQQQQQQQQQH----HQQQQHHQHSQ----PQQQHQH---NQQQQHQHN 244
Query: 597 SGKPQTPPNPTTLVPQ 612
+ Q N +VPQ
Sbjct: 245 QQQHQQQQNQIQMVPQ 260
Score = 75.1 bits (183), Expect = 6e-13
Identities = 52/172 (30%), Positives = 68/172 (39%), Gaps = 28/172 (16%)
Query: 364 DYQNRVFSDDERSDHGVPVGYRKPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDL 423
+YQ ++ +++ P + QQQ Q Q H Q Q Q H Q QQ+
Sbjct: 113 NYQYQLMFMQQQAQQNQPPQQNQQQQHHQQQQQQPQHHQQMQQQQHHQQMQQQQQ----- 167
Query: 424 PSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQ 483
+ +Q+ Q ++ H + Q + Q
Sbjct: 168 -----------------------HHQQMQQQQHHQQMQHHQLQQHQHQHQQQQQQQQHQQ 204
Query: 484 QHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQ 535
QHH+Q QQQ QQQ Q QQQHQ Q QQ QH QQQHQQQQ Q Q
Sbjct: 205 QHHQQQQQQQQQHHQQQQHHQHSQPQQQHQHNQQQQHQHNQQQHQQQQNQIQ 256
Score = 69.7 bits (169), Expect = 3e-11
Identities = 54/157 (34%), Positives = 65/157 (41%), Gaps = 18/157 (11%)
Query: 461 PTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQ--------HLLQQQFDQQQH 512
P+H N+ D Q +I Q+ Y Q F QQQ QQQ QQQ
Sbjct: 86 PSHDGQYPDMPNMVD-QYQIHPNQNPHYNYQYQLMFMQQQAQQNQPPQQNQQQQHHQQQQ 144
Query: 513 QQQQHQQQQHQQQQHQQQQQQ----QQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTH 568
QQ QH QQ QQQ HQQ QQQ QQ QQ H Q HH + QQQ
Sbjct: 145 QQPQHHQQMQQQQHHQQMQQQQQHHQQMQQQQHHQQMQHHQLQQ-----HQHQHQQQQQQ 199
Query: 569 PQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPN 605
QH Q ++ +Q Q + QQ + + Q N
Sbjct: 200 QQHQQQHHQQQQQQQQQHHQQQQHHQHSQPQQQHQHN 236
Score = 67.4 bits (163), Expect = 1e-10
Identities = 58/194 (29%), Positives = 71/194 (35%), Gaps = 50/194 (25%)
Query: 482 VQQHHEQGYLLQQQFDQQQHLLQQQFDQ--QQHQQQQHQQQQHQQQQHQQQQQQQQQQQY 539
+QQ +Q QQ QQ H QQQ Q QQ QQQQH QQ QQQQH QQ QQQQ Q
Sbjct: 121 MQQQAQQNQPPQQNQQQQHHQQQQQQPQHHQQMQQQQHHQQMQQQQQHHQQMQQQQHHQQ 180
Query: 540 IHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGK 599
+ Q H + QQQ QH Q ++ +Q Q + QQ + +
Sbjct: 181 MQHHQLQQHQHQH----------QQQQQQQQHQQQHHQQQQQQQQQHHQQQQHHQHSQPQ 230
Query: 600 PQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQY 659
Q N Q Q NQ QHQQQ
Sbjct: 231 QQHQHN--------------------------------------QQQQHQHNQQQHQQQQ 252
Query: 660 VTYSQIHHPSQSIA 673
+ QS++
Sbjct: 253 NQIQMVPQQPQSLS 266
Score = 32.7 bits (73), Expect = 3.5
Identities = 26/84 (30%), Positives = 37/84 (43%), Gaps = 18/84 (21%)
Query: 470 KLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQ----------------QFDQQQHQ 513
K+ Q+++Q Q+H Q ++ Q +D QQ+ Q Q QQQHQ
Sbjct: 372 KMQQVPQQAQLQPLQNHNQ--IIPQLWDSQQNNSSQNTPPTQPQNNMNQINHQLLQQQHQ 429
Query: 514 QQQHQQQQHQQQQHQQQQQQQQQQ 537
Q Q Q + +QQ Q QQQ
Sbjct: 430 QAQLQAHLNLTASNQQVPPQLQQQ 453
>FSH_DROME (P13709) Female sterile homeotic protein (Fragile-chorion
membrane protein)
Length = 2038
Score = 81.6 bits (200), Expect = 7e-15
Identities = 111/403 (27%), Positives = 163/403 (39%), Gaps = 58/403 (14%)
Query: 231 GVEAGGGGASLPGSFANCKNSKQDVSSV----PDSPMLETSSSFGSTSSSPALANLPPIR 286
GV G S + KN D+S V P + L S G T++ + IR
Sbjct: 1210 GVPVAGAAVSASTGQQHNKNGPNDLSKVQPGGPINAALPPHSFAGGTATVATSQSSGGIR 1269
Query: 287 T----HAEDGGGGGVRVQQQDQKVLGIEEQFAQLGVAAGVGV---GQKQDEGFVVITSPP 339
H G GGG + + G A G G+ G + + S
Sbjct: 1270 IASNLHKPSGLGGGDLGEHHAALAAALTSGINSTGTAGG-GINNNGGSNNNANPLGGSHG 1328
Query: 340 PPPVPTTLSAPAMGV--------PINSAVVSGDYQNRVFSDDERSDHGVPVGY-RKPPTP 390
V +L++ A G+ P+ ++ S ++ +D+ + + +P P
Sbjct: 1329 DAMVNASLASLASGLKQIPQFDDPVEQSLASLEFSAGSTGKSGLTDNFLMQQHLMQPAGP 1388
Query: 391 TPQQQAQQQL---HTQTQSQSHPAQFQQR-----------SSGGTDLPSPDS---VSSDS 433
QQQ QQQ H Q Q Q Q QQ+ S G ++ + ++ +
Sbjct: 1389 QQQQQQQQQQPFGHQQQQQQQQQQQQQQQQQHMDYVTELLSKGAENVGGMNGNHLLNFNL 1448
Query: 434 SLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQ 493
+A A ++ P Q+Q + V LN++ S + ++ +Q + Q
Sbjct: 1449 DMAAAYQQKHP--QQQQQQAHNNGFNVADFGMAGFDGLNMT-AASFLDLEPSLQQQQMQQ 1505
Query: 494 QQFDQQQHLLQQQ---FDQQQHQQQQHQQQQHQ--QQQHQQQQQQQQQQQYIHGTQF--I 546
Q QQ H QQQ QQQHQQQ HQQQQ Q QQQ QQQQQQQQQQQ++ Q
Sbjct: 1506 MQLQQQHHQQQQQQTHQQQQQHQQQHHQQQQQQLTQQQLQQQQQQQQQQQHLQQQQHQQQ 1565
Query: 547 HHNPAN-------AIPAYYPVYPSQQQTHPQHAQVYYVPARQA 582
HH AN I + P P +QQ QH +V +P +Q+
Sbjct: 1566 HHQAANKLLIIPKPIESMMPSPPDKQQLQ-QHQKV--LPPQQS 1605
Score = 68.9 bits (167), Expect = 4e-11
Identities = 103/441 (23%), Positives = 150/441 (33%), Gaps = 95/441 (21%)
Query: 342 PVPTTLSAPAMGVPINSAVVS---GDYQNRVFSDD-ERSDHGVPVGYRKPP--------- 388
P P + GVP+ A VS G N+ +D + G P+ PP
Sbjct: 1199 PAPGNMMHAGAGVPVAGAAVSASTGQQHNKNGPNDLSKVQPGGPINAALPPHSFAGGTAT 1258
Query: 389 TPTPQQQAQQQLHTQTQSQS----------HPAQFQQRSSG----GTDLPSPDSVSSDSS 434
T Q ++ + S H A +SG GT ++ ++
Sbjct: 1259 VATSQSSGGIRIASNLHKPSGLGGGDLGEHHAALAAALTSGINSTGTAGGGINNNGGSNN 1318
Query: 435 LANAM--SRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLL 492
AN + S ++ + SG ++P +PV+ L + ++ L
Sbjct: 1319 NANPLGGSHGDAMVNASLASLASGLKQIPQFDDPVEQSL------ASLEFSAGSTGKSGL 1372
Query: 493 QQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQ--YIHGTQFIHHNP 550
F QQHL+Q QQQ QQQQ Q HQQQQ QQQQQQQQQQQ + T+ +
Sbjct: 1373 TDNFLMQQHLMQPAGPQQQQQQQQQQPFGHQQQQQQQQQQQQQQQQQHMDYVTELLSKGA 1432
Query: 551 ANA--------IPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQT 602
N + + + QQ HPQ +Q QA+N A+ G +G
Sbjct: 1433 ENVGGMNGNHLLNFNLDMAAAYQQKHPQQ--------QQQQAHNNGFNVADFGMAG---- 1480
Query: 603 PPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGG--SPQLVQVPANQLQHQ---- 656
MTAA + Q+ Q+ Q HQ
Sbjct: 1481 ----------------------FDGLNMTAASFLDLEPSLQQQQMQQMQLQQQHHQQQQQ 1518
Query: 657 ---QQYVTYSQIHHPSQSIAPNSAPPANYAYDYVDPAHAQIYYSQPLAPTMPSQYQTMTA 713
QQ + Q HH Q H Q Q Q+
Sbjct: 1519 QTHQQQQQHQQQHHQQQQQQLTQQQLQQQQQQQQQQQHLQQQQHQ-------QQHHQAAN 1571
Query: 714 AAMVLPEGSSAQLPSDGMKQQ 734
+++P+ + +PS KQQ
Sbjct: 1572 KLLIIPKPIESMMPSPPDKQQ 1592
Score = 32.7 bits (73), Expect = 3.5
Identities = 34/118 (28%), Positives = 51/118 (42%), Gaps = 18/118 (15%)
Query: 193 GAGLLNRGFSDSASVNCLLGLDDV-VAGNNNNIEQGGREGVEAGGGGAS-----LPGSFA 246
GAG + SA+V G V+ ++ +Q G +G GGG+S PGS A
Sbjct: 171 GAGASTGSGTSSAAVTSGPGSGSTKVSVAASSAQQSGLQGATGAGGGSSSTPGTQPGSGA 230
Query: 247 NCKNSKQDVSSVPDSPMLETSSSFGSTSSSPALANLPP--------IRTHAEDGGGGG 296
+ + VS++ + SS+ G S P ++ +PP T A GG GG
Sbjct: 231 GGAIAARPVSAMGGT----VSSTAGGAPSIPPISTMPPHTVPGSTNTTTTAMAGGVGG 284
>SNF5_YEAST (P18480) Transcription regulatory protein SNF5 (SWI/SNF
complex component SNF5) (Transcription factor TYE4)
Length = 905
Score = 80.1 bits (196), Expect = 2e-14
Identities = 82/293 (27%), Positives = 108/293 (35%), Gaps = 67/293 (22%)
Query: 336 TSPPPPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSDDERSDHGVPVGYRKPPTPTPQQQ 395
++PPPPP P L VP+ A ++ +P + P T QQQ
Sbjct: 97 STPPPPPAPHNLHPQIGQVPLAPAPIN-----------------LPPQIAQLPLAT-QQQ 138
Query: 396 AQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQS 455
+L Q ++++P + + A Q Q Q+L+Q
Sbjct: 139 VLNKLRQQAIAKNNPQVVNAITVAQQQVQRQIEQQKGQQTAQTQLEQ-----QRQLLVQQ 193
Query: 456 GTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQ 515
+ Q R Q+Q+ +Q + Q QQQ QQQ Q Q QQQ
Sbjct: 194 QQQQ-----------------QLRNQIQRQQQQQFRHHVQIQQQQQKQQQQQQQHQQQQQ 236
Query: 516 QHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVY 575
Q QQQQ QQQQ QQQQQQQQQQQ QQQ Q Q
Sbjct: 237 QQQQQQQQQQQQQQQQQQQQQQQQ---------------------QQQQQQQQGQIPQSQ 275
Query: 576 YVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVP--QTAAYNNPMRNAPLP 626
VP Q ++S Q + Q P P +P QT Y+ P P P
Sbjct: 276 QVP----QVRSMSGQPPTNVQPTIGQLPQLPKLNLPKYQTIQYDPPETKLPYP 324
Score = 45.4 bits (106), Expect = 5e-04
Identities = 51/184 (27%), Positives = 65/184 (34%), Gaps = 26/184 (14%)
Query: 504 QQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPA-----------N 552
QQ + QQ Q QQ+HQQ + QQQQQQQQ H +P
Sbjct: 34 QQLYQSLTPQQLQMIQQRHQQLLRSRLQQQQQQQQQTSPPPQTHQSPPPPPQQSQPIANQ 93
Query: 553 AIPAYYPVYPSQQQTHPQHAQVYYVPA------RQAQAYNLSMQQANMGESGKPQTPPNP 606
+ + P P+ HPQ QV PA + AQ + QQ + NP
Sbjct: 94 SATSTPPPPPAPHNLHPQIGQVPLAPAPINLPPQIAQLPLATQQQVLNKLRQQAIAKNNP 153
Query: 607 TTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQIH 666
+ T A R K + TA QL Q +Q QQQ +QI
Sbjct: 154 QVVNAITVAQQQVQRQIEQQKGQQTA---------QTQLEQQRQLLVQQQQQQQLRNQIQ 204
Query: 667 HPSQ 670
Q
Sbjct: 205 RQQQ 208
>MAM_DROME (P21519) Neurogenic protein mastermind
Length = 1596
Score = 80.1 bits (196), Expect = 2e-14
Identities = 92/315 (29%), Positives = 121/315 (38%), Gaps = 65/315 (20%)
Query: 383 GYRKPP---TPTPQQQAQQQLHTQTQSQSHPAQFQQRSSG-----GTDLPSPDSVSSDSS 434
G+ +PP P QQQ QQQ Q Q H Q G G P+ +
Sbjct: 640 GFPRPPHGMNPQQQQQQQQQQQQQQAQQQHGQMMGQGQPGRYNDYGGGFPNDFGLGP--- 696
Query: 435 LANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQ 494
N +Q+ Q+Q Q + P P ++ Q+ + ++Q E Y Q
Sbjct: 697 --NGPQQQQQQAQQQQPQQQHLPPQFHQQKGP-GPGAGMNVQQNFLDIKQ--ELFYSSQN 751
Query: 495 QFD----QQQHLLQQQFDQQQHQQQQHQ-----------------QQQHQQQQHQQQQQQ 533
FD QQQ +QQQ QQ HQQQQ + QQQ QQQQ QQQQQQ
Sbjct: 752 DFDLKRLQQQQAMQQQQQQQHHQQQQPKMGGVPNFNKQQQQQQVPQQQLQQQQQQQQQQQ 811
Query: 534 QQQQQYIHGTQFIHHNPANAIPAYYPVYP-------------SQQQTHPQHAQVYYVPAR 580
QQQQQY + F + NP NA + P +QQQ P P +
Sbjct: 812 QQQQQY---SPFSNQNP-NAAANFLNCPPRGGPNGNQQPGNLAQQQQQPGAG-----PQQ 862
Query: 581 QAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATG 640
Q Q N Q N +G PN Q + + + + T + + G
Sbjct: 863 QQQRGNAGNGQQNNPNTGPGGNTPNAPQQQQQQQSTTTTL------QMKQTQQLHISQQG 916
Query: 641 GSPQLVQVPANQLQH 655
G Q +QV A Q H
Sbjct: 917 GGAQGIQVSAGQHLH 931
Score = 57.0 bits (136), Expect = 2e-07
Identities = 39/77 (50%), Positives = 42/77 (53%), Gaps = 11/77 (14%)
Query: 481 QVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQH----QQQQHQQQQHQQQQQQQQQ 536
Q QQ +Q +L Q Q L QQ Q Q QQQQH QQQQ Q QQ QQQQQQQQQ
Sbjct: 21 QQQQQQQQQHLNLQLHQQHLGLHLQQQQQLQLQQQQHNAQAQQQQIQVQQQQQQQQQQQQ 80
Query: 537 QQYIHGTQFIHHNPANA 553
QQ H+P NA
Sbjct: 81 QQQ-------QHSPYNA 90
Score = 53.5 bits (127), Expect = 2e-06
Identities = 35/67 (52%), Positives = 39/67 (57%), Gaps = 6/67 (8%)
Query: 473 VSDPQSRIQVQQH-----HEQGYLLQQQFDQQQHLLQQQFDQQQHQQQ-QHQQQQHQQQQ 526
V+ Q + Q QQH H+Q L Q QQ L QQQ + Q QQQ Q QQQQ QQQQ
Sbjct: 18 VAQQQQQQQQQQHLNLQLHQQHLGLHLQQQQQLQLQQQQHNAQAQQQQIQVQQQQQQQQQ 77
Query: 527 HQQQQQQ 533
QQQQQQ
Sbjct: 78 QQQQQQQ 84
Score = 50.8 bits (120), Expect = 1e-05
Identities = 50/168 (29%), Positives = 74/168 (43%), Gaps = 23/168 (13%)
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVL 452
QQQ Q Q Q Q Q QQ S +L + ++ + A P V
Sbjct: 62 QQQQIQVQQQQQQQQQQQQQQQQHSPYNANLGATGGIAGITGGNGAGGPTNP----GAVP 117
Query: 453 IQSGTTRVPTHSNPVDPKLNVSDPQSRIQ----VQQHHEQGYLLQQQFDQQQHLLQQQFD 508
G T +PT PV +L R + V ++ + + +Q +Q+ +LQ++F
Sbjct: 118 TAPGDT-MPTKRMPVVDRLRRRMENYRRRQTDCVPRYEQAFNTVCEQQNQETTVLQKRFL 176
Query: 509 QQQHQQ-------------QQHQQQQHQ-QQQHQQQQQQQQQQQYIHG 542
+ ++++ QQHQQQQHQ QQQHQQ QQ QQ Q + G
Sbjct: 177 ESKNKRAAKKTDKKLPDPSQQHQQQQHQQQQQHQQHQQHQQAQTMLAG 224
Score = 49.3 bits (116), Expect = 4e-05
Identities = 48/165 (29%), Positives = 73/165 (44%), Gaps = 25/165 (15%)
Query: 394 QQAQQQLHTQTQSQSHPAQFQQR---SSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQ 450
Q QQQ+ Q Q Q Q QQ+ S +L + ++ + A P
Sbjct: 60 QAQQQQIQVQQQQQQQQQQQQQQQQHSPYNANLGATGGIAGITGGNGAGGPTNP----GA 115
Query: 451 VLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQ----VQQHHEQGYLLQQQFDQQQHLLQQQ 506
V G T +PT PV +L R + V ++ + + +Q +Q+ +LQ++
Sbjct: 116 VPTAPGDT-MPTKRMPVVDRLRRRMENYRRRQTDCVPRYEQAFNTVCEQQNQETTVLQKR 174
Query: 507 FDQQQHQQ-------------QQHQQQQHQQQQHQQQQQQQQQQQ 538
F + ++++ QQHQQQQHQQQQ QQ QQ QQ Q
Sbjct: 175 FLESKNKRAAKKTDKKLPDPSQQHQQQQHQQQQQHQQHQQHQQAQ 219
Score = 48.1 bits (113), Expect = 8e-05
Identities = 78/308 (25%), Positives = 99/308 (31%), Gaps = 60/308 (19%)
Query: 387 PPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPD--SVSSDSSLANAMSRQKP 444
P QQQ QQQ Q Q Q P Q ++ L P + + N +Q+
Sbjct: 796 PQQQLQQQQQQQQQQQQQQQQYSPFSNQNPNAAANFLNCPPRGGPNGNQQPGNLAQQQQQ 855
Query: 445 VIYQEQVLIQSGTTRVPTHSNP-VDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLL 503
Q Q G +NP P N P + Q QQ LQ + QQ H+
Sbjct: 856 PGAGPQQQQQRGNAGNGQQNNPNTGPGGNT--PNAPQQQQQQQSTTTTLQMKQTQQLHIS 913
Query: 504 QQQ---------------------------------FDQQQHQQQQHQQQQHQQQQHQQQ 530
QQ F QQQ QQQQ QQQ Q
Sbjct: 914 QQGGGAQGIQVSAGQHLHLSGDMKSNVSVAAQQGVFFSQQQAQQQQQQQQPGGTNGPNPQ 973
Query: 531 QQQQQQQQYIHGTQFIHHNPANAIPAYYPV-------YPSQQQTHP-QHAQVYYVPA--- 579
QQQQQ HG N + V P QQQ P Q+ VP+
Sbjct: 974 QQQQQP----HG-----GNAGGGVGVGVGVGVGNGGPNPGQQQQQPNQNMSNANVPSDGF 1024
Query: 580 --RQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRT 637
Q+Q+ N + QQ + + Q PN Q A A + A+G
Sbjct: 1025 SLSQSQSMNFNQQQQQQAAAQQQQVQPNMRQRQTQAQAAAAAAAAAAQAQAAANASGPNV 1084
Query: 638 ATGGSPQL 645
PQ+
Sbjct: 1085 PLMQQPQV 1092
Score = 41.6 bits (96), Expect = 0.008
Identities = 70/282 (24%), Positives = 91/282 (31%), Gaps = 49/282 (17%)
Query: 413 FQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLN 472
F +GGT +P++ + + N MS Q L Q H N +
Sbjct: 587 FPNGPNGGTG-GAPNAGGNGGNSGNLMSEHP---LAAQTLKQMAEQH--QHKNAMGGMGG 640
Query: 473 VSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQ--------------------- 511
P + QQ +Q QQQ QQQH Q Q Q
Sbjct: 641 FPRPPHGMNPQQQQQQQQQQQQQQAQQQH--GQMMGQGQPGRYNDYGGGFPNDFGLGPNG 698
Query: 512 -HQQQQHQQQQHQQQQHQQQQQQQQ-----------QQQYIHGTQFIHHNPAN--AIPAY 557
QQQQ QQQ QQQH Q QQ QQ ++ Q + ++ N +
Sbjct: 699 PQQQQQQAQQQQPQQQHLPPQFHQQKGPGPGAGMNVQQNFLDIKQELFYSSQNDFDLKRL 758
Query: 558 YPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYN 617
QQQ QH Q +N QQ + + Q Q Y
Sbjct: 759 QQQQAMQQQQQQQHHQQQQPKMGGVPNFNKQQQQQQVPQQQLQQQQQQQQQQQQQQQQY- 817
Query: 618 NPMRNAPLPKTEMTAAGY-RTATGGSPQLVQVPANQLQHQQQ 658
+P N + AA + G P Q P N Q QQQ
Sbjct: 818 SPFSN----QNPNAAANFLNCPPRGGPNGNQQPGNLAQQQQQ 855
Score = 41.6 bits (96), Expect = 0.008
Identities = 36/93 (38%), Positives = 39/93 (41%), Gaps = 21/93 (22%)
Query: 509 QQQHQQQQHQQQQHQQQQ---HQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQ 565
QQQ QQQQH Q QQ H QQQQQ Q QQ H Q A V QQ
Sbjct: 22 QQQQQQQQHLNLQLHQQHLGLHLQQQQQLQLQQQQHNAQ--------AQQQQIQVQQQQQ 73
Query: 566 QTHPQHAQVYYVPARQAQAYNLSMQQANMGESG 598
Q Q Q +Q YN AN+G +G
Sbjct: 74 QQQQQQQQ-----QQQHSPYN-----ANLGATG 96
Score = 38.9 bits (89), Expect = 0.049
Identities = 21/39 (53%), Positives = 24/39 (60%), Gaps = 3/39 (7%)
Query: 503 LQQQFDQQQHQQQQHQQQQHQ---QQQHQQQQQQQQQQQ 538
+QQQ +QQ QQQQ QQQQH + Q QQ QQQQ
Sbjct: 1416 MQQQLMRQQQQQQQQQQQQHMGPGAANNMQMQQLLQQQQ 1454
Score = 38.5 bits (88), Expect = 0.064
Identities = 29/81 (35%), Positives = 33/81 (39%), Gaps = 31/81 (38%)
Query: 499 QQHLLQQQFDQQQHQQQQH---------QQQQHQQQQH---------------------- 527
QQ L++QQ QQQ QQQQH Q QQ QQQ
Sbjct: 1417 QQQLMRQQQQQQQQQQQQHMGPGAANNMQMQQLLQQQQSGGGGNMMASQMQMTSMHMTQT 1476
Query: 528 QQQQQQQQQQQYIHGTQFIHH 548
QQQ QQQQQ++ T H
Sbjct: 1477 QQQITMQQQQQFVQSTTTTTH 1497
Score = 37.4 bits (85), Expect = 0.14
Identities = 20/41 (48%), Positives = 23/41 (55%), Gaps = 2/41 (4%)
Query: 477 QSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQH 517
Q ++Q+QQ QQQ QQ QQQ QQQ QQQQH
Sbjct: 47 QQQLQLQQQQHNAQAQQQQIQVQQQ--QQQQQQQQQQQQQH 85
Score = 35.4 bits (80), Expect = 0.55
Identities = 42/188 (22%), Positives = 68/188 (35%), Gaps = 25/188 (13%)
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGG----TDLPSPDSVSSDSSLANAMSRQKPVIYQ 448
QQQ QQQ Q Q Q P ++GG T ++ ++ A P +
Sbjct: 70 QQQQQQQQQQQQQQQHSPYNANLGATGGIAGITGGNGAGGPTNPGAVPTAPGDTMPT--K 127
Query: 449 EQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQF------------ 496
++ R+ + + + +Q +++ +LQ++F
Sbjct: 128 RMPVVDRLRRRMENYRRRQTDCVPRYEQAFNTVCEQQNQETTVLQKRFLESKNKRAAKKT 187
Query: 497 ------DQQQHLLQQQFDQQQHQQ-QQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHN 549
QQH QQ QQQHQQ QQHQQ Q Q QQ+ + + +
Sbjct: 188 DKKLPDPSQQHQQQQHQQQQQHQQHQQHQQAQTMLAGQLQSSVHVQQKFLKRPAEDVDNG 247
Query: 550 PANAIPAY 557
P + P +
Sbjct: 248 PDSFEPPH 255
Score = 35.0 bits (79), Expect = 0.71
Identities = 53/222 (23%), Positives = 78/222 (34%), Gaps = 63/222 (28%)
Query: 352 MGVPINSA---VVSGDYQNRVFSDDERSDHGVPVGYRKPPTPTPQQQAQQQLHTQTQSQS 408
MGVP + G Q + + + + G+P+G QQ+ + + TQ Q+Q
Sbjct: 1307 MGVPYGGGRGGPMGGPQQQQRPPNVQVTPDGMPMG--------SQQEWRHMMMTQQQTQM 1358
Query: 409 HPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVD 468
F GG P + + + N SG + P
Sbjct: 1359 G---FGGPGPGGPMRQGPGGFNGGNFMPNGAPNGAA---------GSGPNAGGMMTGPNV 1406
Query: 469 PKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHL---------LQQQFDQQQ-------- 511
P++ ++ Q + Q+ + +Q QQQ QQQH+ +QQ QQQ
Sbjct: 1407 PQMQLTPAQMQQQLMRQQQQ----QQQQQQQQHMGPGAANNMQMQQLLQQQQSGGGGNMM 1462
Query: 512 ---------HQQQQHQQQQHQQQQ----------HQQQQQQQ 534
H Q QQ QQQQ HQQQQ Q
Sbjct: 1463 ASQMQMTSMHMTQTQQQITMQQQQQFVQSTTTTTHQQQQMMQ 1504
Score = 33.5 bits (75), Expect = 2.1
Identities = 72/274 (26%), Positives = 89/274 (32%), Gaps = 74/274 (27%)
Query: 293 GGGGVRVQQQDQKVLGIEEQFAQLGVAAGVGVG-------------QKQDEGFVVITSPP 339
GGGGV V GV GVGVG Q +T P
Sbjct: 1234 GGGGVGVG----------------GVGVGVGVGGVGGANGGRFPNSAAQAAAMRRMTQQP 1277
Query: 340 PPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSDDERSDHGVPVGY-RKPPTPTPQQQAQQ 398
PP S P M P ++ + Q+ R+ GVP G R P PQQQ Q+
Sbjct: 1278 IPP-----SGPMMR-PQHAMYMQ---QHGGAGGGPRTGMGVPYGGGRGGPMGGPQQQ-QR 1327
Query: 399 QLHTQT----------QSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQ 448
+ Q Q H QQ++ G P P RQ P +
Sbjct: 1328 PPNVQVTPDGMPMGSQQEWRHMMMTQQQTQMGFGGPGP----------GGPMRQGPGGFN 1377
Query: 449 EQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFD 508
+ +G S P N + V Q +QQQ +QQ
Sbjct: 1378 GGNFMPNGAPNGAAGSGP-----NAGGMMTGPNVPQMQLTPAQMQQQLMRQQ-------- 1424
Query: 509 QQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHG 542
QQ QQQQ QQ + Q QQ QQQ G
Sbjct: 1425 -QQQQQQQQQQHMGPGAANNMQMQQLLQQQQSGG 1457
>PCAP_HUMAN (Q96RN5) Positive cofactor 2 glutamine/Q-rich-associated
protein (PC2 glutamine/Q-rich-associated protein)
(TPA-inducible gene-1) (TIG-1) (Activator-recruited
cofactor 105 kDa component) (ARC105) (CTG repeat protein
7a)
Length = 788
Score = 79.3 bits (194), Expect = 3e-14
Identities = 80/257 (31%), Positives = 103/257 (39%), Gaps = 45/257 (17%)
Query: 456 GTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQ 515
GT+ + HS V ++ + PQ+++Q+QQ Q QQQF QQQ QQ QQQ QQQ
Sbjct: 132 GTSGMAPHSMAV---VSTATPQTQLQLQQVALQQQQQQQQFQQQQQAALQQQQQQQQQQQ 188
Query: 516 QHQQQQHQQQQ-----HQQQQQQQQQQQYIHGTQFIHHN------PANAIPAYYPVYPSQ 564
QQ QQQ QQQQ QQQQQQ H + H N + + Q
Sbjct: 189 FQAQQSAMQQQFQAVVQQQQQLQQQQQQQQHLIKLHHQNQQQIQQQQQQLQRIAQLQLQQ 248
Query: 565 QQTHPQHAQVYYVPARQAQ---------------AYNLSMQQANMGESGKPQTPPNPTTL 609
QQ Q Q A QAQ + L Q M + Q PP P
Sbjct: 249 QQQQQQQQQQQQQQALQAQPPIQQPPMQQPQPPPSQALPQQLQQMHHTQHHQPPPQPQ-- 306
Query: 610 VPQTAAYNNPMRNAPLPKTEMTAAGYRTATG------------GSPQLVQVPANQLQHQQ 657
A N P + P +T+ + + G +P +VQ P Q Q QQ
Sbjct: 307 -QPPVAQNQPSQLPPQSQTQPLVSQAQALPGQMLYTQPPLKFVRAPMVVQQPPVQPQVQQ 365
Query: 658 QYVTYSQIHHPSQSIAP 674
Q T Q +Q +AP
Sbjct: 366 QQ-TAVQTAQAAQMVAP 381
Score = 32.3 bits (72), Expect = 4.6
Identities = 59/269 (21%), Positives = 83/269 (29%), Gaps = 29/269 (10%)
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVL 452
QQQ QQQ Q Q+ P Q PS + + Q P Q+ +
Sbjct: 251 QQQQQQQQQQQQALQAQPPIQQPPMQQPQPPPSQALPQQLQQMHHTQHHQPPPQPQQPPV 310
Query: 453 IQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQH 512
Q+ +++P PQS+ Q Q Q + Q +F +
Sbjct: 311 AQNQPSQLP--------------PQSQTQPLVSQAQALPGQMLYTQP----PLKFVRAPM 352
Query: 513 QQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANA---IPAYYPVYPSQQQTHP 569
QQ Q QQQ Q Q Q G Q I A I A +P +
Sbjct: 353 VVQQPPVQPQVQQQQTAVQTAQAAQMVAPGVQMITEALAQGGMHIRARFPPTTAVSAIPS 412
Query: 570 QHAQVYYVPARQAQAYNLSM-------QQANMGESGKPQTPPNPTTLVPQTAAYNNPMRN 622
+ P Q +L M QQ +S P P+P P + N+ + +
Sbjct: 413 SSIPLGRQPMAQVSQSSLPMLSSPSPGQQVQTPQSMPPPPQPSPQPGQPSSQP-NSNVSS 471
Query: 623 APLPKTEMTAAGYRTATGGSPQLVQVPAN 651
P P SP + P N
Sbjct: 472 GPAPSPSSFLPSPSPQPSQSPVTARTPQN 500
>CUDA_DICDI (O00841) Putative transcriptional regulator cudA
Length = 791
Score = 79.3 bits (194), Expect = 3e-14
Identities = 62/175 (35%), Positives = 79/175 (44%), Gaps = 25/175 (14%)
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVL 452
QQ QQQ+ Q QSQ Q QQ+ P N Q+P Q+Q
Sbjct: 600 QQLQQQQIQQQQQSQQQVQQQQQQQM--QQQPQQQQQQPQQQQQNQQQGQQPQQQQQQGQ 657
Query: 453 IQSGTTRVPTHSNPVDPKLNVSDP---------QSRIQVQQHHEQGYLLQQQFDQQQ--- 500
+ ++ +DP+L + + Q QQ++ Q Y LQQQ QQQ
Sbjct: 658 LD--------YTTYIDPQLQMQQAAQQQYMQQTMDQQQQQQYYMQQYHLQQQQQQQQAQR 709
Query: 501 HLLQQQFDQQQHQQQQHQQQQ---HQQQQHQQQQQQQQQQQYIHGTQFIHHNPAN 552
+L+QQQ+ QQQ QQQQ Q QQ QQQQ QQQ QQQQ Q Q + AN
Sbjct: 710 YLMQQQYMQQQAQQQQQQHQQVAIQQQQQQNQQQNQQQQNQSPQNQQSLDFQNAN 764
Score = 69.7 bits (169), Expect = 3e-11
Identities = 55/159 (34%), Positives = 72/159 (44%), Gaps = 8/159 (5%)
Query: 461 PTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQ 520
P++SN V P+ + Q +IQ QQ +Q Q QQQ +QQQ QQQ Q QQ QQ
Sbjct: 588 PSNSNSVVPQQSQQLQQQQIQQQQQSQQ-----QVQQQQQQQMQQQPQQQQQQPQQQQQN 642
Query: 521 QHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYV--P 578
Q Q QQ QQQQQQ Q + + A A Y QQQ + Q Y++
Sbjct: 643 QQQGQQPQQQQQQGQLDYTTYIDPQLQMQQA-AQQQYMQQTMDQQQQQQYYMQQYHLQQQ 701
Query: 579 ARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYN 617
+Q QA MQQ M + + Q + + Q N
Sbjct: 702 QQQQQAQRYLMQQQYMQQQAQQQQQQHQQVAIQQQQQQN 740
Score = 67.0 bits (162), Expect = 2e-10
Identities = 56/162 (34%), Positives = 71/162 (43%), Gaps = 14/162 (8%)
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVL 452
QQQ QQQ+ Q Q Q Q QQ++ P + P + +Q
Sbjct: 618 QQQQQQQMQQQPQQQQQQPQQQQQNQQQGQQPQQQQQQGQLDYTTYID---PQLQMQQAA 674
Query: 453 IQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQH 512
Q + T + + + Q QQ Q YL+QQQ+ QQQ QQQ QQQH
Sbjct: 675 QQQYMQQ--TMDQQQQQQYYMQQYHLQQQQQQQQAQRYLMQQQYMQQQ--AQQQ--QQQH 728
Query: 513 QQQ--QHQQQQHQQQQHQQQQQQQQQQQ---YIHGTQFIHHN 549
QQ Q QQQQ+QQQ QQQ Q Q QQ + + FI N
Sbjct: 729 QQVAIQQQQQQNQQQNQQQQNQSPQNQQSLDFQNANNFIDFN 770
Score = 64.3 bits (155), Expect = 1e-09
Identities = 69/258 (26%), Positives = 92/258 (34%), Gaps = 47/258 (18%)
Query: 370 FSDDERSDHGVPVGYRKPPTPTPQQQAQQQLHTQTQSQSHPAQFQQ--RSSGGTDLPSPD 427
F D G G P P + + +QS+ F S+ + +P
Sbjct: 540 FKDKCGGSSGGACGENNPDNPCSCKVCPYKQKVDHINQSYETYFNMFNPSNSNSVVPQQS 599
Query: 428 SVSSDSSLANAMSRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHE 487
+ Q+ V Q+Q +Q P+ PQ + Q QQ +
Sbjct: 600 QQLQQQQIQQQQQSQQQVQQQQQQQMQQ------------QPQQQQQQPQQQQQNQQQGQ 647
Query: 488 QGYLLQQQFD-----------QQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQ 536
Q QQQ Q Q QQQ+ QQ QQQ QQ QQ QQQQQQQQ
Sbjct: 648 QPQQQQQQGQLDYTTYIDPQLQMQQAAQQQYMQQTMDQQQQQQYYMQQYHLQQQQQQQQA 707
Query: 537 QQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGE 596
Q+Y+ Q++ QQQ Q Q V +Q Q N QQ N +
Sbjct: 708 QRYLMQQQYM-----------------QQQAQQQQQQHQQVAIQQQQQQN---QQQNQQQ 747
Query: 597 SGKPQTPPNPTTLVPQTA 614
Q+P N +L Q A
Sbjct: 748 QN--QSPQNQQSLDFQNA 763
>CLOC_DROME (O61735) Circadian locomoter output cycles Kaput protein
(dCLOCK) (dPAS1)
Length = 1027
Score = 77.8 bits (190), Expect = 1e-13
Identities = 77/234 (32%), Positives = 105/234 (43%), Gaps = 34/234 (14%)
Query: 471 LNVSDPQSRIQVQQ--HHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQ 528
LN D Q +Q QQ H + + LQQQ Q H QQ QQQHQQQQ QQQQ QQQQ Q
Sbjct: 760 LNQQDQQMMMQQQQNLHTQHQHNLQQQ--HQSHSQLQQHTQQQHQQQQQQQQQQQQQQQQ 817
Query: 529 QQQQQQQQQQYIHGTQFIHHNPAN---------AIPAYYPVYP-----SQQQTHPQHAQV 574
QQQQQQQQQQ Q + N I A+ + P SQ +P ++
Sbjct: 818 QQQQQQQQQQQQQQQQQLQLQQQNDILLREDIDDIDAFLNLSPLHSLGSQSTINPFNSS- 876
Query: 575 YYVPARQAQAYN--LSMQQANMGESGKPQTPP---NPTTLVP--QTAAYNNPMRNAPLPK 627
Q+YN ++ N + + PP N +L+ Q A ++P N +
Sbjct: 877 ---SNNNNQSYNGGSNLNNGNQNNNNRSSNPPQNNNEDSLLSCMQMATESSPSINFHM-- 931
Query: 628 TEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQI--HHPSQSIAPNSAPP 679
++ G T + + + +N +Q QQQ QI H QS + S+ P
Sbjct: 932 -GISDDGSETQSEDNKMMHTSGSNLVQQQQQQQQQQQILQQHQQQSNSFFSSNP 984
Score = 62.4 bits (150), Expect = 4e-09
Identities = 42/81 (51%), Positives = 49/81 (59%), Gaps = 6/81 (7%)
Query: 462 THSNPVDPKLN-VSDPQSRIQVQQHHEQGYLLQQQFD---QQQHLLQQQFDQQQHQQQQH 517
T SN V P+ N + Q+ +Q ++QQQ + Q QH LQQQ Q Q QQH
Sbjct: 738 TGSNAVQPQFNQYGFALNSEQMLNQQDQQMMMQQQQNLHTQHQHNLQQQH-QSHSQLQQH 796
Query: 518 QQQQHQQQQHQQQQQQQQQQQ 538
QQQHQQQQ QQQQQQQQQQQ
Sbjct: 797 TQQQHQQQQ-QQQQQQQQQQQ 816
Score = 53.9 bits (128), Expect = 1e-06
Identities = 29/79 (36%), Positives = 39/79 (48%)
Query: 465 NPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQ 524
N D +D + Q H +Q+ ++H + +Q H QQ QQ Q+QQ
Sbjct: 502 NDSDSTSMSTDSVTSRQSMMTHVSSQSQRQRSHHREHHRENHHNQSHHHMQQQQQHQNQQ 561
Query: 525 QQHQQQQQQQQQQQYIHGT 543
QQHQQ QQ QQQ Q+ GT
Sbjct: 562 QQHQQHQQLQQQLQHTVGT 580
Score = 50.8 bits (120), Expect = 1e-05
Identities = 39/122 (31%), Positives = 46/122 (36%), Gaps = 38/122 (31%)
Query: 484 QHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQ------------QQHQQQQHQQQQ 531
Q ++ G+ L + Q Q QQQ+ QHQ QQH QQQHQQQQ
Sbjct: 746 QFNQYGFALNSEQMLNQQDQQMMMQQQQNLHTQHQHNLQQQHQSHSQLQQHTQQQHQQQQ 805
Query: 532 QQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQ 591
QQQQQQ QQQ Q Q +Q Q L +QQ
Sbjct: 806 QQQQQQ--------------------------QQQQQQQQQQQQQQQQQQQQQQQLQLQQ 839
Query: 592 AN 593
N
Sbjct: 840 QN 841
Score = 40.0 bits (92), Expect = 0.022
Identities = 44/190 (23%), Positives = 66/190 (34%), Gaps = 18/190 (9%)
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSS-----------GGTDLPSPDSVSSDSSLA--NAM 439
QQQ QQQ Q Q Q Q QQ++ +L S+ S S++ N+
Sbjct: 817 QQQQQQQQQQQQQQQQQQLQLQQQNDILLREDIDDIDAFLNLSPLHSLGSQSTINPFNSS 876
Query: 440 SRQKPVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQ 499
S Y + +G S+ N S +Q+
Sbjct: 877 SNNNNQSYNGGSNLNNGNQNNNNRSSNPPQNNNEDSLLSCMQMATESSPSINFHMGISDD 936
Query: 500 QHLLQQQFDQQQHQQQQH---QQQQHQQQQHQQQQQQQQQQQYIHGTQFIH--HNPANAI 554
Q + ++ H + QQQQ QQQQ QQ QQQ + F++ + N +
Sbjct: 937 GSETQSEDNKMMHTSGSNLVQQQQQQQQQQQILQQHQQQSNSFFSSNPFLNSQNQNQNQL 996
Query: 555 PAYYPVYPSQ 564
P + P Q
Sbjct: 997 PNDLEILPYQ 1006
Score = 40.0 bits (92), Expect = 0.022
Identities = 23/68 (33%), Positives = 30/68 (43%), Gaps = 8/68 (11%)
Query: 501 HLLQQQFDQQQHQQQQH------QQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAI 554
H+ Q Q+ H ++ H Q H QQQ Q Q QQQQ QQ+ Q + H
Sbjct: 523 HVSSQSQRQRSHHREHHRENHHNQSHHHMQQQQQHQNQQQQHQQHQQLQQQLQHTVGT-- 580
Query: 555 PAYYPVYP 562
P P+ P
Sbjct: 581 PKMVPLLP 588
Score = 37.4 bits (85), Expect = 0.14
Identities = 27/99 (27%), Positives = 35/99 (35%), Gaps = 10/99 (10%)
Query: 503 LQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYP 562
+Q QF+Q Q Q QQ QQQQ Q+ H Q H + +
Sbjct: 743 VQPQFNQYGFALNSEQMLNQQDQQMMMQQQQNLHTQHQHNLQQQHQSHSQL--------- 793
Query: 563 SQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQ 601
QQ T QH Q +Q Q QQ + + Q
Sbjct: 794 -QQHTQQQHQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ 831
>PCAP_MOUSE (Q924H2) Positive cofactor 2 glutamine/Q-rich-associated
protein (PC2 glutamine/Q-rich-associated protein)
(mPcqap)
Length = 792
Score = 77.0 bits (188), Expect = 2e-13
Identities = 81/259 (31%), Positives = 109/259 (41%), Gaps = 45/259 (17%)
Query: 456 GTTRVPTHSNPVDPKLNVSDPQSRIQVQQ---HHEQGYLLQQQFDQQQHLLQQQFDQQQH 512
GT+ + H V ++ + PQ+++Q+QQ +Q QQQF QQQ LQQQ QQQ
Sbjct: 132 GTSGMAPHGMAV---VSTTTPQTQLQLQQVALQQQQQRQQQQQFQQQQAALQQQQQQQQQ 188
Query: 513 QQQQHQ--------QQQHQ----QQQHQQQQQQ--------QQQQQYIHGTQF------- 545
QQQQ Q QQQ Q QQQ QQQQQQ Q QQQ I Q
Sbjct: 189 QQQQQQFQAQQNAMQQQFQAVVQQQQLQQQQQQQHLIKLHHQSQQQQIQQQQLQRMAQLQ 248
Query: 546 ----IHHNPANAIPAYYPV-YPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKP 600
A+ A P+ PS QQ P +Q +P + +Q ++ Q P
Sbjct: 249 LQQQQQQQQQQALQAQPPMQQPSMQQPQPPPSQA--LPQQLSQLHHPQHHQPPPQAQQSP 306
Query: 601 QTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQVP-----ANQLQH 655
P + PQ+ + R LP + AA + +P +VQ P Q+Q
Sbjct: 307 IAQNQPPQIPPQSQSQPLVSRAQALPGPMLYAAQQQLKFVRAPMVVQQPQVQPQVQQVQP 366
Query: 656 QQQYVTYSQIHHPSQSIAP 674
Q Q Q +Q +AP
Sbjct: 367 QVQPQAAVQAAQSAQMVAP 385
>LUG_ARATH (Q9FUY2) Transcriptional corepressor LEUNIG
Length = 931
Score = 77.0 bits (188), Expect = 2e-13
Identities = 56/139 (40%), Positives = 68/139 (48%), Gaps = 23/139 (16%)
Query: 477 QSRIQVQQH----HEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQ 532
+ ++Q QH +Q QQQ QQ LLQ+ QQQ QQQQH Q QQQQ QQQQQ
Sbjct: 88 EQQLQQSQHPQVSQQQQQQQQQQIQMQQLLLQRAQQQQQQQQQQHHHHQQQQQQQQQQQQ 147
Query: 533 QQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQ---AYNLSM 589
QQQQQQ H Q P QQQ+ PQH Q P +Q Q +L+
Sbjct: 148 QQQQQQQQHQNQ-------------PPSQQQQQQSTPQHQQ-QPTPQQQPQRRDGSHLAN 193
Query: 590 QQAN--MGESGKPQTPPNP 606
AN +G + +P NP
Sbjct: 194 GSANGLVGNNSEPVMRQNP 212
Score = 66.6 bits (161), Expect = 2e-10
Identities = 40/91 (43%), Positives = 50/91 (53%)
Query: 447 YQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQ 506
Y E +I++ ++ +P + Q +IQ+QQ Q QQQ QQQH QQ
Sbjct: 78 YIETQMIKAREQQLQQSQHPQVSQQQQQQQQQQIQMQQLLLQRAQQQQQQQQQQHHHHQQ 137
Query: 507 FDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQ 537
QQQ QQQQ QQQQ QQ Q+Q QQQQQQ
Sbjct: 138 QQQQQQQQQQQQQQQQQQHQNQPPSQQQQQQ 168
Score = 39.7 bits (91), Expect = 0.029
Identities = 56/246 (22%), Positives = 83/246 (32%), Gaps = 34/246 (13%)
Query: 497 DQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPA 556
++ + + Q + ++ Q QQ Q Q QQQQQQQQQQ I Q + +
Sbjct: 69 EKHSEVAASYIETQMIKAREQQLQQSQHPQVSQQQQQQQQQQ-IQMQQLL-------LQR 120
Query: 557 YYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAY 616
QQQ H H Q +Q Q QQ +P + PQ
Sbjct: 121 AQQQQQQQQQQHHHHQQ----QQQQQQQQQQQQQQQQQQHQNQPPSQQQQQQSTPQHQQQ 176
Query: 617 NNPMRNAPLPKTEMTAAGYRTATGGS---PQLVQVPANQLQHQQQYVTYSQIHHPSQSIA 673
P + A G G+ P + Q P + + ++ P+Q +
Sbjct: 177 PTPQQQPQRRDGSHLANGSANGLVGNNSEPVMRQNPGSGSSLASK-AYEERVKMPTQRES 235
Query: 674 PNSAPPANYAYD---YVDPAHAQIYYSQPLAPTMPSQYQTMTAAAMVLPEGSSAQLPSDG 730
+ A + + +DP+HA I S AAA P G S G
Sbjct: 236 LDEAAMKRFGDNVGQLLDPSHASILKS---------------AAASGQPAGQVLHSTSGG 280
Query: 731 MKQQTR 736
M Q +
Sbjct: 281 MSPQVQ 286
Score = 33.9 bits (76), Expect = 1.6
Identities = 21/67 (31%), Positives = 32/67 (47%), Gaps = 9/67 (13%)
Query: 383 GYRKPPTPTPQQQAQQQL---HTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAM 439
G PP P PQ Q QL + Q QS +H Q++ GG S++ D S++N+
Sbjct: 458 GGGNPPQPQPQPQPLNQLALTNPQPQSSNHSIHQQEKLGGG------GSITMDGSISNSF 511
Query: 440 SRQKPVI 446
+ V+
Sbjct: 512 RGNEQVL 518
>DOA_DROME (P49762) Serine/threonine-protein kinase Doa (EC
2.7.1.37) (Protein darkener of apricot)
Length = 832
Score = 76.6 bits (187), Expect = 2e-13
Identities = 78/280 (27%), Positives = 105/280 (36%), Gaps = 35/280 (12%)
Query: 461 PTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQ----- 515
PT S+ + + Q + VQ + Q LLQQQ QQQ + + Q
Sbjct: 74 PTSSSSSSSSKYIGESQIPVPVQLYDPQKPLLQQQ--QQQQRICYPIGKSNSTSQLPMGG 131
Query: 516 -----QHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTH-P 569
QHQQQQH QQQ QQ Q+QQQ Q H F++ N P QQ H P
Sbjct: 132 YQRLLQHQQQQHHQQQQQQHQEQQQYPQ--HKRPFLNWNSFACSAMNGASDPFMQQQHMP 189
Query: 570 QHAQVYYVPARQAQAYNLS-------------MQQANMGESGKPQTP--PNPTTLVPQTA 614
H Q ++P + Q+Y+ S Q N + K P P ++PQ
Sbjct: 190 AHQQQQHLPHKLQQSYSSSHVPKQAPKSGLAMFLQKNTNKENKFGQPMQQQPPGMMPQMY 249
Query: 615 AYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAP 674
Y P + + + A +A+ S Q QHQQQ Q HP Q
Sbjct: 250 GYQAPQQQSKIGYPRTGAPLTHSASFSSAQRPTALQFHQQHQQQQHLQQQQQHPQQQQHQ 309
Query: 675 NSA-----PPANYAYDYVDPAHAQIYYSQPLAPTMPSQYQ 709
+S+ NY P + P A P+Q Q
Sbjct: 310 HSSFGVGMMSRNYYNMPKQPERKPLQTFDPYAYPKPNQMQ 349
Score = 45.8 bits (107), Expect = 4e-04
Identities = 72/297 (24%), Positives = 101/297 (33%), Gaps = 63/297 (21%)
Query: 392 PQQQAQQQL-HTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKP----VI 446
P Q QQ L H QS S +Q G + + + ++ M +Q P +
Sbjct: 189 PAHQQQQHLPHKLQQSYSSSHVPKQAPKSGLAMFLQKNTNKENKFGQPMQQQPPGMMPQM 248
Query: 447 YQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQ 506
Y Q Q P P+ + S Q +Q H
Sbjct: 249 YGYQAPQQQSKIGYPRTGAPLTHSASFSSAQRPTALQFH--------------------- 287
Query: 507 FDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQ 566
QQHQQQQH QQQ QQH QQQQ Q + G ++ P P+
Sbjct: 288 ---QQHQQQQHLQQQ---QQH-PQQQQHQHSSFGVGMMSRNYYNMPKQPERKPLQTFDPY 340
Query: 567 THPQHAQVYYVPARQAQAY-NLSMQQANMGESGKP----QTPPNPTTLVPQTAAYNNPMR 621
+P+ Q+ V +Q Q + + Q A+ G G Q PN T + Y +P
Sbjct: 341 AYPKPNQMQPVKYQQQQQHPHTQFQNASAGGGGGGAAGLQYDPNTNTQL----FYASP-- 394
Query: 622 NAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQ---IHHPSQSIAPN 675
A+ S + Q P Q Q QQ + S +H Q P+
Sbjct: 395 ----------------ASSSSNKQPQQPQQQQQQQQSQLQQSNSVIFNHSGQQHQPH 435
Score = 43.1 bits (100), Expect = 0.003
Identities = 56/184 (30%), Positives = 66/184 (35%), Gaps = 53/184 (28%)
Query: 385 RKPPTPTPQQQAQQQLHTQTQSQSHPAQFQ-QRSSGGTDLPSPDSVSSDSSLANAMSRQK 443
++P QQ QQQ H Q Q Q HP Q Q Q SS G + S + + +K
Sbjct: 279 QRPTALQFHQQHQQQQHLQ-QQQQHPQQQQHQHSSFGVGMMSRNYYNMPKQ-----PERK 332
Query: 444 PVIYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQH-HEQ----------GYLL 492
P+ T PK N P Q QQH H Q G
Sbjct: 333 PL---------------QTFDPYAYPKPNQMQPVKYQQQQQHPHTQFQNASAGGGGGGAA 377
Query: 493 QQQFDQQQHLL-------------QQQFDQQQHQQQQHQQQQ-------HQQQQHQQQQQ 532
Q+D + Q Q QQQ QQQQ Q QQ H QQHQ QQ
Sbjct: 378 GLQYDPNTNTQLFYASPASSSSNKQPQQPQQQQQQQQSQLQQSNSVIFNHSGQQHQPHQQ 437
Query: 533 QQQQ 536
QQ +
Sbjct: 438 QQNE 441
>FXP2_PANTR (Q8MJA0) Forkhead box protein P2
Length = 716
Score = 75.9 bits (185), Expect = 4e-13
Identities = 85/300 (28%), Positives = 123/300 (40%), Gaps = 52/300 (17%)
Query: 400 LHTQTQSQSHPAQ---FQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSG 456
LH Q Q A+ QQ++SG L SP S +Q+P+ V + +
Sbjct: 50 LHLQQQQALQAARQLLLQQQTSG---LKSPKS----------SDKQRPLQVPVSVAMMTP 96
Query: 457 TTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQ-----FDQQQH-----LLQQQ 506
P + + +S Q + +QQ +Q +LQQQ + +QQ LLQQQ
Sbjct: 97 QVITPQQMQQILQQQVLSPQQLQALLQQ--QQAVMLQQQQLQEFYKKQQEQLHLQLLQQQ 154
Query: 507 FDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQ 566
QQQ QQQQ QQQQ QQQQ QQQQQQQQQQQ Q +P QQQ
Sbjct: 155 QQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ----QQQQQQHPGKQAKEQQQQQQQQQQ 210
Query: 567 THPQ----HAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRN 622
Q Q+ + Q Q + LS+Q+ G PP L Q+
Sbjct: 211 LAAQQLVFQQQLLQMQQLQQQQHLLSLQR-----QGLISIPPGQAALPVQS--------- 256
Query: 623 APLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPNSAPPANY 682
LP+ ++ A + + + N ++H +T + + S ++PP +
Sbjct: 257 --LPQAGLSPAEIQQLWKEVTGVHSMEDNGIKHGGLDLTTNNSSSTTSSTTSKASPPITH 314
Score = 46.6 bits (109), Expect = 2e-04
Identities = 54/218 (24%), Positives = 76/218 (34%), Gaps = 52/218 (23%)
Query: 326 QKQDEGFVVITSPPPPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSDDERSDHGVPVGYR 385
Q+Q G + SP L P + V++ ++ S + +
Sbjct: 67 QQQTSG---LKSPKSSDKQRPLQVPVSVAMMTPQVITPQQMQQILQQQVLSPQQLQALLQ 123
Query: 386 KPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPV 445
+ QQQ Q+ + + Q Q H QQ+ +Q+
Sbjct: 124 QQQAVMLQQQQLQEFYKKQQEQLHLQLLQQQQQ--------------------QQQQQQQ 163
Query: 446 IYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQ 505
Q+Q Q + Q QQ +Q QQQ QQQH +Q
Sbjct: 164 QQQQQ--------------------------QQQQQQQQQQQQQQQQQQQQQQQQHPGKQ 197
Query: 506 QFDQQQHQQQQHQ---QQQHQQQQHQQQQQQQQQQQYI 540
+QQQ QQQQ Q QQ QQQ Q QQ QQQQ +
Sbjct: 198 AKEQQQQQQQQQQLAAQQLVFQQQLLQMQQLQQQQHLL 235
>FXP2_PANPA (Q8HZ00) Forkhead box protein P2
Length = 716
Score = 75.9 bits (185), Expect = 4e-13
Identities = 85/300 (28%), Positives = 123/300 (40%), Gaps = 52/300 (17%)
Query: 400 LHTQTQSQSHPAQ---FQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSG 456
LH Q Q A+ QQ++SG L SP S +Q+P+ V + +
Sbjct: 50 LHLQQQQALQAARQLLLQQQTSG---LKSPKS----------SDKQRPLQVPVSVAMMTP 96
Query: 457 TTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQ-----FDQQQH-----LLQQQ 506
P + + +S Q + +QQ +Q +LQQQ + +QQ LLQQQ
Sbjct: 97 QVITPQQMQQILQQQVLSPQQLQALLQQ--QQAVMLQQQQLQEFYKKQQEQLHLQLLQQQ 154
Query: 507 FDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQ 566
QQQ QQQQ QQQQ QQQQ QQQQQQQQQQQ Q +P QQQ
Sbjct: 155 QQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ----QQQQQQHPGKQAKEQQQQQQQQQQ 210
Query: 567 THPQ----HAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRN 622
Q Q+ + Q Q + LS+Q+ G PP L Q+
Sbjct: 211 LAAQQLVFQQQLLQMQQLQQQQHLLSLQR-----QGLISIPPGQAALPVQS--------- 256
Query: 623 APLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPNSAPPANY 682
LP+ ++ A + + + N ++H +T + + S ++PP +
Sbjct: 257 --LPQAGLSPAEIQQLWKEVTGVHSMEDNGIKHGGLDLTTNNSSSTTSSTTSKASPPITH 314
Score = 46.6 bits (109), Expect = 2e-04
Identities = 54/218 (24%), Positives = 76/218 (34%), Gaps = 52/218 (23%)
Query: 326 QKQDEGFVVITSPPPPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSDDERSDHGVPVGYR 385
Q+Q G + SP L P + V++ ++ S + +
Sbjct: 67 QQQTSG---LKSPKSSDKQRPLQVPVSVAMMTPQVITPQQMQQILQQQVLSPQQLQALLQ 123
Query: 386 KPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPV 445
+ QQQ Q+ + + Q Q H QQ+ +Q+
Sbjct: 124 QQQAVMLQQQQLQEFYKKQQEQLHLQLLQQQQQ--------------------QQQQQQQ 163
Query: 446 IYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQ 505
Q+Q Q + Q QQ +Q QQQ QQQH +Q
Sbjct: 164 QQQQQ--------------------------QQQQQQQQQQQQQQQQQQQQQQQQHPGKQ 197
Query: 506 QFDQQQHQQQQHQ---QQQHQQQQHQQQQQQQQQQQYI 540
+QQQ QQQQ Q QQ QQQ Q QQ QQQQ +
Sbjct: 198 AKEQQQQQQQQQQLAAQQLVFQQQLLQMQQLQQQQHLL 235
>FXP2_HUMAN (O15409) Forkhead box protein P2 (CAG repeat protein 44)
(Trinucleotide repeat-containing gene 10 protein)
Length = 715
Score = 75.5 bits (184), Expect = 5e-13
Identities = 76/231 (32%), Positives = 100/231 (42%), Gaps = 42/231 (18%)
Query: 400 LHTQTQSQSHPAQ---FQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSG 456
LH Q Q A+ QQ++SG L SP S +Q+P+ V + +
Sbjct: 50 LHLQQQQALQAARQLLLQQQTSG---LKSPKS----------SDKQRPLQVPVSVAMMTP 96
Query: 457 TTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQ-----FDQQQH-----LLQQQ 506
P + + +S Q + +QQ +Q +LQQQ + +QQ LLQQQ
Sbjct: 97 QVITPQQMQQILQQQVLSPQQLQALLQQ--QQAVMLQQQQLQEFYKKQQEQLHLQLLQQQ 154
Query: 507 FDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQ 566
QQQ QQQQ QQQQ QQQQ QQQQQQQQQQQ Q +P QQQ
Sbjct: 155 QQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ-----QQQQQHPGKQAKEQQQQQQQQQQ 209
Query: 567 THPQ----HAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQT 613
Q Q+ + Q Q + LS+Q+ G PP L Q+
Sbjct: 210 LAAQQLVFQQQLLQMQQLQQQQHLLSLQR-----QGLISIPPGQAALPVQS 255
Score = 46.2 bits (108), Expect = 3e-04
Identities = 54/218 (24%), Positives = 76/218 (34%), Gaps = 53/218 (24%)
Query: 326 QKQDEGFVVITSPPPPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSDDERSDHGVPVGYR 385
Q+Q G + SP L P + V++ ++ S + +
Sbjct: 67 QQQTSG---LKSPKSSDKQRPLQVPVSVAMMTPQVITPQQMQQILQQQVLSPQQLQALLQ 123
Query: 386 KPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPV 445
+ QQQ Q+ + + Q Q H QQ+ +Q+
Sbjct: 124 QQQAVMLQQQQLQEFYKKQQEQLHLQLLQQQQ---------------------QQQQQQQ 162
Query: 446 IYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQ 505
Q+Q Q + Q QQ +Q QQQ QQQH +Q
Sbjct: 163 QQQQQ--------------------------QQQQQQQQQQQQQQQQQQQQQQQQHPGKQ 196
Query: 506 QFDQQQHQQQQHQ---QQQHQQQQHQQQQQQQQQQQYI 540
+QQQ QQQQ Q QQ QQQ Q QQ QQQQ +
Sbjct: 197 AKEQQQQQQQQQQLAAQQLVFQQQLLQMQQLQQQQHLL 234
>FXP2_GORGO (Q8MJ99) Forkhead box protein P2
Length = 713
Score = 75.1 bits (183), Expect = 6e-13
Identities = 84/300 (28%), Positives = 122/300 (40%), Gaps = 55/300 (18%)
Query: 400 LHTQTQSQSHPAQ---FQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSG 456
LH Q Q A+ QQ++SG L SP S +Q+P+ V + +
Sbjct: 50 LHLQQQQALQAARQLLLQQQTSG---LKSPKS----------SDKQRPLQVPVSVAMMTP 96
Query: 457 TTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQ-----FDQQQH-----LLQQQ 506
P + + +S Q + +QQ +Q +LQQQ + +QQ LLQQQ
Sbjct: 97 QVITPQQMQQILQQQVLSPQQLQALLQQ--QQAVMLQQQQLQEFYKKQQEQLHLQLLQQQ 154
Query: 507 FDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQ 566
QQQ QQQQ QQQQ QQQQ QQQQQQQQQQQ +P QQQ
Sbjct: 155 QQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ-------QQHPGKQAKEQQQQQQQQQQ 207
Query: 567 THPQ----HAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRN 622
Q Q+ + Q Q + LS+Q+ G PP L Q+
Sbjct: 208 LAAQQLVFQQQLLQMQQLQQQQHLLSLQR-----QGLISIPPGQAALPVQS--------- 253
Query: 623 APLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPNSAPPANY 682
LP+ ++ A + + + N ++H +T + + S ++PP +
Sbjct: 254 --LPQAGLSPAEIQQLWKEVTGVHSMEDNGIKHGGLDLTTNNSSSTTSSTTSKASPPITH 311
Score = 45.4 bits (106), Expect = 5e-04
Identities = 54/218 (24%), Positives = 76/218 (34%), Gaps = 55/218 (25%)
Query: 326 QKQDEGFVVITSPPPPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSDDERSDHGVPVGYR 385
Q+Q G + SP L P + V++ ++ S + +
Sbjct: 67 QQQTSG---LKSPKSSDKQRPLQVPVSVAMMTPQVITPQQMQQILQQQVLSPQQLQALLQ 123
Query: 386 KPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPV 445
+ QQQ Q+ + + Q Q H QQ+ +Q+
Sbjct: 124 QQQAVMLQQQQLQEFYKKQQEQLHLQLLQQQ-----------------------QQQQQQ 160
Query: 446 IYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQ 505
Q+Q Q + Q QQ +Q QQQ QQQH +Q
Sbjct: 161 QQQQQ--------------------------QQQQQQQQQQQQQQQQQQQQQQQQHPGKQ 194
Query: 506 QFDQQQHQQQQHQ---QQQHQQQQHQQQQQQQQQQQYI 540
+QQQ QQQQ Q QQ QQQ Q QQ QQQQ +
Sbjct: 195 AKEQQQQQQQQQQLAAQQLVFQQQLLQMQQLQQQQHLL 232
>SPEN_DROME (Q8SX83) Split ends protein
Length = 5560
Score = 74.7 bits (182), Expect = 8e-13
Identities = 92/371 (24%), Positives = 136/371 (35%), Gaps = 43/371 (11%)
Query: 326 QKQDEGFVVITSPP-----PPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSDDERSDHGV 380
Q+ + +ITSP P PTT + P S V S YQ
Sbjct: 3689 QQTQQQHALITSPQSSNISPLASPTTRVLSSSNSPTTSKVNS--YQ-------------- 3732
Query: 381 PVGYRKPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMS 440
P + P P+P+ A+ Q T Q + P Q T + P + S +
Sbjct: 3733 PRNQQVPQQPSPKSVAEVQ--TTPQLMTIPLQKM------TPIQVPHHPTIISKVVTVQP 3784
Query: 441 RQKPVIYQEQVLIQSGTTRVPTHSNP-VDPKLNVSDPQ--SRIQVQQH--HEQGYLLQQQ 495
+Q Q QV +P H N ++ N PQ +++ QH H Q ++ QQ
Sbjct: 3785 QQAT---QSQVASSPPLGSLPPHKNVHLNAHQNQQQPQVIAKMTAHQHQQHMQQFMHQQM 3841
Query: 496 FDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIP 555
+QQH+ QQQ Q Q Q Q QQ QQQQQ QQ+++ +P
Sbjct: 3842 IQRQQHMQQQQLHGQSQQITSAPQHQMHQQHQAQQQQQHHNQQHLNQQLHAQQHPTQKQH 3901
Query: 556 AYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTL--VPQT 613
+ Q Q H Q QAQ +LS QQ + Q L + +
Sbjct: 3902 QAQQQFNQQIQQHQSQQQHQVQQQNQAQQQHLSQQQHQSQQQLNQQHQAQQQQLQQIQKL 3961
Query: 614 AAYNNPMRNAPLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQ--QYVTYSQIHHPSQS 671
+ P + P+ G + Q+PA + QQ Q +++S P+
Sbjct: 3962 QQMHGPQQQQKSPQGVGHLGGSTSIFASQQHNSQLPARGVPQQQHPQQLSHSSPCKPNTL 4021
Query: 672 IAPNSA--PPA 680
++ N PPA
Sbjct: 4022 VSVNQGVQPPA 4032
>FXP2_PONPY (Q8MJ98) Forkhead box protein P2
Length = 714
Score = 74.3 bits (181), Expect = 1e-12
Identities = 84/300 (28%), Positives = 122/300 (40%), Gaps = 54/300 (18%)
Query: 400 LHTQTQSQSHPAQ---FQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVLIQSG 456
LH Q Q A+ QQ++SG L SP S +Q+P+ V + +
Sbjct: 50 LHLQQQQALQAARQLLLQQQTSG---LKSPKS----------SDKQRPLQVPVSVAMMTP 96
Query: 457 TTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQ-----FDQQQH-----LLQQQ 506
P + + +S Q + +QQ +Q +LQQQ + +QQ LLQQQ
Sbjct: 97 QVITPQQMQQILQQQVLSPQQLQALLQQ--QQAVMLQQQQLQEFYKKQQEQLHLQLLQQQ 154
Query: 507 FDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQ 566
QQQ QQQQ QQQQ QQQQ QQQQQQQQQQQ +P QQQ
Sbjct: 155 QQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ------QQQHPGKQAKEQQQQQQQQQQ 208
Query: 567 THPQ----HAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRN 622
Q Q+ + Q Q + LS+Q+ G PP L Q+
Sbjct: 209 LAAQQLVFQQQLLQMQQLQQQQHLLSLQR-----QGLISIPPGQAALPVQS--------- 254
Query: 623 APLPKTEMTAAGYRTATGGSPQLVQVPANQLQHQQQYVTYSQIHHPSQSIAPNSAPPANY 682
LP+ ++ A + + + N ++H +T + + S ++PP +
Sbjct: 255 --LPQAGLSPAEIQQLWKEVTGVHSMEDNGIKHGGLDLTTNNSSSTTSSTTSKASPPITH 312
Score = 45.8 bits (107), Expect = 4e-04
Identities = 54/218 (24%), Positives = 76/218 (34%), Gaps = 54/218 (24%)
Query: 326 QKQDEGFVVITSPPPPPVPTTLSAPAMGVPINSAVVSGDYQNRVFSDDERSDHGVPVGYR 385
Q+Q G + SP L P + V++ ++ S + +
Sbjct: 67 QQQTSG---LKSPKSSDKQRPLQVPVSVAMMTPQVITPQQMQQILQQQVLSPQQLQALLQ 123
Query: 386 KPPTPTPQQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPV 445
+ QQQ Q+ + + Q Q H QQ+ +Q+
Sbjct: 124 QQQAVMLQQQQLQEFYKKQQEQLHLQLLQQQ----------------------QQQQQQQ 161
Query: 446 IYQEQVLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQ 505
Q+Q Q + Q QQ +Q QQQ QQQH +Q
Sbjct: 162 QQQQQ--------------------------QQQQQQQQQQQQQQQQQQQQQQQQHPGKQ 195
Query: 506 QFDQQQHQQQQHQ---QQQHQQQQHQQQQQQQQQQQYI 540
+QQQ QQQQ Q QQ QQQ Q QQ QQQQ +
Sbjct: 196 AKEQQQQQQQQQQLAAQQLVFQQQLLQMQQLQQQQHLL 233
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.127 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 96,597,826
Number of Sequences: 164201
Number of extensions: 4707876
Number of successful extensions: 76635
Number of sequences better than 10.0: 1051
Number of HSP's better than 10.0 without gapping: 664
Number of HSP's successfully gapped in prelim test: 406
Number of HSP's that attempted gapping in prelim test: 28340
Number of HSP's gapped (non-prelim): 11488
length of query: 738
length of database: 59,974,054
effective HSP length: 118
effective length of query: 620
effective length of database: 40,598,336
effective search space: 25170968320
effective search space used: 25170968320
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0328.16


