

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0327.1
(91 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
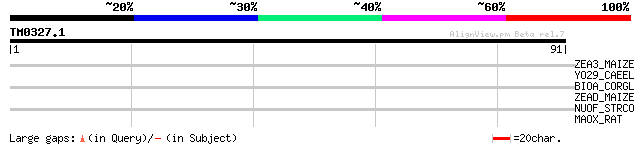
Score E
Sequences producing significant alignments: (bits) Value
ZEA3_MAIZE (P06674) Zein-alpha precursor (19 kDa) (Clone 19A2) (... 30 1.4
YO29_CAEEL (P34274) Hypothetical protein ZK688.9 in chromosome III 28 3.1
BIOA_CORGL (P46395) Adenosylmethionine-8-amino-7-oxononanoate am... 28 4.1
ZEAD_MAIZE (P24450) Zein-alpha precursor (19 kDa) (PMS2) 28 5.3
NUOF_STRCO (Q9XAQ9) NADH-quinone oxidoreductase chain F (EC 1.6.... 27 6.9
MAOX_RAT (P13697) NADP-dependent malic enzyme (EC 1.1.1.40) (NAD... 27 9.1
>ZEA3_MAIZE (P06674) Zein-alpha precursor (19 kDa) (Clone 19A2)
(Fragment)
Length = 230
Score = 29.6 bits (65), Expect = 1.4
Identities = 13/35 (37%), Positives = 20/35 (57%)
Query: 52 QALSAKGTPFDPAVYNDPSIISQRLPIVKQMTQNL 86
QA++A P P PS + Q+LP+V + QN+
Sbjct: 57 QAIAAGILPLSPLFLQQPSALLQQLPLVHLLAQNI 91
>YO29_CAEEL (P34274) Hypothetical protein ZK688.9 in chromosome III
Length = 429
Score = 28.5 bits (62), Expect = 3.1
Identities = 22/73 (30%), Positives = 35/73 (47%), Gaps = 7/73 (9%)
Query: 9 LVLMQLRVDGVLIRLRDTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYND 68
L+ +RVD VL+R+ DTR + G + ++RE RE+ + L DP D
Sbjct: 351 LLRFYMRVDNVLLRVCDTR---IVGNEFDGHVIREWQLREAKYGNLG----HVDPEELLD 403
Query: 69 PSIISQRLPIVKQ 81
LP+V++
Sbjct: 404 VDRAWMHLPVVEE 416
>BIOA_CORGL (P46395) Adenosylmethionine-8-amino-7-oxononanoate
aminotransferase (EC 2.6.1.62) (7,8-diamino-pelargonic
acid aminotransferase) (DAPA aminotransferase)
Length = 423
Score = 28.1 bits (61), Expect = 4.1
Identities = 16/54 (29%), Positives = 25/54 (45%), Gaps = 6/54 (11%)
Query: 25 DTRMHCVFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIISQRLPI 78
DT H +FGG T+ ++ T + L+ G FD Y+D +S + I
Sbjct: 72 DTMSHVMFGGLTHEPAIK------LTHKLLNLTGNSFDHVFYSDSGSVSVEVAI 119
>ZEAD_MAIZE (P24450) Zein-alpha precursor (19 kDa) (PMS2)
Length = 233
Score = 27.7 bits (60), Expect = 5.3
Identities = 12/35 (34%), Positives = 19/35 (54%)
Query: 52 QALSAKGTPFDPAVYNDPSIISQRLPIVKQMTQNL 86
QA++ P P PS + Q+LP+V + QN+
Sbjct: 60 QAIATGILPLSPLFLQQPSALLQQLPLVHLVAQNI 94
>NUOF_STRCO (Q9XAQ9) NADH-quinone oxidoreductase chain F (EC
1.6.99.5) (NADH dehydrogenase I, chain F) (NDH-1, chain
F)
Length = 449
Score = 27.3 bits (59), Expect = 6.9
Identities = 12/36 (33%), Positives = 20/36 (55%), Gaps = 1/36 (2%)
Query: 30 CVFG-GSTNPVILRESCWRESTFQALSAKGTPFDPA 64
C G G+ +P+ +RE + ++ +G PFDPA
Sbjct: 401 CALGDGAASPIFSSLKYFREEYEEHITGRGCPFDPA 436
>MAOX_RAT (P13697) NADP-dependent malic enzyme (EC 1.1.1.40)
(NADP-ME) (Malic enzyme 1)
Length = 572
Score = 26.9 bits (58), Expect = 9.1
Identities = 10/26 (38%), Positives = 16/26 (61%)
Query: 43 ESCWRESTFQALSAKGTPFDPAVYND 68
E C++ + +A+ A G+PFDP D
Sbjct: 418 EECYKVTKGRAIFASGSPFDPVTLPD 443
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.331 0.144 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,723,552
Number of Sequences: 164201
Number of extensions: 337098
Number of successful extensions: 927
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 924
Number of HSP's gapped (non-prelim): 6
length of query: 91
length of database: 59,974,054
effective HSP length: 67
effective length of query: 24
effective length of database: 48,972,587
effective search space: 1175342088
effective search space used: 1175342088
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0327.1


