

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0322.5
(210 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
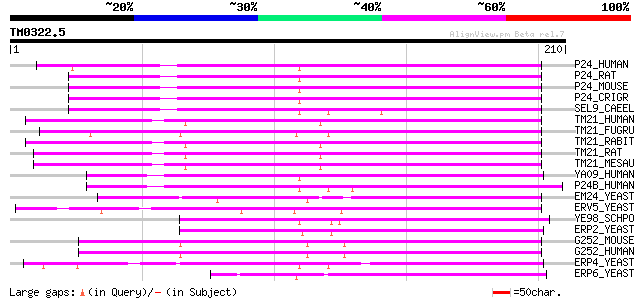
Score E
Sequences producing significant alignments: (bits) Value
P24_HUMAN (Q15363) Cop-coated vesicle membrane protein p24 precu... 99 7e-21
P24_RAT (Q63524) Cop-coated vesicle membrane protein p24 precurs... 99 9e-21
P24_MOUSE (Q9R0Q3) Cop-coated vesicle membrane protein p24 precu... 99 9e-21
P24_CRIGR (P49020) Cop-coated vesicle membrane protein p24 precu... 99 9e-21
SEL9_CAEEL (O17528) Suppressor/enhancer of lin-12 protein 9 prec... 83 4e-16
TM21_HUMAN (P49755) Transmembrane protein Tmp21 precursor (21 kD... 83 5e-16
TM21_FUGRU (Q90515) Transmembrane protein Tmp21 precursor (S31II... 82 9e-16
TM21_RABIT (Q28735) Transmembrane protein Tmp21 precursor (21 kD... 81 2e-15
TM21_RAT (Q63584) Transmembrane protein Tmp21 precursor (21 kDa ... 77 2e-14
TM21_MESAU (O35587) Transmembrane protein Tmp21 precursor (21 kD... 77 3e-14
YA09_HUMAN (Q9Y3B3) Hypothetical protein CGI-109 precursor 75 8e-14
P24B_HUMAN (Q9Y3Q3) Membrane protein p24B precursor (UNQ5357/PRO... 69 8e-12
EM24_YEAST (P32803) Endosomal P24B protein precursor (24 kDa end... 69 8e-12
ERV5_YEAST (P54837) ERV25 protein precursor 65 9e-11
YE98_SCHPO (O13770) Hypothetical protein C17A5.08 in chromosome ... 64 3e-10
ERP2_YEAST (P39704) ERP2 protein precursor 60 4e-09
G252_MOUSE (Q99KF1) Glycoprotein 25L2 precursor 60 5e-09
G252_HUMAN (Q9BVK6) Glycoprotein 25L2 precursor 59 6e-09
ERP4_YEAST (Q12450) ERP4 protein precursor 54 3e-07
ERP6_YEAST (P53198) ERP6 protein precursor 53 5e-07
>P24_HUMAN (Q15363) Cop-coated vesicle membrane protein p24
precursor (p24A)
Length = 201
Score = 99.0 bits (245), Expect = 7e-21
Identities = 58/195 (29%), Positives = 97/195 (49%), Gaps = 10/195 (5%)
Query: 11 VIFFLGTLIYNV--FSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFT 68
++ L L+ V + +SI + EC E SG + F V + ID
Sbjct: 7 LLVLLAALLATVSGYFVSIDAHAEECFFERVTSGTKMGLIFEVAEGGFL------DIDVE 60
Query: 69 VTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMST--PETVSFYIHVGHIPSEHD 126
+T P ++ S K+ F A +G YKFCF N MST P+ V F I +G P D
Sbjct: 61 ITGPDNKGIYKGDRESSGKYTFAAHMDGTYKFCFSNRMSTMTPKIVMFTIDIGEAPKGQD 120
Query: 127 LAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILL 186
+ + H + + I EL A+ ++ EQ+Y++ R+ HR N++T +RV+ ++ E ++L
Sbjct: 121 METEAHQNKLEEMINELAVAMTAVKHEQEYMEVRERIHRAINDNTNSRVVLWSFFEALVL 180
Query: 187 AVTSILQVVYIRRLF 201
++ Q+ Y++R F
Sbjct: 181 VAMTLGQIYYLKRFF 195
>P24_RAT (Q63524) Cop-coated vesicle membrane protein p24 precursor
(p24A) (RNP21.4)
Length = 201
Score = 98.6 bits (244), Expect = 9e-21
Identities = 55/181 (30%), Positives = 91/181 (49%), Gaps = 8/181 (4%)
Query: 23 FSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKG 82
+ +SI + EC E SG + F V + ID +T P ++
Sbjct: 21 YFVSIDAHAEECFFERVTSGTKMGLIFEVAEGGFL------DIDVEITGPDNKGIYKGDR 74
Query: 83 TSGDKFEFKAPQNGIYKFCFHNPMST--PETVSFYIHVGHIPSEHDLAKDEHLDPINVKI 140
S K+ F A +G YKFCF N MST P+ V F I +G P D+ + H + + I
Sbjct: 75 ESSGKYTFAAHMDGTYKFCFSNRMSTMTPKIVMFTIDIGEAPKGQDMETEAHQNKLEEMI 134
Query: 141 AELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYIRRL 200
EL A+ ++ EQ+Y++ R+ HR N++T +RV+ ++ E ++L ++ Q+ Y++R
Sbjct: 135 NELAVAMTAVKHEQEYMEVRERIHRAINDNTNSRVVLWSFFEALVLVAMTLGQIYYLKRF 194
Query: 201 F 201
F
Sbjct: 195 F 195
>P24_MOUSE (Q9R0Q3) Cop-coated vesicle membrane protein p24
precursor (p24A) (Sid 394)
Length = 201
Score = 98.6 bits (244), Expect = 9e-21
Identities = 55/181 (30%), Positives = 91/181 (49%), Gaps = 8/181 (4%)
Query: 23 FSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKG 82
+ +SI + EC E SG + F V + ID +T P ++
Sbjct: 21 YFVSIDAHAEECFFERVTSGTKMGLIFEVAEGGFL------DIDVEITGPDNKGIYKGDR 74
Query: 83 TSGDKFEFKAPQNGIYKFCFHNPMST--PETVSFYIHVGHIPSEHDLAKDEHLDPINVKI 140
S K+ F A +G YKFCF N MST P+ V F I +G P D+ + H + + I
Sbjct: 75 ESSGKYTFAAHMDGTYKFCFSNRMSTMTPKIVMFTIDIGEAPKGQDMETEAHQNKLEEMI 134
Query: 141 AELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYIRRL 200
EL A+ ++ EQ+Y++ R+ HR N++T +RV+ ++ E ++L ++ Q+ Y++R
Sbjct: 135 NELAVAMTAVKHEQEYMEVRERIHRAINDNTNSRVVLWSFFEALVLVAMTLGQIYYLKRF 194
Query: 201 F 201
F
Sbjct: 195 F 195
>P24_CRIGR (P49020) Cop-coated vesicle membrane protein p24
precursor (Fragment)
Length = 196
Score = 98.6 bits (244), Expect = 9e-21
Identities = 55/181 (30%), Positives = 91/181 (49%), Gaps = 8/181 (4%)
Query: 23 FSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKG 82
+ +SI + EC E SG + F V + ID +T P ++
Sbjct: 16 YFVSIDAHAEECFFERVTSGTKMGLIFEVAEGGFL------DIDVEITGPDNKGIYKGDR 69
Query: 83 TSGDKFEFKAPQNGIYKFCFHNPMST--PETVSFYIHVGHIPSEHDLAKDEHLDPINVKI 140
S K+ F A +G YKFCF N MST P+ V F I +G P D+ + H + + I
Sbjct: 70 ESSGKYTFAAHMDGTYKFCFSNRMSTMTPKIVMFTIDIGEAPKGQDMETEAHQNKLEEMI 129
Query: 141 AELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYIRRL 200
EL A+ ++ EQ+Y++ R+ HR N++T +RV+ ++ E ++L ++ Q+ Y++R
Sbjct: 130 NELAVAMTAVKHEQEYMEVRERIHRAINDNTNSRVVLWSFFEALVLVAMTLGQIYYLKRF 189
Query: 201 F 201
F
Sbjct: 190 F 190
>SEL9_CAEEL (O17528) Suppressor/enhancer of lin-12 protein 9
precursor
Length = 203
Score = 83.2 bits (204), Expect = 4e-16
Identities = 51/185 (27%), Positives = 89/185 (47%), Gaps = 12/185 (6%)
Query: 23 FSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKG 82
+ + + N+ +C + SG + F V + ID +T P ++ +
Sbjct: 19 YFIHVDANEEQCFFDRLTSGTKMGLMFEVAEGGFL------DIDVKITGPDNKEIYKGER 72
Query: 83 TSGDKFEFKAPQNGIYKFCFHNPMST--PETVSFYIHVG--HIPSEHDLAKDEHLDPINV 138
S KF F A +G+Y +CF N MST P+ V F + + H + A + D +
Sbjct: 73 ESSGKFTFAAHMDGVYTYCFGNKMSTMTPKAVMFTVEITEPHQQAPGAAANQDAADNAKL 132
Query: 139 K--IAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVY 196
+ + EL AL S+ EQ+Y++ R+ HR NE+T +RV+ + E +L ++ Q+ Y
Sbjct: 133 EEMVRELSSALMSVKHEQEYMEVRERVHRNINENTNSRVVMWAAFEAFVLVGMTVGQIFY 192
Query: 197 IRRLF 201
++R F
Sbjct: 193 LKRFF 197
>TM21_HUMAN (P49755) Transmembrane protein Tmp21 precursor (21 kDa
Transmembrane trafficking protein) (p24delta)
(S31III125) (S31I125) (Tmp-21-I)
Length = 219
Score = 82.8 bits (203), Expect = 5e-16
Identities = 53/201 (26%), Positives = 94/201 (46%), Gaps = 10/201 (4%)
Query: 7 ICTLVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGI- 65
+ L++F LG + S + +N +C+ E VTG + + D S G+
Sbjct: 16 LALLLLFLLGPRLVLAISFHLPINSRKCLREEIHKDLLVTGAYEISDQ----SGGAGGLR 71
Query: 66 -DFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYI----HVGH 120
+T+ G+ ++S + + KF F +++ CF + + I H
Sbjct: 72 SHLKITDSAGHILYSKEDATKGKFAFTTEDYDMFEVCFESKGTGRIPDQLVILDMKHGVE 131
Query: 121 IPSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTV 180
+ ++AK E L P+ V++ L + ESI+ + Y+K R+ R TNEST TRV+++++
Sbjct: 132 AKNYEEIAKVEKLKPLEVELRRLEDLSESIVNDFAYMKKREEEMRDTNESTNTRVLYFSI 191
Query: 181 LEYILLAVTSILQVVYIRRLF 201
L + QV Y+RR F
Sbjct: 192 FSMFCLIGLATWQVFYLRRFF 212
>TM21_FUGRU (Q90515) Transmembrane protein Tmp21 precursor
(S31III125)
Length = 213
Score = 82.0 bits (201), Expect = 9e-16
Identities = 54/200 (27%), Positives = 97/200 (48%), Gaps = 10/200 (5%)
Query: 12 IFFLGTLIYNVFSLSISL--NDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHP----GI 65
+ FL LI + FS+S L N +C+ E VTG + + + + + G
Sbjct: 7 LLFLPVLIESAFSISFFLPVNTRKCLREEIHKDVLVTGEYEISEQVVTVHTSSTVVGDGS 66
Query: 66 DFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMS---TPETVSFYIHVG-HI 121
F +T+ + ++S + + KF F +++ CF + + + V+ + G
Sbjct: 67 IFKITDSSSHTLYSKEDATKGKFAFTTEDYDMFEVCFESKCTGRVPDQLVNLDMKHGVEA 126
Query: 122 PSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVL 181
+ ++AK E L P+ V++ L + ESI+ + Y+K R+ R TNEST TRV+++++
Sbjct: 127 KNYEEIAKVEKLKPLEVELRRLEDLSESIVNDFAYMKKREEEMRDTNESTNTRVLYFSIF 186
Query: 182 EYILLAVTSILQVVYIRRLF 201
L + QV Y+RR F
Sbjct: 187 SMFCLIGLATWQVFYLRRFF 206
>TM21_RABIT (Q28735) Transmembrane protein Tmp21 precursor (21 kDa
Transmembrane trafficking protein) (Integral membrane
protein p23)
Length = 219
Score = 81.3 bits (199), Expect = 2e-15
Identities = 54/201 (26%), Positives = 92/201 (44%), Gaps = 10/201 (4%)
Query: 7 ICTLVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGI- 65
+ L +F LG S + +N +C+ E VTG + + D S G+
Sbjct: 16 LALLFLFLLGPSSVLAISFHLPVNSRKCLREEIHKDLLVTGAYEITDQ----SGGAGGLR 71
Query: 66 -DFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYI----HVGH 120
+T+ G+ ++S + S KF F +++ CF + + I H
Sbjct: 72 THLKITDSAGHILYSKEDASKGKFAFTTEDYDMFEVCFESKGTGRIPDQLVILDMKHGVE 131
Query: 121 IPSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTV 180
+ ++AK E L P+ V++ L + ESI+ + Y+K R+ R TNEST TRV+++++
Sbjct: 132 AKNYEEIAKVEKLKPLEVELRRLEDLSESIVNDFAYMKKREEEMRDTNESTNTRVLYFSI 191
Query: 181 LEYILLAVTSILQVVYIRRLF 201
L + QV Y+RR F
Sbjct: 192 FSMFCLIGLATWQVFYLRRFF 212
>TM21_RAT (Q63584) Transmembrane protein Tmp21 precursor (21 kDa
Transmembrane trafficking protein) (Fragment)
Length = 203
Score = 77.4 bits (189), Expect = 2e-14
Identities = 52/198 (26%), Positives = 91/198 (45%), Gaps = 10/198 (5%)
Query: 10 LVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGI--DF 67
L +F LG S + +N +C+ E VTG + + D S G+
Sbjct: 3 LFLFLLGPSSVLGISFHLPVNSRKCLREEIHKDLLVTGAYEITDQ----SGGAGGLRTHL 58
Query: 68 TVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYI----HVGHIPS 123
+T+ G+ +++ + + KF F +++ CF + + I H +
Sbjct: 59 KITDSAGHILYAKEDATKGKFAFTTEDYDMFEVCFESKGTGRIPDQLVILDMKHGVEAKN 118
Query: 124 EHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEY 183
++AK E L P+ V++ L + ESI+ + Y+K R+ R TNEST TRV+++++
Sbjct: 119 YEEIAKVEKLKPLEVELRRLEDLSESIVNDFAYMKKREEEMRDTNESTNTRVLYFSIFSM 178
Query: 184 ILLAVTSILQVVYIRRLF 201
L + QV Y+RR F
Sbjct: 179 FCLIGLATWQVFYLRRFF 196
>TM21_MESAU (O35587) Transmembrane protein Tmp21 precursor (21 kDa
Transmembrane trafficking protein) (Integral membrane
protein p23)
Length = 219
Score = 77.0 bits (188), Expect = 3e-14
Identities = 51/198 (25%), Positives = 91/198 (45%), Gaps = 10/198 (5%)
Query: 10 LVIFFLGTLIYNVFSLSISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGI--DF 67
L + LG S + +N +C+ E VTG + + D S G+
Sbjct: 19 LFLLLLGPSSVLAISFHLPVNSRKCLREEIHKDLLVTGAYEITDQ----SGGAGGLRTHL 74
Query: 68 TVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTPETVSFYI----HVGHIPS 123
+T+ G+ +++ + + KF F +++ CF + + I H +
Sbjct: 75 KITDSAGHILYAKEDATKGKFAFTTEDYDMFEVCFESKGTGRIPDQLVILDMKHGVEAKN 134
Query: 124 EHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEY 183
++AK E L P+ V++ L + ESI+ + Y+K R+ R TNEST TRV+++++
Sbjct: 135 YEEIAKVEKLKPLEVELRRLEDLSESIVNDFAYMKKREEEMRDTNESTNTRVLYFSIFSM 194
Query: 184 ILLAVTSILQVVYIRRLF 201
+ L + QV Y+RR F
Sbjct: 195 LCLIGLATWQVFYLRRFF 212
>YA09_HUMAN (Q9Y3B3) Hypothetical protein CGI-109 precursor
Length = 215
Score = 75.5 bits (184), Expect = 8e-14
Identities = 53/175 (30%), Positives = 80/175 (45%), Gaps = 8/175 (4%)
Query: 30 NDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGTSGDKFE 89
N +C E G T F V+ + H +D + +P G ++ D F
Sbjct: 36 NAKQCFYEDIAQGTKSTLEFQVI------TGGHYDVDCRLEDPDGKVLYKEMKKQYDSFT 89
Query: 90 FKAPQNGIYKFCFHNPMST--PETVSFYIHVGHIPSEHDLAKDEHLDPINVKIAELREAL 147
F A +NG YKFCF N ST +TV F VG + + L + + EAL
Sbjct: 90 FTASKNGTYKFCFSNEFSTFTHKTVYFDFQVGETHLCFLVDRVSALTQMESACVSIHEAL 149
Query: 148 ESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYIRRLFS 202
+S+I Q + + R+ + R E TRV +++V E ++L V SI QV ++ FS
Sbjct: 150 KSVIDYQTHFRLREAQGRSRAEDLNTRVAYWSVGEALILLVVSIGQVFLLKSFFS 204
>P24B_HUMAN (Q9Y3Q3) Membrane protein p24B precursor
(UNQ5357/PRO1078)
Length = 217
Score = 68.9 bits (167), Expect = 8e-12
Identities = 50/184 (27%), Positives = 87/184 (47%), Gaps = 10/184 (5%)
Query: 30 NDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGTSGDKFE 89
N +C E + G + ++ V+ + H +D V +P GN ++ D F
Sbjct: 36 NAKQCFHEEVEQGVKFSLDYQVI------TGGHYDVDCYVEDPQGNTIYRETKKQYDSFT 89
Query: 90 FKAPQNGIYKFCFHNPMST--PETVSFYIHVG-HIPSEHDLA-KDEHLDPINVKIAELRE 145
++A G+Y+FCF N ST +TV F VG P D+ + L + + E
Sbjct: 90 YRAEVKGVYQFCFSNEFSTFSHKTVYFDFQVGDEPPILPDMGNRVTALTQMESACVTIHE 149
Query: 146 ALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYIRRLFSKSF 205
AL+++I Q + + R+ + R E +RV +++V E I L V S QV+ ++ F++
Sbjct: 150 ALKTVIDSQTHYRLREAQDRARAEDLNSRVSYWSVGETIALFVVSFSQVLLLKSFFTEKR 209
Query: 206 AYNR 209
+R
Sbjct: 210 PISR 213
>EM24_YEAST (P32803) Endosomal P24B protein precursor (24 kDa
endomembrane protein) (Basic 24 kDa late endocytic
intermediate component)
Length = 203
Score = 68.9 bits (167), Expect = 8e-12
Identities = 49/171 (28%), Positives = 86/171 (49%), Gaps = 8/171 (4%)
Query: 34 CVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAV-HSLKGTSGDKFEFKA 92
C E GD ++ +F D + SS G DF + P + V +++ TS + A
Sbjct: 32 CFFEDLSKGDELSISFQFGDRNPQSSSQLTG-DFIIYGPERHEVLKTVRDTSHGEITLSA 90
Query: 93 PQNGIYKFCFHNPMSTPET--VSFYIHVGHIPSEHDLAKDEHLDPINVKIAELREALESI 150
P G +++CF N + ET V+F IH G + + D D + + ++ + +L + +
Sbjct: 91 PYKGHFQYCFLNENTGIETKDVTFNIH-GVVYVDLD---DPNTNTLDSAVRKLSKLTREV 146
Query: 151 ITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQVVYIRRLF 201
EQ Y+ R+ HR T EST RV ++++ + ++ S+ Q+ Y+RR F
Sbjct: 147 KDEQSYIVIRERTHRNTAESTNDRVKWWSIFQLGVVIANSLFQIYYLRRFF 197
>ERV5_YEAST (P54837) ERV25 protein precursor
Length = 211
Score = 65.5 bits (158), Expect = 9e-11
Identities = 53/207 (25%), Positives = 92/207 (43%), Gaps = 16/207 (7%)
Query: 3 WFSKICTLVIFFLGTLIYNVFSLSISLNDVE-CVSEHAQSGDSVTGNFVVMDHDIFWSSD 61
W + + +LV+ G F ++ S + + C+ + G V + H D
Sbjct: 7 WLTTLISLVVAVQGLH----FDIAASTDPEQVCIRDFVTEGQLVVADI----HSDGSVGD 58
Query: 62 HPGIDFTVTNPGGNAVHSLKGTSGD-KFEFKAPQNGIYKFCFHNPM-----STPETVSFY 115
++ V + GN + +GD + F AP + + CF N S +
Sbjct: 59 GQKLNLFVRDSVGNEYRRKRDFAGDVRVAFTAPSSTAFDVCFENQAQYRGRSLSRAIELD 118
Query: 116 IHVGHIPSE-HDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTR 174
I G + + ++ +E L PI V++ + E + I+ E YLK R+ R R TNEST R
Sbjct: 119 IESGAEARDWNKISANEKLKPIEVELRRVEEITDEIVDELTYLKNREERLRDTNESTNRR 178
Query: 175 VIFYTVLEYILLAVTSILQVVYIRRLF 201
V +++L I+L+ + QV Y++ F
Sbjct: 179 VRNFSILVIIVLSSLGVWQVNYLKNYF 205
>YE98_SCHPO (O13770) Hypothetical protein C17A5.08 in chromosome I
precursor
Length = 210
Score = 63.5 bits (153), Expect = 3e-10
Identities = 43/146 (29%), Positives = 72/146 (48%), Gaps = 6/146 (4%)
Query: 65 IDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMST--PETVSFYIHVGH-- 120
+D+ + P V K F + G Y+FCF N MST + V+ I + +
Sbjct: 60 VDYMIKAPSQKTVALGKKRRQADVFFTLEEKGEYEFCFDNHMSTFTDKIVTMEITMENEL 119
Query: 121 -IPS-EHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFY 178
+P+ D AKD D + + E+ AL I Q Y K R++R+ T +ST+ R+ ++
Sbjct: 120 SLPALTRDEAKDYKKDSMQSTVLEISTALSEIDRVQNYFKTREHRNYSTVKSTQARIFWF 179
Query: 179 TVLEYILLAVTSILQVVYIRRLFSKS 204
++ E I++ S LQV ++ F +S
Sbjct: 180 SLAESIMVVALSALQVFIVKTFFKRS 205
>ERP2_YEAST (P39704) ERP2 protein precursor
Length = 215
Score = 60.1 bits (144), Expect = 4e-09
Identities = 44/140 (31%), Positives = 69/140 (48%), Gaps = 2/140 (1%)
Query: 65 IDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMSTP-ETVSFYIHVGH-IP 122
IDF +T P G+ + S K F K+ G Y FCF N T + V + +
Sbjct: 69 IDFDITAPDGSVITSEKQKKYSDFLLKSFGVGKYTFCFSNNYGTALKKVEITLEKEKTLT 128
Query: 123 SEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLE 182
EH+ + N + E+ L I YL+AR+ R+ T ST++R+ + ++L
Sbjct: 129 DEHEADVNNDDIIANNAVEEIDRNLNKITKTLNYLRAREWRNMSTVNSTESRLTWLSILI 188
Query: 183 YILLAVTSILQVVYIRRLFS 202
I++AV SI QV+ I+ LF+
Sbjct: 189 IIIIAVISIAQVLLIQFLFT 208
>G252_MOUSE (Q99KF1) Glycoprotein 25L2 precursor
Length = 214
Score = 59.7 bits (143), Expect = 5e-09
Identities = 44/188 (23%), Positives = 86/188 (45%), Gaps = 13/188 (6%)
Query: 27 ISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHP-----GIDFTVTNPGGNAVHSLK 81
I + +C E V GN+ +D P G+ V +P + + +
Sbjct: 21 IGETEKKCFIEEIPDETMVIGNYRTQLYDKQREEYQPATPGLGMFVEVKDPEDKVILARQ 80
Query: 82 GTSGDKFEFKAPQNGIYKFCFHNPMSTPET-------VSFYIHVGHIPSEH-DLAKDEHL 133
S +F F + G ++ C H+ + V I VG +++ ++A + L
Sbjct: 81 YGSEGRFTFTSHTPGEHQICLHSNSTKFSLFAGGMLRVHLDIQVGEHANDYAEIAAKDKL 140
Query: 134 DPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQ 193
+ +++ +L E +E I EQ Y + R+ R R T+EST RV++++VL+ ++L + Q
Sbjct: 141 SELQLRVRQLVEQVEQIQKEQNYQRWREERFRQTSESTNQRVLWWSVLQTLILLAIGVCQ 200
Query: 194 VVYIRRLF 201
+ +++ F
Sbjct: 201 MRHLKSFF 208
>G252_HUMAN (Q9BVK6) Glycoprotein 25L2 precursor
Length = 214
Score = 59.3 bits (142), Expect = 6e-09
Identities = 43/188 (22%), Positives = 86/188 (44%), Gaps = 13/188 (6%)
Query: 27 ISLNDVECVSEHAQSGDSVTGNFVVMDHDIFWSSDHP-----GIDFTVTNPGGNAVHSLK 81
I + +C E V GN+ +D P G+ V +P + + +
Sbjct: 21 IGETEKKCFIEEIPDETMVIGNYRTQLYDKQREEYQPATPGLGMFVEVKDPEDKVILARQ 80
Query: 82 GTSGDKFEFKAPQNGIYKFCFHNPMSTPET-------VSFYIHVGHIPSEH-DLAKDEHL 133
S +F F + G ++ C H+ + V I VG +++ ++A + L
Sbjct: 81 YGSEGRFTFTSHTPGEHQICLHSNSTKFSLFAGGMLRVHLDIQVGEHANDYAEIAAKDKL 140
Query: 134 DPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSILQ 193
+ +++ +L E +E I EQ Y + R+ R R T+EST RV+++++L+ ++L + Q
Sbjct: 141 SELQLRVRQLVEQVEQIQKEQNYQRWREERFRQTSESTNQRVLWWSILQTLILVAIGVWQ 200
Query: 194 VVYIRRLF 201
+ +++ F
Sbjct: 201 MRHLKSFF 208
>ERP4_YEAST (Q12450) ERP4 protein precursor
Length = 207
Score = 53.9 bits (128), Expect = 3e-07
Identities = 54/207 (26%), Positives = 94/207 (45%), Gaps = 18/207 (8%)
Query: 6 KICTLV-IFFLGTLIYNVFS-------LSISLNDVECVSEHAQSGDSVTGNFVVMDHDIF 57
++ TL+ I F +L+ + FS +S+ EC+ S V +V+ + +
Sbjct: 2 RVFTLIAILFSSSLLTHAFSSNYAPVGISLPAFTKECLYYDLSSDKDV----LVVSYQVL 57
Query: 58 WSSDHPGIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCFHNPMST-PETVSFYI 116
+ IDF +T P G+ + + + F K+ G Y FC N T P+ V +
Sbjct: 58 TGGNFE-IDFDITAPDGSVIVTERQKKHSDFLLKSFGIGKYTFCLSNNYGTSPKKVEITL 116
Query: 117 HVG-HIPSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRV 175
I S H+ +D N I E+ L I YL+AR+ R+ YT ST++R+
Sbjct: 117 EKEKEIVSSHESKEDIIA---NNAIEEIDRNLNKITKTMDYLRAREWRNMYTVSSTESRL 173
Query: 176 IFYTVLEYILLAVTSILQVVYIRRLFS 202
+ ++L ++ SI+Q + I+ F+
Sbjct: 174 TWLSLLIMGVMVGISIVQALIIQFFFT 200
>ERP6_YEAST (P53198) ERP6 protein precursor
Length = 216
Score = 53.1 bits (126), Expect = 5e-07
Identities = 36/132 (27%), Positives = 65/132 (48%), Gaps = 7/132 (5%)
Query: 77 VHSLKGTSGDKFEFKAPQNGIYKFCFHNPMS-----TPETVSFYIHVGHIPSEHDLAKDE 131
VH SGD F F A ++G YK C + ++ T + VG + D+ + E
Sbjct: 83 VHQQGSPSGD-FSFLALESGEYKICLQSRVNNWVGKTKTKLEIEFEVG-FEAMLDMQRKE 140
Query: 132 HLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYILLAVTSI 191
L+ ++ K++ L + I EQ+ ++ R+ R +ES +R +++TV + LL + +
Sbjct: 141 TLESLHGKVSILNSKIVDIRREQQLMREREESFRDISESVNSRAMWWTVTQVTLLIIICV 200
Query: 192 LQVVYIRRLFSK 203
Q+ +R F K
Sbjct: 201 WQMKSLRSFFVK 212
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,009,405
Number of Sequences: 164201
Number of extensions: 1051138
Number of successful extensions: 2564
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 16
Number of HSP's that attempted gapping in prelim test: 2516
Number of HSP's gapped (non-prelim): 42
length of query: 210
length of database: 59,974,054
effective HSP length: 105
effective length of query: 105
effective length of database: 42,732,949
effective search space: 4486959645
effective search space used: 4486959645
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0322.5


