

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0322.2
(856 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
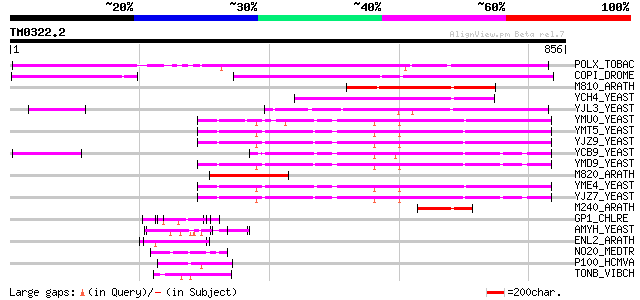
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 453 e-126
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 338 3e-92
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 208 4e-53
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 162 3e-39
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 139 3e-32
YMU0_YEAST (Q04670) Transposon Ty1 protein B 127 1e-28
YMT5_YEAST (Q04214) Transposon Ty1 protein B 125 5e-28
YJZ9_YEAST (P47100) Transposon Ty1 protein B 124 8e-28
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 124 1e-27
YMD9_YEAST (Q03434) Transposon Ty1 protein B 122 3e-27
M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820... 122 5e-27
YME4_YEAST (Q04711) Transposon Ty1 protein B 121 9e-27
YJZ7_YEAST (P47098) Transposon Ty1 protein B 119 3e-26
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 68 1e-10
GP1_CHLRE (Q9FPQ6) Vegetative cell wall protein gp1 precursor (H... 55 8e-07
AMYH_YEAST (P08640) Glucoamylase S1/S2 precursor (EC 3.2.1.3) (G... 52 7e-06
ENL2_ARATH (Q9T076) Early nodulin-like protein 2 precursor (Phyt... 50 3e-05
NO20_MEDTR (P93329) Early nodulin 20 precursor (N-20) 50 3e-05
P100_HCMVA (P08318) Large structural phosphoprotein (pp150) (150... 46 4e-04
TONB_VIBCH (O52042) TonB protein 45 6e-04
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 453 bits (1165), Expect = e-126
Identities = 293/851 (34%), Positives = 438/851 (51%), Gaps = 69/851 (8%)
Query: 5 TRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFTNLGIV 62
+R WV+ LK K + F FH ++E + +K+K +++D GGE+ + Y ++ GI
Sbjct: 511 SRKLWVYILKTKDQVFQVFQKFHALVERETGRKLKRLRSDNGGEYTSREFEEYCSSHGIR 570
Query: 63 HRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLD 122
H T P T NG+ ER +R IVE ++L A + +W A T YLINR P+V L
Sbjct: 571 HEKTVPGTPQHNGVAERMNRTIVEKVRSMLRMAKLPKSFWGEAVQTACYLINRSPSVPLA 630
Query: 123 NESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD 182
E P L++FGC + ++ KL S C+F+GY Y+ D
Sbjct: 631 FEIPERVWTNKEVSYSHLKVFGCRAFAHVPKEQRTKLDDKSIPCIFIGYGDEEFGYRLWD 690
Query: 183 PTGRLYI-SKDVLFNEFRFPYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPH 241
P + I S+DV+F E ++ + + + P T+ S
Sbjct: 691 PVKKKVIRSRDVVFRE--------------SEVRTAADMSEKVKNGIIPNFVTIPST--- 733
Query: 242 SPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPIV 301
S++P S ST T V + P Q D V +
Sbjct: 734 --------SNNPTSAEST------TDEVSEQGEQP-----GEVIEQGEQLDEG--VEEVE 772
Query: 302 NP---HNTHSMVTRAKDGIVKPRLVPT----LLLSQAEPTSAKLALKDPK---WYSAMQD 351
+P H + R++ V+ R P+ L+ EP S K L P+ AMQ+
Sbjct: 773 HPTQGEEQHQPLRRSERPRVESRRYPSTEYVLISDDREPESLKEVLSHPEKNQLMKAMQE 832
Query: 352 EFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYK 411
E +L N T+ LV LP ++ + CKWVF++K++ D + RYKARLV KG+ Q +G D+
Sbjct: 833 EMESLQKNGTYKLVELPKGKRPLKCKWVFKLKKDGDCKLVRYKARLVVKGFEQKKGIDFD 892
Query: 412 ETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFE-AKDRSL 470
E FSPVVK ++R ILSLA S + QLDV AFL+G LEEE+YM QP GFE A + +
Sbjct: 893 EIFSPVVKMTSIRTILSLAASLDLEVEQLDVKTAFLHGDLEEEIYMEQPEGFEVAGKKHM 952
Query: 471 VCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSL-FTLYSSGCSVYVLVYVDDI 529
VC+LNK+LYGLKQAPR W+ + + + + + +P + F +S + +L+YVDD+
Sbjct: 953 VCKLNKSLYGLKQAPRQWYMKFDSFMKSQTYLKTYSDPCVYFKRFSENNFIILLLYVDDM 1012
Query: 530 IITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQ-VQHLHDKSLLLTQSKYIKDLLER 588
+I G ++ L L+ F +K LG LG++ V+ + L L+Q KYI+ +LER
Sbjct: 1013 LIVGKDKGLIAKLKGDLSKSFDMKDLGPAQQILGMKIVRERTSRKLWLSQEKYIERVLER 1072
Query: 589 ASMAYAKGISTPMISNQKLTK--------HGANYMADPSLYRSIVGALQYVTI-TRPEIS 639
+M AK +STP+ + KL+K N P Y S VG+L Y + TRP+I+
Sbjct: 1073 FNMKNAKPVSTPLAGHLKLSKKMCPTTVEEKGNMAKVP--YSSAVGSLMYAMVCTRPDIA 1130
Query: 640 YSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAAD 699
++V V +FL P +EHW AVK ILRYL+GT L C L + DAD A D
Sbjct: 1131 HAVGVVSRFLENPGKEHWEAVKWILRYLRGTTGDCL----CFGGSDPILKGYTDADMAGD 1186
Query: 700 PDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHI 759
D+ +S++G L +SW SK Q VA S+TEAEY T E++W++ LQEL +
Sbjct: 1187 IDNRKSSTGYLFTFSGGAISWQSKLQKCVALSTTEAEYIAATETGKEMIWLKRFLQELGL 1246
Query: 760 PHKTPLILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDI 819
K ++ CD+ S + L+ N + H RTKH+++ ++RE V ++SL + + + + D+
Sbjct: 1247 HQKEYVVYCDSQSAIDLSKNSMYHARTKHIDVRYHWIREMVDDESLKVLKISTNENPADM 1306
Query: 820 LTKALSPLRFE 830
LTK + +FE
Sbjct: 1307 LTKVVPRNKFE 1317
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 338 bits (868), Expect = 3e-92
Identities = 190/500 (38%), Positives = 293/500 (58%), Gaps = 11/500 (2%)
Query: 345 WYSAMQDEFNALHSNQTWTLVPLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQ 404
W A+ E NA N TWT+ P ++ + +WVF +K N G+ RYKARLVA+G++Q
Sbjct: 906 WEEAINTELNAHKINNTWTITKRPENKNIVDSRWVFSVKYNELGNPIRYKARLVARGFTQ 965
Query: 405 VEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFE 464
DY+ETF+PV + + R ILSL + +HQ+DV AFLNG L+EE+YM P G
Sbjct: 966 KYQIDYEETFAPVARISSFRFILSLVIQYNLKVHQMDVKTAFLNGTLKEEIYMRLPQGIS 1025
Query: 465 AKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGC--SVYV 522
+ VC+LNKA+YGLKQA R WFE + AL + F S + ++ L ++YV
Sbjct: 1026 CNSDN-VCKLNKAIYGLKQAARCWFEVFEQALKECEFVNSSVDRCIYILDKGNINENIYV 1084
Query: 523 LVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQSKYI 582
L+YVDD++I + + + L +F + L + +F+G++++ DK + L+QS Y+
Sbjct: 1085 LLYVDDVVIATGDMTRMNNFKRYLMEKFRMTDLNEIKHFIGIRIEMQEDK-IYLSQSAYV 1143
Query: 583 KDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSL-YRSIVGALQYVTI-TRPEISY 640
K +L + +M +STP+ S K+ N D + RS++G L Y+ + TRP+++
Sbjct: 1144 KKILSKFNMENCNAVSTPLPS--KINYELLNSDEDCNTPCRSLIGCLMYIMLCTRPDLTT 1201
Query: 641 SVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADP 700
+VN + ++ S+ E W +KR+LRYLKGT+D L K +L +I + D+DWA
Sbjct: 1202 AVNILSRYSSKNNSELWQNLKRVLRYLKGTIDMKLIFKK-NLAFENKIIGYVDSDWAGSE 1260
Query: 701 DDHRSTSGSLV-FLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHI 759
D +ST+G L NL+ W +K+Q+ VA SSTEAEY L V E LW++ LL ++I
Sbjct: 1261 IDRKSTTGYLFKMFDFNLICWNTKRQNSVAASSTEAEYMALFEAVREALWLKFLLTSINI 1320
Query: 760 PHKTPL-ILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDD 818
+ P+ I DN +++ +NP H R KH+++ F RE+V N + ++++P+ +Q D
Sbjct: 1321 KLENPIKIYEDNQGCISIANNPSCHKRAKHIDIKYHFAREQVQNNVICLEYIPTENQLAD 1380
Query: 819 ILTKALSPLRFEDLRTKLNV 838
I TK L RF +LR KL +
Sbjct: 1381 IFTKPLPAARFVELRDKLGL 1400
Score = 89.4 bits (220), Expect = 4e-17
Identities = 58/199 (29%), Positives = 96/199 (48%), Gaps = 8/199 (4%)
Query: 4 YTRYTWVFPLKKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFTNLGI 61
+T Y + +K KSD ++ F +F E N KV + D G E+ + + GI
Sbjct: 510 FTHYCVTYLIKYKSDVFSMFQDFVAKSEAHFNLKVVYLYIDNGREYLSNEMRQFCVKKGI 569
Query: 62 VHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPT--V 119
+ T PHT NG+ ER R I E +++ A + +W A LT YLINR+P+ +
Sbjct: 570 SYHLTVPHTPQLNGVSERMIRTITEKARTMVSGAKLDKSFWGEAVLTATYLINRIPSRAL 629
Query: 120 VLDNESPFTKLFQVSFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYK 179
V +++P+ +K LR+FG Y +++ K S + +F+GY +K
Sbjct: 630 VDSSKTPYEMWHNKKPYLKHLRVFGATVYVHIKNKQG-KFDDKSFKSIFVGYEP--NGFK 686
Query: 180 CLDPTGRLYI-SKDVLFNE 197
D +I ++DV+ +E
Sbjct: 687 LWDAVNEKFIVARDVVVDE 705
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 208 bits (530), Expect = 4e-53
Identities = 111/230 (48%), Positives = 156/230 (67%), Gaps = 5/230 (2%)
Query: 520 VYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSLLLTQS 579
+Y+L+YVDDI++TG+S +L LI +L++ FS+K LG + YFLG+Q++ H L L+Q+
Sbjct: 1 MYLLLYVDDILLTGSSNTLLNMLIFQLSSTFSMKDLGPVHYFLGIQIK-THPSGLFLSQT 59
Query: 580 KYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTITRPEIS 639
KY + +L A M K +STP+ + A Y DPS +RSIVGALQY+T+TRP+IS
Sbjct: 60 KYAEQILNNAGMLDCKPMSTPLPLKLNSSVSTAKY-PDPSDFRSIVGALQYLTLTRPDIS 118
Query: 640 YSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAAD 699
Y+VN VCQ + +P + +KR+LRY+KGT+ GL++ S L++ AFCD+DWA
Sbjct: 119 YAVNIVCQRMHEPTLADFDLLKRVLRYVKGTIFHGLYIHKNS---KLNVQAFCDSDWAGC 175
Query: 700 PDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLW 749
RST+G FLG N++SW +K+Q V+RSSTE EYR LA T AEL W
Sbjct: 176 TSTRRSTTGFCTFLGCNIISWSAKRQPTVSRSSTETEYRALALTAAELTW 225
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 162 bits (411), Expect = 3e-39
Identities = 103/311 (33%), Positives = 163/311 (52%), Gaps = 6/311 (1%)
Query: 440 LDVNNAFLNGKLEEEVYMVQPPGF-EAKDRSLVCRLNKALYGLKQAPRAWFERLKAALLK 498
+DV+ AFLN ++E +Y+ QPPGF ++ V L +YGLKQAP W E + L K
Sbjct: 1 MDVDTAFLNSTMDEPIYVKQPPGFVNERNPDYVWELYGGMYGLKQAPLLWNEHINNTLKK 60
Query: 499 FGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTL 558
GF L+ +S +Y+ VYVDD+++ S + + +L +S+K LG +
Sbjct: 61 IGFCRHEGEHGLYFRSTSDGPIYIGVYVDDLLVAAPSPKIYDRVKQELTKLYSMKDLGKV 120
Query: 559 DYFLGVQVQHLHDKSLLLTQSKYIKDLLERASMAYAKGISTPMISNQKLTKHGANYMADP 618
D FLG+ + + + L+ YI + + K TP+ +++ L + + ++ D
Sbjct: 121 DKFLGLNIHQSTNGDITLSLQDYIAKAASESEINTFKLTQTPLCNSKPLFETTSPHLKDI 180
Query: 619 SLYRSIVGALQYVTIT-RPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHL 677
+ Y+SIVG L + T RP+ISY V+ + +FL +P H + +R+LRYL T L
Sbjct: 181 TPYQSIVGQLLFCANTGRPDISYPVSLLSRFLREPRAIHLESARRVLRYLYTTRSMCLKY 240
Query: 678 KPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKK-QSLVARSSTEAE 736
+ S ++L +CDA A D ST G + L V+W SKK + ++ STEAE
Sbjct: 241 RSGS---QVALTVYCDASHGAIHDLPHSTGGYVTLLAGAPVTWSSKKLKGVIPVPSTEAE 297
Query: 737 YRNLANTVAEL 747
Y + TV E+
Sbjct: 298 YITASETVMEI 308
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 139 bits (350), Expect = 3e-32
Identities = 125/457 (27%), Positives = 214/457 (46%), Gaps = 30/457 (6%)
Query: 393 YKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLE 452
YKAR+V +G +Q Y + + ++I L +A ++ + LD+N+AFL KLE
Sbjct: 1337 YKARIVCRGDTQSPD-TYSVITTESLNHNHIKIFLMIANNRNMFMKTLDINHAFLYAKLE 1395
Query: 453 EEVYMVQPPGFEAKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFT 512
EE+Y+ P DR V +LNKALYGLKQ+P+ W + L+ L G + P L+
Sbjct: 1396 EEIYIPHP-----HDRRCVVKLNKALYGLKQSPKEWNDHLRQYLNGIGLKDNSYTPGLY- 1449
Query: 513 LYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTL-DYFLGVQV--QHL 569
+ ++ + VYVDD +I ++ L I KL + F LK GTL D L + L
Sbjct: 1450 -QTEDKNLMIAVYVDDCVIAASNEQRLDEFINKLKSNFELKITGTLIDDVLDTDILGMDL 1508
Query: 570 HDKSLLLTQSKYIKDLLERASMAYAKGI------STPMISNQKL-TKHGANYMADPSL-- 620
L T +K + R Y + + S P +S K+ K M++
Sbjct: 1509 VYNKRLGTIDLTLKSFINRMDKKYNEELKKIRKSSIPHMSTYKIDPKKDVLQMSEEEFRQ 1568
Query: 621 ----YRSIVGALQYVT-ITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGL 675
+ ++G L YV R +I ++V KV + ++ P E + + +I++YL D G+
Sbjct: 1569 GVLKLQQLLGELNYVRHKCRYDIEFAVKKVARLVNYPHERVFYMIYKIIQYLVRYKDIGI 1628
Query: 676 HL-KPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTE 734
H + C ++ +IA DA ++ D +S G +++ G N+ + S K + SSTE
Sbjct: 1629 HYDRDC--NKDKKVIAITDASVGSE-YDAQSRIGVILWYGMNIFNVYSNKSTNRCVSSTE 1685
Query: 735 AEYRNLANTVAELLWVQSLLQELHIPHKTPLI-LCDNMSTVALTHNPVLHTRTKHMELDI 793
AE + A+ ++ L+EL ++ + D+ + + + K +
Sbjct: 1686 AELHAIYEGYADSETLKVTLKELGEGDNNDIVMITDSKPAIQGLNRSYQQPKEKFTWIKT 1745
Query: 794 FFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRFE 830
++EK+ KS+ + + D+LTK +S F+
Sbjct: 1746 EIIKEKIKEKSIKLLKITGKGNIADLLTKPVSASDFK 1782
Score = 50.4 bits (119), Expect = 2e-05
Identities = 27/89 (30%), Positives = 46/89 (51%), Gaps = 2/89 (2%)
Query: 30 IELQLNKKVKVVQTDGGGEFK--PLTPYFTNLGIVHRFTCPHTHHQNGIVERKHRHIVEM 87
+E Q ++KV+ + +D G EF + YF + GI H T H NG ER R I+
Sbjct: 682 VETQFDRKVREINSDRGTEFTNDQIEEYFISKGIHHILTSTQDHAANGRAERYIRTIITD 741
Query: 88 GLALLAHAHISLEYWDHAFLTVVYLINRL 116
LL +++ +++W++A + + N L
Sbjct: 742 ATTLLRQSNLRVKFWEYAVTSATNIRNYL 770
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 127 bits (320), Expect = 1e-28
Identities = 140/575 (24%), Positives = 253/575 (43%), Gaps = 48/575 (8%)
Query: 290 STDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAM 349
+T + S+ P + H + +KP + T L T K + K+ A
Sbjct: 770 NTKNMRSLEPPRSKKRIHLIAAVKAVKSIKP--IRTTLRYDEAITYNKDIKEKEKYIQAY 827
Query: 350 QDEFNALHSNQTWTLVPLPSHRKAIGCKWV----FRIKENADGSVNRYKARLVAKGYSQV 405
E N L +TW RK I K V F DG+ +KAR VA+G Q
Sbjct: 828 HKEVNQLLKMKTWD-TDRYYDRKEIDPKRVINSMFIFNRKRDGT---HKARFVARGDIQ- 882
Query: 406 EGFDYKETFSPVVKPITVR-----IILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQP 460
+ +T+ P ++ TV LSLA+ ++I QLD+++A+L ++EE+Y+ P
Sbjct: 883 ----HPDTYDPGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEELYIRPP 938
Query: 461 PGFEAKDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSL--FTLYSSGC 518
P D+ + RL K+LYGLKQ+ W+E +K+ L+K C + ++
Sbjct: 939 PHLGMNDKLI--RLKKSLYGLKQSGANWYETIKSYLIK-----QCGMEEVRGWSCVFKNS 991
Query: 519 SVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLK--QLGTLDY-----FLGVQVQHLHD 571
V + ++VDD+I+ L + +I L ++ K LG D LG+++++
Sbjct: 992 QVTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRG 1051
Query: 572 KSLLLTQSKYIKDLLERASMAY---AKGISTP-----MISNQKLTKHGANYMADPSLYRS 623
K + L + + + + ++ + +S P I Q+L +Y +
Sbjct: 1052 KYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQK 1111
Query: 624 IVGALQYVTIT-RPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSL 682
++G YV R ++ Y +N + Q + P ++ +++++ T D L
Sbjct: 1112 LIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQLIWHKSKP 1171
Query: 683 HQPLS-LIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLA 741
+P + L+ DA + P ++S G++ L ++ S K SL S+TEAE ++
Sbjct: 1172 VKPTNKLVVISDASYGNQPY-YKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIHAIS 1230
Query: 742 NTVAELLWVQSLLQELHIPHKTPLILCDNMSTVA-LTHNPVLHTRTKHMELDIFFVREKV 800
+V L + L+QEL+ T +L D+ ST++ + N R + +R++V
Sbjct: 1231 ESVPLLNNLSHLVQELNKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTKAMRLRDEV 1290
Query: 801 LNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTK 835
L + ++ + D++TK L F+ L K
Sbjct: 1291 SGNHLHVCYIETKKNIADVMTKPLPIKTFKLLTNK 1325
Score = 38.9 bits (89), Expect = 0.058
Identities = 25/89 (28%), Positives = 43/89 (48%), Gaps = 10/89 (11%)
Query: 5 TRYTWVFPLKKKS-----DTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFT 57
T++ WV+PL + D +TT + F I+ Q V V+Q D G E+ + L +
Sbjct: 268 TKFRWVYPLHDRREDSILDVFTTILAF---IKNQFQASVLVIQMDRGSEYTNRTLHKFLE 324
Query: 58 NLGIVHRFTCPHTHHQNGIVERKHRHIVE 86
GI +T +G+ ER +R +++
Sbjct: 325 KNGITPCYTTTADSRAHGVAERLNRTLLD 353
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 125 bits (314), Expect = 5e-28
Identities = 138/570 (24%), Positives = 248/570 (43%), Gaps = 38/570 (6%)
Query: 290 STDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAM 349
+T + S+ P + H + +KP + T L T K + K+ A
Sbjct: 770 NTKNMRSLEPPRSKKRIHLIAAVKAVKSIKP--IRTTLRYDEAITYNKDIKEKEKYIEAY 827
Query: 350 QDEFNALHSNQTWTLVPLPSHRKAIGCKWV----FRIKENADGSVNRYKARLVAKGYSQV 405
E N L +TW RK I K V F DG+ +KAR VA+G Q
Sbjct: 828 HKEVNQLLKMKTWDTDKYYD-RKEIDPKRVINSMFIFNRKRDGT---HKARFVARGDIQH 883
Query: 406 EGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEA 465
S V + LSLA+ ++I QLD+++A+L ++EE+Y+ PP
Sbjct: 884 PDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGM 943
Query: 466 KDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSL--FTLYSSGCSVYVL 523
D+ + RL K+LYGLKQ+ W+E +K+ L+K C + ++ V +
Sbjct: 944 NDK--LIRLKKSLYGLKQSGANWYETIKSYLIK-----QCGMEEVRGWSCVFENSQVTIC 996
Query: 524 VYVDDIIITGNSLPVLQHLIAKLNAEFSLK--QLGTLDY-----FLGVQVQHLHDKSLLL 576
++VDD+++ +L + +I KL ++ K LG D LG+++++ K + L
Sbjct: 997 LFVDDMVLFSKNLNSNKRIIDKLKMQYDTKIINLGESDEEIQYDILGLEIKYQRGKYMKL 1056
Query: 577 TQSKYIKDLLERASMAY---AKGISTP-----MISNQKLTKHGANYMADPSLYRSIVGAL 628
+ + + + ++ + +S P I Q+L +Y + ++G
Sbjct: 1057 GMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKLIGLA 1116
Query: 629 QYVTIT-RPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLS 687
YV R ++ Y +N + Q + P ++ +++++ T D L +P +
Sbjct: 1117 SYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQLIWHKSKPVKPTN 1176
Query: 688 -LIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAE 746
L+ DA + P ++S G++ L ++ S K SL S+TEAE ++ +V
Sbjct: 1177 KLVVISDASYGNQP-YYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPL 1235
Query: 747 LLWVQSLLQELHIPHKTPLILCDNMSTVA-LTHNPVLHTRTKHMELDIFFVREKVLNKSL 805
L + L+QEL T +L D+ ST++ + N R + +R++V L
Sbjct: 1236 LNNLSYLIQELDKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTKAMRLRDEVSGNHL 1295
Query: 806 IIQHVPSLDQRDDILTKALSPLRFEDLRTK 835
+ ++ + D++TK L F+ L K
Sbjct: 1296 HVCYIETKKNIADVMTKPLPIKTFKLLTNK 1325
Score = 38.9 bits (89), Expect = 0.058
Identities = 25/89 (28%), Positives = 43/89 (48%), Gaps = 10/89 (11%)
Query: 5 TRYTWVFPLKKKS-----DTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFT 57
T++ WV+PL + D +TT + F I+ Q V V+Q D G E+ + L +
Sbjct: 268 TKFRWVYPLHDRREDSILDVFTTILAF---IKNQFQASVLVIQMDRGSEYTNRTLHKFLE 324
Query: 58 NLGIVHRFTCPHTHHQNGIVERKHRHIVE 86
GI +T +G+ ER +R +++
Sbjct: 325 KNGITPCYTTTADSRAHGVAERLNRTLLD 353
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 124 bits (312), Expect = 8e-28
Identities = 137/570 (24%), Positives = 249/570 (43%), Gaps = 38/570 (6%)
Query: 290 STDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAM 349
+T + S+ P + H + +KP + T L T K + K+ A
Sbjct: 1197 NTKNMRSLEPPRSKKRIHLIAAVKAVKSIKP--IRTTLRYDEAITYNKDIKEKEKYIEAY 1254
Query: 350 QDEFNALHSNQTWTLVPLPSHRKAIGCKWV----FRIKENADGSVNRYKARLVAKGYSQV 405
E N L +TW RK I K V F + DG+ +KAR VA+G Q
Sbjct: 1255 HKEVNQLLKMKTWDTDEYYD-RKEIDPKRVINSMFIFNKKRDGT---HKARFVARGDIQH 1310
Query: 406 EGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEA 465
S V + LSLA+ ++I QLD+++A+L ++EE+Y+ PP
Sbjct: 1311 PDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGM 1370
Query: 466 KDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSL--FTLYSSGCSVYVL 523
D+ + RL K+LYGLKQ+ W+E +K+ L++ C + ++ V +
Sbjct: 1371 NDK--LIRLKKSLYGLKQSGANWYETIKSYLIQ-----QCGMEEVRGWSCVFKNSQVTIC 1423
Query: 524 VYVDDIIITGNSLPVLQHLIAKLNAEFSLK--QLGTLDY-----FLGVQVQHLHDKSLLL 576
++VDD+++ +L + +I KL ++ K LG D LG+++++ K + L
Sbjct: 1424 LFVDDMVLFSKNLNSNKRIIEKLKMQYDTKIINLGESDEEIQYDILGLEIKYQRGKYMKL 1483
Query: 577 TQSKYIKDLLERASMAY---AKGISTP-----MISNQKLTKHGANYMADPSLYRSIVGAL 628
+ + + + ++ + +S P I Q+L +Y + ++G
Sbjct: 1484 GMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQELELEEDDYKMKVHEMQKLIGLA 1543
Query: 629 QYVTIT-RPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLS 687
YV R ++ Y +N + Q + P ++ +++++ T D L +P +
Sbjct: 1544 SYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWNTRDKQLIWHKSKPVKPTN 1603
Query: 688 -LIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAE 746
L+ DA + P ++S G++ L ++ S K SL S+TEAE ++ +V
Sbjct: 1604 KLVVISDASYGNQP-YYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPL 1662
Query: 747 LLWVQSLLQELHIPHKTPLILCDNMSTVA-LTHNPVLHTRTKHMELDIFFVREKVLNKSL 805
L + L+QEL T +L D+ ST++ + N R + +R++V L
Sbjct: 1663 LNNLSYLIQELDKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTKAMRLRDEVSGNHL 1722
Query: 806 IIQHVPSLDQRDDILTKALSPLRFEDLRTK 835
+ ++ + D++TK L F+ L K
Sbjct: 1723 HVCYIETKKNIADVMTKPLPIKTFKLLTNK 1752
Score = 38.9 bits (89), Expect = 0.058
Identities = 25/89 (28%), Positives = 43/89 (48%), Gaps = 10/89 (11%)
Query: 5 TRYTWVFPLKKKS-----DTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFT 57
T++ WV+PL + D +TT + F I+ Q V V+Q D G E+ + L +
Sbjct: 695 TKFRWVYPLHDRREDSILDVFTTILAF---IKNQFQASVLVIQMDRGSEYTNRTLHKFLE 751
Query: 58 NLGIVHRFTCPHTHHQNGIVERKHRHIVE 86
GI +T +G+ ER +R +++
Sbjct: 752 KNGITPCYTTTADSRAHGVAERLNRTLLD 780
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 124 bits (310), Expect = 1e-27
Identities = 127/494 (25%), Positives = 224/494 (44%), Gaps = 51/494 (10%)
Query: 371 RKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRIILSLA 430
+K I ++F K DG+ +KAR VA+G Q + S V + LS+A
Sbjct: 1296 KKVINSMFIFNKKR--DGT---HKARFVARGDIQHPDTYDSDMQSNTVHHYALMTSLSIA 1350
Query: 431 VSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEAKDRSLVCRLNKALYGLKQAPRAWFE 490
+ ++I QLD+++A+L ++EE+Y+ PP D+ L RL K+LYGLKQ+ W+E
Sbjct: 1351 LDNDYYITQLDISSAYLYADIKEELYIRPPPHLGLNDKLL--RLRKSLYGLKQSGANWYE 1408
Query: 491 RLKAALLKFGFTASCCNPSLFTLYS---SGCSVYVLVYVDDIIITGNSLPVLQHLIAKLN 547
+K+ L+ +CC+ +S V + ++VDD+I+ L + +I L
Sbjct: 1409 TIKSYLI------NCCDMQEVRGWSCVFKNSQVTICLFVDDMILFSKDLNANKKIITTLK 1462
Query: 548 AEFSLK--QLGTLDY-----FLGVQVQHLHDKSLLLTQSKYIKDLLERASMAY------- 593
++ K LG D LG+++++ K + L K + + L + ++
Sbjct: 1463 KQYDTKIINLGESDNEIQYDILGLEIKYQRSKYMKLGMEKSLTEKLPKLNVPLNPKGKKL 1522
Query: 594 -AKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTIT-RPEISYSVNKVCQFLSQ 651
A G I +L Y + ++G YV R ++ Y +N + Q +
Sbjct: 1523 RAPGQPGHYIDQDELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILF 1582
Query: 652 PLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQP-LSLIAFCDADWAADPDDHRSTSGSL 710
P + +++++ T D L +P L+A DA + P ++S G++
Sbjct: 1583 PSRQVLDMTYELIQFMWDTRDKQLIWHKNKPTKPDNKLVAISDASYGNQP-YYKSQIGNI 1641
Query: 711 VFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPLI---L 767
L ++ S K SL S+TEAE ++ + L + L+QEL +K P+I L
Sbjct: 1642 FLLNGKVIGGKSTKASLTCTSTTEAEIHAVSEAIPLLNNLSHLVQEL---NKKPIIKGLL 1698
Query: 768 CDNMSTVAL---THNPVLHTR---TKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILT 821
D+ ST+++ T+ R TK M L R++V +L + ++ + D++T
Sbjct: 1699 TDSRSTISIIKSTNEEKFRNRFFGTKAMRL-----RDEVSGNNLYVYYIETKKNIADVMT 1753
Query: 822 KALSPLRFEDLRTK 835
K L F+ L K
Sbjct: 1754 KPLPIKTFKLLTNK 1767
Score = 52.4 bits (124), Expect = 5e-06
Identities = 34/112 (30%), Positives = 55/112 (48%), Gaps = 6/112 (5%)
Query: 5 TRYTWVFPL--KKKSDTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFTNLG 60
TR+ WV+PL +++ F + I+ Q N +V V+Q D G E+ K L +FTN G
Sbjct: 691 TRFQWVYPLHDRREESILNVFTSILAFIKNQFNARVLVIQMDRGSEYTNKTLHKFFTNRG 750
Query: 61 IVHRFTCPHTHHQNGIVERKHRHIVEMGLALLAHAHISLEYWDHA--FLTVV 110
I +T +G+ ER +R ++ LL + + W A F T++
Sbjct: 751 ITACYTTTADSRAHGVAERLNRTLLNDCRTLLHCSGLPNHLWFSAVEFSTII 802
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 122 bits (307), Expect = 3e-27
Identities = 144/578 (24%), Positives = 251/578 (42%), Gaps = 54/578 (9%)
Query: 290 STDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAM 349
+T + S+ P + H + +KP + T L T K + K+ A
Sbjct: 770 NTKNMRSLEPPRSKKRIHLIAAVKAVKSIKP--IRTTLRYDEAITYNKDIKEKEKYIEAY 827
Query: 350 QDEFNALHSNQTWTLVPLPSHRKAIGCKWV----FRIKENADGSVNRYKARLVAKGYSQV 405
E N L +TW RK I K V F + DG+ +KAR VA+G Q
Sbjct: 828 HKEVNQLLKMKTWDTDEYYD-RKEIDPKRVINSMFIFNKKRDGT---HKARFVARGDIQH 883
Query: 406 EGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEA 465
S V + LSLA+ ++I QLD+++A+L ++EE+Y+ PP
Sbjct: 884 PDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGM 943
Query: 466 KDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSL--FTLYSSGCSVYVL 523
D+ + RL K+LYGLKQ+ W+E +K+ L+K C + ++ V +
Sbjct: 944 NDK--LIRLKKSLYGLKQSGANWYETIKSYLIK-----QCGMEEVRGWSCVFKNSQVTIC 996
Query: 524 VYVDDIIITGNSLPVLQHLIAKLNAEFSLK--QLGTLDY-----FLGVQVQHLHDKSLLL 576
++VDD+I+ L + +I L ++ K LG D LG+++++ K + L
Sbjct: 997 LFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGKYMKL 1056
Query: 577 TQSKYIKDLLERASMAY---AKGISTP-----MISNQKLTKHGANYMADPSLYRSIVGAL 628
+ + + + ++ + +S P I +L Y + ++G
Sbjct: 1057 GMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHEMQKLIGLA 1116
Query: 629 QYVTIT-RPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQP-L 686
YV R ++ Y +N + Q + P + +++++ T D L +P
Sbjct: 1117 SYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPTEPDN 1176
Query: 687 SLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAE 746
L+A DA + P ++S G++ L ++ S K SL S+TEAE ++ +V
Sbjct: 1177 KLVAISDASYGNQP-YYKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPL 1235
Query: 747 LLWVQSLLQELHIPHKTPLI---LCDNMSTVAL---THNPVLHTR---TKHMELDIFFVR 797
L + L+QEL +K P+I L D+ ST+++ T+ R TK M L R
Sbjct: 1236 LNNLSYLIQEL---NKKPIIKGLLTDSRSTISIIKSTNEEKFRNRFFGTKAMRL-----R 1287
Query: 798 EKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTK 835
++V +L + ++ + D++TK L F+ L K
Sbjct: 1288 DEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKLLTNK 1325
Score = 38.9 bits (89), Expect = 0.058
Identities = 25/89 (28%), Positives = 43/89 (48%), Gaps = 10/89 (11%)
Query: 5 TRYTWVFPLKKKS-----DTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFT 57
T++ WV+PL + D +TT + F I+ Q V V+Q D G E+ + L +
Sbjct: 268 TKFRWVYPLHDRREDSILDVFTTILAF---IKNQFQASVLVIQMDRGSEYTNRTLHKFLE 324
Query: 58 NLGIVHRFTCPHTHHQNGIVERKHRHIVE 86
GI +T +G+ ER +R +++
Sbjct: 325 KNGITPCYTTTADSRAHGVAERLNRTLLD 353
>M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820
(ORF170)
Length = 170
Score = 122 bits (305), Expect = 5e-27
Identities = 63/125 (50%), Positives = 88/125 (70%), Gaps = 3/125 (2%)
Query: 309 MVTRAKDGIVK--PRLVPTLLLS-QAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLV 365
M+TR+K GI K P+ T+ + + EP S ALKDP W AMQ+E +AL N+TW LV
Sbjct: 1 MLTRSKAGINKLNPKYSLTITTTIKKEPKSVIFALKDPGWCQAMQEELDALSRNKTWILV 60
Query: 366 PLPSHRKAIGCKWVFRIKENADGSVNRYKARLVAKGYSQVEGFDYKETFSPVVKPITVRI 425
P P ++ +GCKWVF+ K ++DG+++R KARLVAKG+ Q EG + ET+SPVV+ T+R
Sbjct: 61 PPPVNQNILGCKWVFKTKLHSDGTLDRLKARLVAKGFHQEEGIYFVETYSPVVRTATIRT 120
Query: 426 ILSLA 430
IL++A
Sbjct: 121 ILNVA 125
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 121 bits (303), Expect = 9e-27
Identities = 138/570 (24%), Positives = 244/570 (42%), Gaps = 38/570 (6%)
Query: 290 STDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAM 349
+T + S+ P + H + +KP + T L T K + K+ A
Sbjct: 770 NTKNMRSLEPPRSKKRIHLIAAVKAVKSIKP--IRTTLRYDEAITYNKDIKEKEKYIEAY 827
Query: 350 QDEFNALHSNQTWTLVPLPSHRKAIGCKWV----FRIKENADGSVNRYKARLVAKGYSQV 405
E N L TW RK I K V F DG+ +KAR VA+G Q
Sbjct: 828 HKEVNQLLKMNTWDTDKYYD-RKEIDPKRVINSMFIFNRKRDGT---HKARFVARGDIQH 883
Query: 406 EGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEA 465
S V + LSLA+ ++I QLD+++A+L ++EE+Y+ PP
Sbjct: 884 PDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGM 943
Query: 466 KDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSL--FTLYSSGCSVYVL 523
D+ + RL K+LYGLKQ+ W+E +K+ L+K C + ++ V +
Sbjct: 944 NDKLI--RLKKSLYGLKQSGANWYETIKSYLIK-----QCGMEEVRGWSCVFKNSQVTIC 996
Query: 524 VYVDDIIITGNSLPVLQHLIAKLNAEFSLK--QLGTLDY-----FLGVQVQHLHDKSLLL 576
++VDD+I+ L + +I L ++ K LG D LG+++++ K + L
Sbjct: 997 LFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGKYMKL 1056
Query: 577 TQSKYIKDLLERASMAY---AKGISTP-----MISNQKLTKHGANYMADPSLYRSIVGAL 628
+ + + + ++ + +S P I +L Y + ++G
Sbjct: 1057 GMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHEMQKLIGLA 1116
Query: 629 QYVTIT-RPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLS 687
YV R ++ Y +N + Q + P + +++++ T D L +P +
Sbjct: 1117 SYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPTEPDN 1176
Query: 688 -LIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAE 746
L+A DA + P ++S G++ L ++ S K SL S+TEAE ++ +V
Sbjct: 1177 KLVAISDASYGNQPY-YKSQIGNIYLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPL 1235
Query: 747 LLWVQSLLQELHIPHKTPLILCDNMSTVA-LTHNPVLHTRTKHMELDIFFVREKVLNKSL 805
L + L+QEL+ T +L D+ ST++ + N R + +R++V L
Sbjct: 1236 LNNLSHLVQELNKKPITKGLLTDSKSTISIIISNNEEKFRNRFFGTKAMRLRDEVSGNHL 1295
Query: 806 IIQHVPSLDQRDDILTKALSPLRFEDLRTK 835
+ ++ + D++TK L F+ L K
Sbjct: 1296 HVCYIETKKNIADVMTKPLPIKTFKLLTNK 1325
Score = 38.9 bits (89), Expect = 0.058
Identities = 25/89 (28%), Positives = 43/89 (48%), Gaps = 10/89 (11%)
Query: 5 TRYTWVFPLKKKS-----DTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFT 57
T++ WV+PL + D +TT + F I+ Q V V+Q D G E+ + L +
Sbjct: 268 TKFRWVYPLHDRREDSILDVFTTILAF---IKNQFQASVLVIQMDRGSEYTNRTLHKFLE 324
Query: 58 NLGIVHRFTCPHTHHQNGIVERKHRHIVE 86
GI +T +G+ ER +R +++
Sbjct: 325 KNGITPCYTTTADSRAHGVAERLNRTLLD 353
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 119 bits (299), Expect = 3e-26
Identities = 143/578 (24%), Positives = 250/578 (42%), Gaps = 54/578 (9%)
Query: 290 STDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAM 349
+T + S+ P + H + +KP + T L T K + K+ A
Sbjct: 1197 NTKNMRSLEPPRSKKRIHLIAAVKAVKSIKP--IRTTLRYDEAITYNKDIKEKEKYIEAY 1254
Query: 350 QDEFNALHSNQTWTLVPLPSHRKAIGCKWV----FRIKENADGSVNRYKARLVAKGYSQV 405
E N L +TW RK I K V F + DG+ +KAR VA+G Q
Sbjct: 1255 HKEVNQLLKMKTWDTDEYYD-RKEIDPKRVINSMFIFNKKRDGT---HKARFVARGDIQH 1310
Query: 406 EGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFEA 465
S V + LSLA+ ++I QLD+++A+L ++EE+Y+ PP
Sbjct: 1311 PDTYDTGMQSNTVHHYALMTSLSLALDNNYYITQLDISSAYLYADIKEELYIRPPPHLGM 1370
Query: 466 KDRSLVCRLNKALYGLKQAPRAWFERLKAALLKFGFTASCCNPSL--FTLYSSGCSVYVL 523
D+ + RL K+ YGLKQ+ W+E +K+ L+K C + ++ V +
Sbjct: 1371 NDK--LIRLKKSHYGLKQSGANWYETIKSYLIK-----QCGMEEVRGWSCVFKNSQVTIC 1423
Query: 524 VYVDDIIITGNSLPVLQHLIAKLNAEFSLK--QLGTLDY-----FLGVQVQHLHDKSLLL 576
++VDD+I+ L + +I L ++ K LG D LG+++++ K + L
Sbjct: 1424 LFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRGKYMKL 1483
Query: 577 TQSKYIKDLLERASMAY---AKGISTP-----MISNQKLTKHGANYMADPSLYRSIVGAL 628
+ + + + ++ + +S P I +L Y + ++G
Sbjct: 1484 GMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQDELEIDEDEYKEKVHEMQKLIGLA 1543
Query: 629 QYVTIT-RPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQP-L 686
YV R ++ Y +N + Q + P + +++++ T D L +P
Sbjct: 1544 SYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPTEPDN 1603
Query: 687 SLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYRNLANTVAE 746
L+A DA + P ++S G++ L ++ S K SL S+TEAE ++ +V
Sbjct: 1604 KLVAISDASYGNQP-YYKSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAEIHAISESVPL 1662
Query: 747 LLWVQSLLQELHIPHKTPLI---LCDNMSTVAL---THNPVLHTR---TKHMELDIFFVR 797
L + L+QEL +K P+I L D+ ST+++ T+ R TK M L R
Sbjct: 1663 LNNLSYLIQEL---NKKPIIKGLLTDSRSTISIIKSTNEEKFRNRFFGTKAMRL-----R 1714
Query: 798 EKVLNKSLIIQHVPSLDQRDDILTKALSPLRFEDLRTK 835
++V +L + ++ + D++TK L F+ L K
Sbjct: 1715 DEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKLLTNK 1752
Score = 38.9 bits (89), Expect = 0.058
Identities = 25/89 (28%), Positives = 43/89 (48%), Gaps = 10/89 (11%)
Query: 5 TRYTWVFPLKKKS-----DTYTTFVNFHKMIELQLNKKVKVVQTDGGGEF--KPLTPYFT 57
T++ WV+PL + D +TT + F I+ Q V V+Q D G E+ + L +
Sbjct: 695 TKFRWVYPLHDRREDSILDVFTTILAF---IKNQFQASVLVIQMDRGSEYTNRTLHKFLE 751
Query: 58 NLGIVHRFTCPHTHHQNGIVERKHRHIVE 86
GI +T +G+ ER +R +++
Sbjct: 752 KNGITPCYTTTADSRAHGVAERLNRTLLD 780
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 67.8 bits (164), Expect = 1e-10
Identities = 41/90 (45%), Positives = 55/90 (60%), Gaps = 8/90 (8%)
Query: 630 YVTITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHLKPCSLHQPLSLI 689
Y+TITRP+++++VN++ QF S AV ++L Y+KGTV GL S L L
Sbjct: 2 YLTITRPDLTFAVNRLSQFSSASRTAQMQAVYKVLHYVKGTVGQGLFYSATS---DLQLK 58
Query: 690 AFCDADWAADPDDHRSTSG--SLV---FLG 714
AF D+DWA+ PD RS +G SLV FLG
Sbjct: 59 AFADSDWASCPDTRRSVTGFCSLVPLWFLG 88
>GP1_CHLRE (Q9FPQ6) Vegetative cell wall protein gp1 precursor
(Hydroxyproline-rich glycoprotein 1)
Length = 555
Score = 55.1 bits (131), Expect = 8e-07
Identities = 33/65 (50%), Positives = 35/65 (53%), Gaps = 2/65 (3%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQS--SPAPISPALSSPAPQSSSTDSS 294
SPAP SP P SPA S P P P SP P S SPAP SPA SPAP S + S
Sbjct: 42 SPAPPSPAPPSPAPPSPAPPSPAPPSPGPPSPAPPSPPSPAPPSPAPPSPAPPSPAPPSP 101
Query: 295 SSVSP 299
+ SP
Sbjct: 102 APPSP 106
Score = 53.9 bits (128), Expect = 2e-06
Identities = 29/63 (46%), Positives = 32/63 (50%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSS 296
SPAP SP P SPA S P P P SP P S P+P P+ S PAP S S S +
Sbjct: 85 SPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPPSPAPPSPSPPAPPSPSPPSPAP 144
Query: 297 VSP 299
P
Sbjct: 145 PLP 147
Score = 53.1 bits (126), Expect = 3e-06
Identities = 41/103 (39%), Positives = 47/103 (44%), Gaps = 7/103 (6%)
Query: 205 LFGCASSPQLSSVGSVHSSIPLFSFPQLTTLS----SPAPHSPNTTLPESSSPASQSSTP 260
+F C SP S + P + P S SPAP SP + P SPA S P
Sbjct: 36 IFNCPPSPAPPSPAPPSPAPPSPAPPSPAPPSPGPPSPAPPSPPS--PAPPSPAPPSPAP 93
Query: 261 ILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPIVNP 303
P P SP P SPAP SPA SPAP S + + S SP P
Sbjct: 94 PSPAPPSPAP-PSPAPPSPAPPSPAPPSPPSPAPPSPSPPAPP 135
Score = 51.2 bits (121), Expect = 1e-05
Identities = 30/69 (43%), Positives = 36/69 (51%), Gaps = 2/69 (2%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQS--SPAPISPALSSPAPQSSSTDSS 294
SPAP SP P SPA S P P P SP P S P+P SPA SP+P + + S
Sbjct: 80 SPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPPSPAPPSPSPPAPPSPSP 139
Query: 295 SSVSPIVNP 303
S +P + P
Sbjct: 140 PSPAPPLPP 148
Score = 50.4 bits (119), Expect = 2e-05
Identities = 30/67 (44%), Positives = 38/67 (55%), Gaps = 2/67 (2%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSS 296
SP+P SP LP S +P S S P+ P P+ PVP S PAP SP SP+P + + S
Sbjct: 136 SPSPPSPAPPLPPSPAPPSPSP-PVPPSPSPPVPPS-PAPPSPTPPSPSPPVPPSPAPPS 193
Query: 297 VSPIVNP 303
+P V P
Sbjct: 194 PAPPVPP 200
Score = 50.1 bits (118), Expect = 3e-05
Identities = 28/67 (41%), Positives = 35/67 (51%), Gaps = 4/67 (5%)
Query: 237 SPAPHSPNTTLPESSSPASQSS----TPILPEPTSPVPQSSPAPISPALSSPAPQSSSTD 292
SPAP SP +P S +P S + +P P P SP P S P+P P+ S PAP S
Sbjct: 188 SPAPPSPAPPVPPSPAPPSPAPPVPPSPAPPSPPSPAPPSPPSPAPPSPSPPAPPSPVPP 247
Query: 293 SSSSVSP 299
S + SP
Sbjct: 248 SPAPPSP 254
Score = 49.7 bits (117), Expect = 3e-05
Identities = 32/69 (46%), Positives = 35/69 (50%), Gaps = 4/69 (5%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSS--PAPISPALSSPAPQSSSTDSS 294
SPAP SP P SPA S P P P SP P S PAP SP+ SPAP + +
Sbjct: 95 SPAPPSPAPPSPAPPSPAPPSPAP--PSPPSPAPPSPSPPAPPSPSPPSPAPPLPPSPAP 152
Query: 295 SSVSPIVNP 303
S SP V P
Sbjct: 153 PSPSPPVPP 161
Score = 49.3 bits (116), Expect = 4e-05
Identities = 31/67 (46%), Positives = 36/67 (53%), Gaps = 2/67 (2%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSS 296
SPAP SP+ +P S SP S P P PT P P S P P SPA SPAP + + S
Sbjct: 149 SPAPPSPSPPVPPSPSPPVPPS-PAPPSPTPPSP-SPPVPPSPAPPSPAPPVPPSPAPPS 206
Query: 297 VSPIVNP 303
+P V P
Sbjct: 207 PAPPVPP 213
Score = 49.3 bits (116), Expect = 4e-05
Identities = 27/68 (39%), Positives = 35/68 (50%), Gaps = 1/68 (1%)
Query: 237 SPAPHSPNTTLPESSSPASQSS-TPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSS 295
SP P SP+ +P S +P S + P P P SP P P+P P+ SPAP S + +
Sbjct: 175 SPTPPSPSPPVPPSPAPPSPAPPVPPSPAPPSPAPPVPPSPAPPSPPSPAPPSPPSPAPP 234
Query: 296 SVSPIVNP 303
S SP P
Sbjct: 235 SPSPPAPP 242
Score = 48.5 bits (114), Expect = 7e-05
Identities = 30/65 (46%), Positives = 34/65 (52%), Gaps = 3/65 (4%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSS--PAPISPALSSPAPQSSSTDSS 294
SPAP SP +P S +P S S P P P SP P S PAP SP SPAP S + S
Sbjct: 201 SPAPPSPAPPVPPSPAPPSPPS-PAPPSPPSPAPPSPSPPAPPSPVPPSPAPPSPAPPSP 259
Query: 295 SSVSP 299
+P
Sbjct: 260 KPPAP 264
Score = 47.4 bits (111), Expect = 2e-04
Identities = 26/63 (41%), Positives = 31/63 (48%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSS 296
SPAP SP P SPA S P P P SP + P+P PA SP+P S + S
Sbjct: 90 SPAPPSPAPPSPAPPSPAPPSPAPPSPAPPSPPSPAPPSPSPPAPPSPSPPSPAPPLPPS 149
Query: 297 VSP 299
+P
Sbjct: 150 PAP 152
Score = 47.4 bits (111), Expect = 2e-04
Identities = 30/69 (43%), Positives = 35/69 (50%), Gaps = 6/69 (8%)
Query: 237 SPAPHSPNTTLPESSSPASQSS----TPILPEPTSPVPQSS--PAPISPALSSPAPQSSS 290
SPAP SP+ P S SP S + +P P P+ PVP S P P SPA SP P S S
Sbjct: 123 SPAPPSPSPPAPPSPSPPSPAPPLPPSPAPPSPSPPVPPSPSPPVPPSPAPPSPTPPSPS 182
Query: 291 TDSSSSVSP 299
S +P
Sbjct: 183 PPVPPSPAP 191
Score = 47.0 bits (110), Expect = 2e-04
Identities = 31/76 (40%), Positives = 39/76 (50%), Gaps = 3/76 (3%)
Query: 237 SPAPHSPNTT-LPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDS-S 294
SPAP +P T P SP S P+ P P P P SP P SPA +P+P S + S S
Sbjct: 296 SPAPPTPPTPPSPSPPSPVPPSPAPVPPSPAPPSPAPSPPP-SPAPPTPSPSPSPSPSPS 354
Query: 295 SSVSPIVNPHNTHSMV 310
S SP +P + S +
Sbjct: 355 PSPSPSPSPSPSPSPI 370
Score = 46.2 bits (108), Expect = 4e-04
Identities = 33/79 (41%), Positives = 40/79 (49%), Gaps = 5/79 (6%)
Query: 229 FPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVP-QSSPAPISPALS---SP 284
FP T + P+P SP + + P S +P P P SP P SPAP SPA S SP
Sbjct: 280 FPANTPMP-PSPPSPPPSPAPPTPPTPPSPSPPSPVPPSPAPVPPSPAPPSPAPSPPPSP 338
Query: 285 APQSSSTDSSSSVSPIVNP 303
AP + S S S SP +P
Sbjct: 339 APPTPSPSPSPSPSPSPSP 357
Score = 45.8 bits (107), Expect = 5e-04
Identities = 32/68 (47%), Positives = 35/68 (51%), Gaps = 4/68 (5%)
Query: 237 SPAPHSPNTTLPESS-SPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSS 295
SPAP SP + P S SPA S +P P P SPVP S PAP SPA SP P + S
Sbjct: 214 SPAPPSPPSPAPPSPPSPAPPSPSP--PAPPSPVPPS-PAPPSPAPPSPKPPAPPPPPSP 270
Query: 296 SVSPIVNP 303
P P
Sbjct: 271 PPPPPPRP 278
Score = 45.1 bits (105), Expect = 8e-04
Identities = 31/88 (35%), Positives = 40/88 (45%), Gaps = 2/88 (2%)
Query: 238 PAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALS-SPAPQSSSTDS-SS 295
P P +P + P S P S + P P P SP P P+P P S SP+P S + S S
Sbjct: 300 PTPPTPPSPSPPSPVPPSPAPVPPSPAPPSPAPSPPPSPAPPTPSPSPSPSPSPSPSPSP 359
Query: 296 SVSPIVNPHNTHSMVTRAKDGIVKPRLV 323
S SP +P S + V +LV
Sbjct: 360 SPSPSPSPSPIPSPSPKPSPSPVAVKLV 387
Score = 45.1 bits (105), Expect = 8e-04
Identities = 27/81 (33%), Positives = 37/81 (45%), Gaps = 6/81 (7%)
Query: 225 PLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSS------TPILPEPTSPVPQSSPAPIS 278
PL P + S P P SP+ +P S +P S + P P P SP P P+P
Sbjct: 145 PLPPSPAPPSPSPPVPPSPSPPVPPSPAPPSPTPPSPSPPVPPSPAPPSPAPPVPPSPAP 204
Query: 279 PALSSPAPQSSSTDSSSSVSP 299
P+ + P P S + S S +P
Sbjct: 205 PSPAPPVPPSPAPPSPPSPAP 225
Score = 44.3 bits (103), Expect = 0.001
Identities = 23/63 (36%), Positives = 30/63 (47%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSS 296
SPAP SP P+ +P S P P P P P ++P P SP P+P + + S
Sbjct: 248 SPAPPSPAPPSPKPPAPPPPPSPPPPPPPRPPFPANTPMPPSPPSPPPSPAPPTPPTPPS 307
Query: 297 VSP 299
SP
Sbjct: 308 PSP 310
Score = 44.3 bits (103), Expect = 0.001
Identities = 29/67 (43%), Positives = 35/67 (51%), Gaps = 5/67 (7%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSS 296
SP P SP P + P+ S P P P +P P SP+P SP SP+P S S S S
Sbjct: 312 SPVPPSPAPVPPSPAPPSPAPSPPPSPAPPTPSPSPSPSP-SP---SPSP-SPSPSPSPS 366
Query: 297 VSPIVNP 303
SPI +P
Sbjct: 367 PSPIPSP 373
Score = 44.3 bits (103), Expect = 0.001
Identities = 24/64 (37%), Positives = 32/64 (49%), Gaps = 2/64 (3%)
Query: 236 SSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSS 295
S P+P P+ + P SP+ S P P P SP P S P+ P+ S P P S + S +
Sbjct: 120 SPPSPAPPSPSPPAPPSPSPPSPAP--PLPPSPAPPSPSPPVPPSPSPPVPPSPAPPSPT 177
Query: 296 SVSP 299
SP
Sbjct: 178 PPSP 181
Score = 41.2 bits (95), Expect = 0.012
Identities = 25/65 (38%), Positives = 33/65 (50%), Gaps = 3/65 (4%)
Query: 237 SPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSS--PAPISPALSSPAPQSSSTDSS 294
SPAP SP + P S SP + S P P P P+P S P+P P SP+P + +
Sbjct: 115 SPAPPSPPSPAPPSPSPPAPPS-PSPPSPAPPLPPSPAPPSPSPPVPPSPSPPVPPSPAP 173
Query: 295 SSVSP 299
S +P
Sbjct: 174 PSPTP 178
Score = 41.2 bits (95), Expect = 0.012
Identities = 39/120 (32%), Positives = 48/120 (39%), Gaps = 10/120 (8%)
Query: 230 PQLTTLSSPAPHSP--NTTLPESSSPASQSSTPI-LPEPTSPVPQSSPAPISPALSSPAP 286
P T SP+P SP + P SPA S P P P P P SP+P SP+P
Sbjct: 300 PTPPTPPSPSPPSPVPPSPAPVPPSPAPPSPAPSPPPSPAPPTPSPSPSPSPSPSPSPSP 359
Query: 287 QSSSTDSSS---SVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDP 343
S + S S S SP +P + A D I L T P SA + +P
Sbjct: 360 SPSPSPSPSPIPSPSPKPSPSPVAVKLVWADDAIAFDDLNGT----STRPGSASRMVGEP 415
Score = 40.0 bits (92), Expect = 0.026
Identities = 26/71 (36%), Positives = 32/71 (44%), Gaps = 8/71 (11%)
Query: 237 SPAPHSPNTTLPESS---SPASQSSTPILPEPTSPVPQSSPAPISP-----ALSSPAPQS 288
SPAP SP+ P S SPA S P P+P +P P SP P P ++P P S
Sbjct: 230 SPAPPSPSPPAPPSPVPPSPAPPSPAPPSPKPPAPPPPPSPPPPPPPRPPFPANTPMPPS 289
Query: 289 SSTDSSSSVSP 299
+ S P
Sbjct: 290 PPSPPPSPAPP 300
Score = 38.1 bits (87), Expect = 0.100
Identities = 19/49 (38%), Positives = 23/49 (46%)
Query: 236 SSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSP 284
S P+P P+ + P SP S P P P SP P + P P SP P
Sbjct: 227 SPPSPAPPSPSPPAPPSPVPPSPAPPSPAPPSPKPPAPPPPPSPPPPPP 275
Score = 38.1 bits (87), Expect = 0.100
Identities = 27/66 (40%), Positives = 30/66 (44%), Gaps = 5/66 (7%)
Query: 238 PAPHSP---NTTLPESS-SPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDS 293
P P P NT +P S SP + P P P SP P S P P SPA P+P S
Sbjct: 274 PPPRPPFPANTPMPPSPPSPPPSPAPPTPPTPPSPSPPS-PVPPSPAPVPPSPAPPSPAP 332
Query: 294 SSSVSP 299
S SP
Sbjct: 333 SPPPSP 338
Score = 37.4 bits (85), Expect = 0.17
Identities = 22/47 (46%), Positives = 24/47 (50%), Gaps = 1/47 (2%)
Query: 253 PASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSP 299
P + P P P SP P SPAP SPA SPAP S S + SP
Sbjct: 33 PGGIFNCPPSPAPPSPAP-PSPAPPSPAPPSPAPPSPGPPSPAPPSP 78
Score = 35.4 bits (80), Expect = 0.65
Identities = 27/72 (37%), Positives = 33/72 (45%), Gaps = 6/72 (8%)
Query: 236 SSPAPHSPNTTLPESSS-PASQSSTPILPEPTSPVPQS--SPAPISPALSSPAP-QSSST 291
S P P P P ++ P S S P P P P P + SP+P SP SPAP S
Sbjct: 269 SPPPPPPPRPPFPANTPMPPSPPSPP--PSPAPPTPPTPPSPSPPSPVPPSPAPVPPSPA 326
Query: 292 DSSSSVSPIVNP 303
S + SP +P
Sbjct: 327 PPSPAPSPPPSP 338
>AMYH_YEAST (P08640) Glucoamylase S1/S2 precursor (EC 3.2.1.3)
(Glucan 1,4-alpha-glucosidase) (1,4-alpha-D-glucan
glucohydrolase)
Length = 1367
Score = 52.0 bits (123), Expect = 7e-06
Identities = 49/163 (30%), Positives = 73/163 (44%), Gaps = 11/163 (6%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSP 268
+SS SS V SS S + T SS S + +P SS ++SS+ +P P+S
Sbjct: 615 SSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSS 674
Query: 269 VPQSSPAPI--SPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTL 326
+SS AP+ S SS AP +SST SSS +P+ P ++ + + A VPT
Sbjct: 675 TTESSSAPVTSSTTESSSAPVTSSTTESSS-APVPTPSSSTTESSSAP--------VPTP 725
Query: 327 LLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPS 369
S E +SA + + + + + VP PS
Sbjct: 726 SSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPS 768
Score = 51.6 bits (122), Expect = 9e-06
Identities = 46/134 (34%), Positives = 62/134 (45%), Gaps = 13/134 (9%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTT----LPESSSPASQSSTPILPE 264
A P SS + SS P+ + TT SS AP + +TT P +SS SS P+ P
Sbjct: 651 APVPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAPV-PT 709
Query: 265 PTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRL-- 322
P+S +SS AP+ P P SS+T+SSS+ P + T S +
Sbjct: 710 PSSSTTESSSAPV------PTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAP 763
Query: 323 VPTLLLSQAEPTSA 336
VPT S E +SA
Sbjct: 764 VPTPSSSTTESSSA 777
Score = 51.6 bits (122), Expect = 9e-06
Identities = 38/111 (34%), Positives = 60/111 (53%), Gaps = 14/111 (12%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTT------LPESSSPASQSSTPIL 262
A P SS + SS P+ + TT SS AP + +TT +P SS ++SS+ +
Sbjct: 720 APVPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPV 779
Query: 263 PEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRA 313
P P+S +SS AP+ P P SS+T+ SSV+P+ P ++ ++ + A
Sbjct: 780 PTPSSSTTESSSAPV------PTPSSSTTE--SSVAPVPTPSSSSNITSSA 822
Score = 50.1 bits (118), Expect = 3e-05
Identities = 49/139 (35%), Positives = 69/139 (49%), Gaps = 23/139 (16%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTT------LPESSSPASQSSTPIL 262
A P SS + SS P+ S TT SS AP + +TT +P SS ++SS+ +
Sbjct: 666 APVPTPSSSTTESSSAPVTSS---TTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPV 722
Query: 263 PEPTSPVPQSSPAPI-----SPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGI 317
P P+S +SS AP+ S SS AP +SST SSS +P+ P ++ + + A
Sbjct: 723 PTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSS-APVPTPSSSTTESSSAP--- 778
Query: 318 VKPRLVPTLLLSQAEPTSA 336
VPT S E +SA
Sbjct: 779 -----VPTPSSSTTESSSA 792
Score = 50.1 bits (118), Expect = 3e-05
Identities = 41/132 (31%), Positives = 63/132 (47%), Gaps = 12/132 (9%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSP 268
+SS SS V SS S + T SS S + +P SS ++SS+ P P+S
Sbjct: 558 SSSTTESSSTPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPAPTPSSS 617
Query: 269 VPQSSPAPISPAL----SSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVP 324
+SS AP++ + S+P P SS+ + SS +P+ P ++ + + A VP
Sbjct: 618 TTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAP--------VP 669
Query: 325 TLLLSQAEPTSA 336
T S E +SA
Sbjct: 670 TPSSSTTESSSA 681
Score = 50.1 bits (118), Expect = 3e-05
Identities = 33/104 (31%), Positives = 52/104 (49%), Gaps = 10/104 (9%)
Query: 211 SPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPI----LPEPT 266
+P S+ S + +P S + S+P P ++T SS+P + S+T +P P+
Sbjct: 709 TPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPS 768
Query: 267 SPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMV 310
S +SS AP+ P P SS+T+SSS+ P + T S V
Sbjct: 769 SSTTESSSAPV------PTPSSSTTESSSAPVPTPSSSTTESSV 806
Score = 49.7 bits (117), Expect = 3e-05
Identities = 46/173 (26%), Positives = 79/173 (45%), Gaps = 13/173 (7%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPI----LPE 264
+SS ++S + SS P+ S ++ S+P P ++T SS+P + S+T +P
Sbjct: 429 SSSAPVTSSTTESSSAPVTSSTTESS-SAPVPTPSSSTTESSSAPVTSSTTESSSAPVPT 487
Query: 265 PTSPVPQSSPAPISPALSS------PAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIV 318
P+S +SS AP++ + + P P SS+T+SSS+ +P + T S
Sbjct: 488 PSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPAPTPSSSTTESSSAPVTSSTT 547
Query: 319 KPRL--VPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPS 369
+ VPT S E +S + + SA ++ + + VP PS
Sbjct: 548 ESSSAPVPTPSSSTTESSSTPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPS 600
Score = 47.4 bits (111), Expect = 2e-04
Identities = 51/192 (26%), Positives = 82/192 (42%), Gaps = 42/192 (21%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSP 268
+SS SS V SS S + T SS S + +P SS ++SS+ +P P+S
Sbjct: 741 SSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVPTPSSS 800
Query: 269 VPQSSPAPI-SPALSS-----------------------PAPQSSSTDSS---------- 294
+SS AP+ +P+ SS P P SS+T+SS
Sbjct: 801 TTESSVAPVPTPSSSSNITSSAPSSTPFSSSTESSSVPVPTPSSSTTESSSAPVSSSTTE 860
Query: 295 SSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFN 354
SSV+P+ P ++ ++ + A P +P +++ T + K Y Q E +
Sbjct: 861 SSVAPVPTPSSSSNITSSA------PSSIPFSSTTESFSTGTTVTPSSSK-YPGSQTETS 913
Query: 355 ALHSNQTWTLVP 366
+ +T T+VP
Sbjct: 914 VSSTTET-TIVP 924
Score = 47.0 bits (110), Expect = 2e-04
Identities = 44/144 (30%), Positives = 69/144 (47%), Gaps = 30/144 (20%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTT------LPESSSPASQSSTPIL 262
+SS ++S + SS P+ + TT SS P + +TT +P SS ++SS+ +
Sbjct: 537 SSSAPVTSSTTESSSAPVPTPSSSTTESSSTPVTSSTTESSSAPVPTPSSSTTESSSAPV 596
Query: 263 PEPTSPVPQSSPAPISPALSSPAPQSSSTDSS----------SSVSPIVNPHNTHSMVTR 312
P P+S +SS AP +P P SS+T+SS SS +P+ P ++ + +
Sbjct: 597 PTPSSSTTESSSAP------APTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSS 650
Query: 313 AKDGIVKPRLVPTLLLSQAEPTSA 336
A VPT S E +SA
Sbjct: 651 AP--------VPTPSSSTTESSSA 666
Score = 45.8 bits (107), Expect = 5e-04
Identities = 47/166 (28%), Positives = 75/166 (44%), Gaps = 17/166 (10%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEP-TS 267
+SS ++S + SS P+ + TT SS AP + +TT SS+P + S+T P TS
Sbjct: 378 SSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTT-ESSSAPVTSSTTESSSAPVTS 436
Query: 268 PVPQSSPAPISPAL----SSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLV 323
+SS AP++ + S+P P SS+ + SS +P+ + S V
Sbjct: 437 STTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAP-----------V 485
Query: 324 PTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPS 369
PT S E +SA + + SA ++ + + P PS
Sbjct: 486 PTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPAPTPS 531
Score = 44.7 bits (104), Expect = 0.001
Identities = 31/89 (34%), Positives = 49/89 (54%), Gaps = 7/89 (7%)
Query: 224 IPLFSFPQLTTLSSPAPHSPNTTLPESSSPA----SQSSTPILPEPTSPVPQSSPAPI-- 277
+P S + S+P P ++T SS+P ++SS+ +P P+S +SS AP+
Sbjct: 314 VPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVTS 373
Query: 278 SPALSSPAPQSSSTDSSSSVSPIVNPHNT 306
S SS AP +SST SSS +P+ P ++
Sbjct: 374 STTESSSAPVTSSTTESSS-APVPTPSSS 401
Score = 44.7 bits (104), Expect = 0.001
Identities = 33/111 (29%), Positives = 54/111 (47%), Gaps = 9/111 (8%)
Query: 233 TTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSS------PAP 286
TT S T +P SS ++SS+ +P P+S +SS AP++ + + P P
Sbjct: 300 TTTSKTCTKKTTTPVPTPSSSTTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTP 359
Query: 287 QSSSTDSSSS-VSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSA 336
SS+T+SSS+ V+ ++ + + + P VPT S E +SA
Sbjct: 360 SSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAP--VPTPSSSTTESSSA 408
Score = 42.7 bits (99), Expect = 0.004
Identities = 47/176 (26%), Positives = 77/176 (43%), Gaps = 28/176 (15%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPI------- 261
+SS ++S + SS P+ + TT SS AP +P SSS SS P+
Sbjct: 495 SSSAPVTSSTTESSSAPVPTPSSSTTESSSAP-APTP----SSSTTESSSAPVTSSTTES 549
Query: 262 ----LPEPTSPVPQSSPAPISPAL----SSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRA 313
+P P+S +SS P++ + S+P P SS+ + SS +P+ P ++ + + A
Sbjct: 550 SSAPVPTPSSSTTESSSTPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPSSSTTESSSA 609
Query: 314 KDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPS 369
PT S E +SA + + SA ++ + + VP PS
Sbjct: 610 P--------APTPSSSTTESSSAPVTSSTTESSSAPVPTPSSSTTESSSAPVPTPS 657
Score = 42.4 bits (98), Expect = 0.005
Identities = 44/146 (30%), Positives = 64/146 (43%), Gaps = 19/146 (13%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSST-----PI-- 261
A P SS + SS P+ S ++ S+P P ++T SS+P + S+T P+
Sbjct: 327 APVPTPSSSTTESSSAPVTSSTTESS-SAPVPTPSSSTTESSSAPVTSSTTESSSAPVTS 385
Query: 262 ---------LPEPTSPVPQSSPAPI--SPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMV 310
+P P+S +SS AP+ S SS AP +SST SSS + + S
Sbjct: 386 STTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAPVTSSTTESSSAP 445
Query: 311 TRAKDGIVKPRLVPTLLLSQAEPTSA 336
+ VPT S E +SA
Sbjct: 446 VTSSTTESSSAPVPTPSSSTTESSSA 471
Score = 40.0 bits (92), Expect = 0.026
Identities = 38/135 (28%), Positives = 59/135 (43%), Gaps = 10/135 (7%)
Query: 209 ASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTT--------LPESSSPASQSSTP 260
+S+P SS S SS+P+ + TT SS AP S +TT P SSS + S+
Sbjct: 824 SSTPFSSSTES--SSVPVPTPSSSTTESSSAPVSSSTTESSVAPVPTPSSSSNITSSAPS 881
Query: 261 ILPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKP 320
+P ++ S+ ++P+ S + T SS+ + P T + VT +
Sbjct: 882 SIPFSSTTESFSTGTTVTPSSSKYPGSQTETSVSSTTETTIVPTKTTTSVTTPSTTTITT 941
Query: 321 RLVPTLLLSQAEPTS 335
+ T S E TS
Sbjct: 942 TVCSTGTNSAGETTS 956
Score = 35.0 bits (79), Expect = 0.84
Identities = 36/161 (22%), Positives = 60/161 (36%), Gaps = 9/161 (5%)
Query: 215 SSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSP 274
++ G SS S + +T +S S TT S S + SST +S ++P
Sbjct: 203 NNCGGTKSSTTTSSTSESSTTTSSTSESSTTTSSTSESSTTTSSTSESSTSSSTTAPATP 262
Query: 275 APISPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVK------PRLVPTLLL 328
S P P ++++ + +P PH+ + T+ K K VPT
Sbjct: 263 TTTSCTKEKPTPPTTTSCTKEKPTP---PHHDTTPCTKKKTTTSKTCTKKTTTPVPTPSS 319
Query: 329 SQAEPTSAKLALKDPKWYSAMQDEFNALHSNQTWTLVPLPS 369
S E +SA + + + + + VP PS
Sbjct: 320 STTESSSAPVPTPSSSTTESSSAPVTSSTTESSSAPVPTPS 360
>ENL2_ARATH (Q9T076) Early nodulin-like protein 2 precursor
(Phytocyanin-like protein)
Length = 349
Score = 50.1 bits (118), Expect = 3e-05
Identities = 39/108 (36%), Positives = 56/108 (51%), Gaps = 7/108 (6%)
Query: 207 GCASSPQLSSVGSVHSS-----IPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPI 261
G A SP+ SS S +S P P T S PAP + + SS+P + P+
Sbjct: 182 GGAHSPKSSSAVSPATSPPGSMAPKSGSPVSPTTSPPAPPKSTSPVSPSSAPMTSPPAPM 241
Query: 262 LPEPTSPVPQSSPAPISPALSSPAPQSSSTDSSS-SVSPIVNPHNTHS 308
P+ +S +P SS AP++ S AP+SSS S+S +VSP + P + S
Sbjct: 242 APKSSSTIPPSS-APMTSPPGSMAPKSSSPVSNSPTVSPSLAPGGSTS 288
Score = 47.0 bits (110), Expect = 2e-04
Identities = 28/105 (26%), Positives = 51/105 (47%), Gaps = 9/105 (8%)
Query: 201 PYKELFGCASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTP 260
P+ G ++ ++ G HS P+ ++ SP P +T P + + +SS+
Sbjct: 140 PHAAAPGSSTPGSMTPPGGAHS-------PKSSSPVSPTTSPPGSTTPPGGAHSPKSSSA 192
Query: 261 ILPEPTSP--VPQSSPAPISPALSSPAPQSSSTDSSSSVSPIVNP 303
+ P + P + S +P+SP S PAP S++ S S +P+ +P
Sbjct: 193 VSPATSPPGSMAPKSGSPVSPTTSPPAPPKSTSPVSPSSAPMTSP 237
Score = 38.5 bits (88), Expect = 0.076
Identities = 31/94 (32%), Positives = 46/94 (47%), Gaps = 5/94 (5%)
Query: 207 GCASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSP--NTTLPESSSPASQSSTPILPE 264
G SP S S+ P+ S S PAP +P ++T+P SS+P + + P+
Sbjct: 208 GSPVSPTTSPPAPPKSTSPV-SPSSAPMTSPPAPMAPKSSSTIPPSSAPMTSPPGSMAPK 266
Query: 265 PTSPVPQSSPAPISPALSSPAPQSSSTDSSSSVS 298
+SPV S+ +SP+L+ SSS S S S
Sbjct: 267 SSSPV--SNSPTVSPSLAPGGSTSSSPSDSPSGS 298
Score = 37.0 bits (84), Expect = 0.22
Identities = 28/106 (26%), Positives = 47/106 (43%), Gaps = 6/106 (5%)
Query: 233 TTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAPISPALSSPAPQSSSTD 292
+T SP +P ++ P S +P + +P P SP +SP P ++P + S
Sbjct: 135 STAQSPHAAAPGSSTPGSMTPPGGAHSPKSSSPVSPT--TSP----PGSTTPPGGAHSPK 188
Query: 293 SSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKL 338
SSS+VSP +P + + + + P S P+SA +
Sbjct: 189 SSSAVSPATSPPGSMAPKSGSPVSPTTSPPAPPKSTSPVSPSSAPM 234
>NO20_MEDTR (P93329) Early nodulin 20 precursor (N-20)
Length = 268
Score = 49.7 bits (117), Expect = 3e-05
Identities = 42/120 (35%), Positives = 52/120 (43%), Gaps = 10/120 (8%)
Query: 217 VGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSSPAP 276
V V SS P P S+P PH P +LP SP S S +P SP P+S+P P
Sbjct: 129 VAPVLSSPPPPPSPPTPRSSTPIPHPPRRSLP---SPPSPSPSPSPSPSPSPSPRSTPIP 185
Query: 277 ISPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSA 336
P SPA S S S S SP +P S+ D + L P+ S P+ A
Sbjct: 186 -HPRKRSPASPSPSPSLSKSPSPSESP----SLAPSPSDSVAS--LAPSSSPSDESPSPA 238
Score = 43.9 bits (102), Expect = 0.002
Identities = 33/101 (32%), Positives = 51/101 (49%), Gaps = 5/101 (4%)
Query: 230 PQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSP--VPQSSPAPISPALSSPAPQ 287
P + SP+P S P SPAS S +P L + SP P +P+P S +++S AP
Sbjct: 169 PSPSPSPSPSPRSTPIPHPRKRSPASPSPSPSLSKSPSPSESPSLAPSP-SDSVASLAPS 227
Query: 288 SSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLL 328
SS +D S S +P +P ++ S A G ++ + + L
Sbjct: 228 SSPSDESPSPAP--SPSSSGSKGGGAGHGFLEVSIAMMMFL 266
>P100_HCMVA (P08318) Large structural phosphoprotein (pp150) (150
kDa matrix phosphoprotein) (Basic phosphoprotein) (BPP)
Length = 1048
Score = 46.2 bits (108), Expect = 4e-04
Identities = 42/130 (32%), Positives = 57/130 (43%), Gaps = 18/130 (13%)
Query: 228 SFPQLTTLSSPAPH-SPNTTLPESSSPASQ-SSTPI---LPEPTSPVPQSSPAPISPALS 282
S P TT +S P P +S+ A +STP+ L + S P + PA L
Sbjct: 764 SSPMTTTSTSQKPVLGKRVATPHASARAQTVTSTPVQGRLEKQVSGTPSTVPA----TLL 819
Query: 283 SPAPQSSSTDSS---------SSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEP 333
P P SS T SS SS S P + S+++ +D +V P P +LS A P
Sbjct: 820 QPQPASSKTTSSRNVTSGAGTSSASSARQPSASASVLSPTEDDVVSPATSPLSMLSSASP 879
Query: 334 TSAKLALKDP 343
+ AK A P
Sbjct: 880 SPAKSAPPSP 889
>TONB_VIBCH (O52042) TonB protein
Length = 244
Score = 45.4 bits (106), Expect = 6e-04
Identities = 38/139 (27%), Positives = 59/139 (42%), Gaps = 26/139 (18%)
Query: 223 SIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILP-----------EPTSPVPQ 271
SI + S P++ AP P T E S P S TP+ +P P PQ
Sbjct: 42 SINMVSMPKV------APAQPEQTQTEQSKPESVKPTPVQEVVKTPPAKPKVKPHKPKPQ 95
Query: 272 SSPAPI------SPALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIV-KPRLV- 323
+ P+ S +++ P P+ + ++ P P+ S T A GI +P LV
Sbjct: 96 TVKKPVPAEQVPSKSVAKPQPEKVERTAETAQKPAPTPNQQPSQPTAASQGITSQPILVD 155
Query: 324 -PTLLLSQAEPTSAKLALK 341
P L+ +Q +P ++A K
Sbjct: 156 KPALVSAQVQPRYPRIARK 174
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 102,421,621
Number of Sequences: 164201
Number of extensions: 4524184
Number of successful extensions: 28987
Number of sequences better than 10.0: 768
Number of HSP's better than 10.0 without gapping: 174
Number of HSP's successfully gapped in prelim test: 629
Number of HSP's that attempted gapping in prelim test: 21303
Number of HSP's gapped (non-prelim): 4039
length of query: 856
length of database: 59,974,054
effective HSP length: 119
effective length of query: 737
effective length of database: 40,434,135
effective search space: 29799957495
effective search space used: 29799957495
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0322.2


