

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0321.6
(497 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
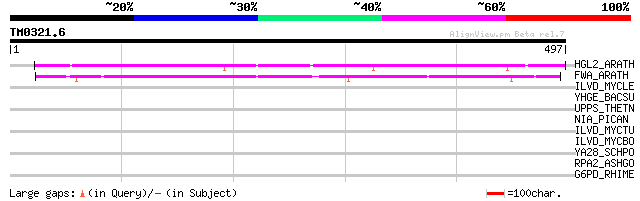
Score E
Sequences producing significant alignments: (bits) Value
HGL2_ARATH (P46607) Homeobox protein GLABRA2 (Homeobox-leucine z... 231 4e-60
FWA_ARATH (Q9FVI6) Homeobox protein FWA 206 1e-52
ILVD_MYCLE (O06069) Dihydroxy-acid dehydratase (EC 4.2.1.9) (DAD) 33 1.7
YHGE_BACSU (P32399) Hypothetical protein yhgE (ORFB) 32 2.9
UPPS_THETN (Q8RA26) Undecaprenyl pyrophosphate synthetase (EC 2.... 32 3.8
NIA_PICAN (P49050) Nitrate reductase [NADPH] (EC 1.7.1.3) (NR) 32 4.9
ILVD_MYCTU (P65154) Dihydroxy-acid dehydratase (EC 4.2.1.9) (DAD) 32 4.9
ILVD_MYCBO (P65155) Dihydroxy-acid dehydratase (EC 4.2.1.9) (DAD) 32 4.9
YA28_SCHPO (Q09699) Hypothetical protein C2F7.08c in chromosome I 31 6.5
RPA2_ASHGO (Q75DS1) DNA-directed RNA polymerase I polypeptide 2 ... 31 8.4
G6PD_RHIME (Q9Z3S2) Glucose-6-phosphate 1-dehydrogenase (EC 1.1.... 31 8.4
>HGL2_ARATH (P46607) Homeobox protein GLABRA2 (Homeobox-leucine
zipper protein ATHB-10) (HD-ZIP protein ATHB-10)
Length = 745
Score = 231 bits (588), Expect = 4e-60
Identities = 152/500 (30%), Positives = 263/500 (52%), Gaps = 30/500 (6%)
Query: 23 VEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYAKKFPRI-TNSENLY 81
+EKS++ ++N A EL + + EP+W++S + +L + Y K+FP+ +S
Sbjct: 250 LEKSRIAEISNRATLELQKMATSGEPMWLRSV-ETGREILNYDEYLKEFPQAQASSFPGR 308
Query: 82 VCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENH-SGALR 140
E+S+D+ +V +L + F+D +W + ++SKA T+ V G + GA++
Sbjct: 309 KTIEASRDAGIVFMDAHKLAQSFMDVGQWKETFACLISKAATVDVIRQGEGPSRIDGAIQ 368
Query: 141 LMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVA----KKFPS 196
LM+ E+ +L+P+VP+RE +F+R C+Q+ +W IVDVS+ + + + +K PS
Sbjct: 369 LMFGEMQLLTPVVPTREVYFVRSCRQLSPEKWAIVDVSVSVEDSNTEKEASLLKCRKLPS 428
Query: 197 GCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKAC 256
GC I + NG VTWVE ++V S + L R +V +A+GA+ W+ LQ E+
Sbjct: 429 GCIIEDTSNGHSKVTWVEHLDVSAS-TVQPLFRSLVNTGLAFGARHWVATLQLHCERLVF 487
Query: 257 SEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIK 316
+ + + L V L GRKSV+ ++ RM SF++ + + + ++
Sbjct: 488 FMATNVPTKDSLG--VTTLAGRKSVLKMAQRMTQSFYRAIAASSYHQWTKITTKTGQDMR 545
Query: 317 LIAHRNTI----LDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAH 372
+ + +N G++V A SS+WLPV+P LF F +DE R+ EWD L+ G ++ A+
Sbjct: 546 VSSRKNLHDPGEPTGVIVCASSSLWLPVSPALLFDFFRDEARRHEWDALSNGAHVQSIAN 605
Query: 373 ISINESNHVSI-LQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKK 431
+S + S+ +QT + + + Q+SS + VVYAP+D +++ G D S
Sbjct: 606 LSKGQDRGNSVAIQTVKSREKSIWVLQDSSTNSYESVVVYAPVDINTTQLVLAGHDPSNI 665
Query: 432 LILPSGFTISRDG--------------REGNEGSLLTMCFQVLVDEEGSSSQVRKKTLES 477
ILPSGF+I DG R GSLLT+ Q L++ ++++ +++ES
Sbjct: 666 QILPSGFSIIPDGVESRPLVITSTQDDRNSQGGSLLTLALQTLIN-PSPAAKLNMESVES 724
Query: 478 IDSSCMLMLQRIKDALHCSD 497
+ + + L IK +L D
Sbjct: 725 VTNLVSVTLHNIKRSLQIED 744
>FWA_ARATH (Q9FVI6) Homeobox protein FWA
Length = 686
Score = 206 bits (523), Expect = 1e-52
Identities = 146/486 (30%), Positives = 236/486 (48%), Gaps = 28/486 (5%)
Query: 24 EKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVV---GFVLVREIYAKKFPRITNSENL 80
E S L +A A++EL+ L P W+ + +V G + E Y F +T
Sbjct: 209 ETSIFLNLAITALRELITLGEVDCPFWM--IDPIVRSKGVSKIYEKYRSSFNNVTKPPGQ 266
Query: 81 YVCEESSKDSLLVMASGMELVKVFLDSDKWADILPTIVSKAKTIQVFDVGSIENHSGALR 140
V E+S+ LV + + LVK +D+ KW ++ IV A T +V GS SG+L+
Sbjct: 267 IV--EASRAKGLVPMTCVTLVKTLMDTGKWVNVFAPIVPVASTHKVISTGSGGTKSGSLQ 324
Query: 141 LMWNEVHVLSPLVPSREYFFLRYCQQVEAGEWLIVDVSLDSFGNCASRSVAKKFPSGCRI 200
+ E V+SPLVP R+ F+RYC+++ G W++VDV+ +K+ PSG I
Sbjct: 325 QIQAEFQVISPLVPKRKVTFIRYCKEIRQGLWVVVDVTPTQNPTLLPYGCSKRLPSGLII 384
Query: 201 RELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEK-KACSEF 259
+L NG VTW+EQ E +S + L + ++ I GAKRWL LQR E S
Sbjct: 385 DDLSNGYSQVTWIEQAEYNES-HIHQLYQPLIGYGIGLGAKRWLATLQRHCESLSTLSST 443
Query: 260 SYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQ-----NLLDPSNQLTTS 314
+ T + GL + G ++ L+ RM ++++ + + + + + + ++
Sbjct: 444 NLTEISPGLSAK-----GATEIVKLAQRMTLNYYRGITSPSVDKWQKIQVENVAQNMSFM 498
Query: 315 IKLIAHRNTILDGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS 374
I+ + L GIV+ A +S+WLPV LF F+ + EWD L M + I
Sbjct: 499 IRKNVNEPGELTGIVLSASTSVWLPVNQHTLFAFISHLSFRHEWDILTNDTTMEETIRIQ 558
Query: 375 INESNHVSILQTSSIIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLIL 434
H +I+ I+ M++ QE D G VVYAP++ ++ G +S L
Sbjct: 559 -KAKRHGNIISLLKIVNNGMLVLQEIWNDASGAMVVYAPVETNSIELVKRGENSDSVKFL 617
Query: 435 PSGFTISRDGREGN-------EGSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLMLQ 487
PSGF+I DG G+ G LLT Q+LV +++ + + T++S+++ +
Sbjct: 618 PSGFSIVPDGVNGSYHRGNTGGGCLLTFGLQILVGINPTAALI-QGTVKSVETLMAHTIV 676
Query: 488 RIKDAL 493
+IK AL
Sbjct: 677 KIKSAL 682
>ILVD_MYCLE (O06069) Dihydroxy-acid dehydratase (EC 4.2.1.9) (DAD)
Length = 564
Score = 33.1 bits (74), Expect = 1.7
Identities = 33/101 (32%), Positives = 48/101 (46%), Gaps = 14/101 (13%)
Query: 331 IAVSSIWLPVTPKNLFLFLKDEFRKA--EWDFLAGGRPMRQKAHISINESNHVSILQTSS 388
I V+S W +TP NL L D KA E F AGG P+ +I+ S+ +S+
Sbjct: 41 IGVASSWNEITPCNLSL---DRLAKAVKEGVFSAGGYPLE---FGTISVSDGISMGHQ-- 92
Query: 389 IIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSS 429
G H + I D V+ A ++LD +++ GCD S
Sbjct: 93 --GMHFSLVSREVIADSVETVMQA--ERLDGSVLLAGCDKS 129
>YHGE_BACSU (P32399) Hypothetical protein yhgE (ORFB)
Length = 775
Score = 32.3 bits (72), Expect = 2.9
Identities = 26/82 (31%), Positives = 42/82 (50%), Gaps = 3/82 (3%)
Query: 246 ELQRTFEK-KACSEFSYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNL 304
E + F K KA SE T S++ G KLLDG K V + S+++ + G+ L
Sbjct: 474 EYNKQFAKAKAGSEQLVTGSSQVSGGLFKLLDGSKQVQSGSSKLADGSASL--DTGLGKL 531
Query: 305 LDPSNQLTTSIKLIAHRNTILD 326
LD + +L++ +K A + +D
Sbjct: 532 LDGTGELSSKLKDAADQTGDID 553
>UPPS_THETN (Q8RA26) Undecaprenyl pyrophosphate synthetase (EC
2.5.1.31) (UPP synthetase)
(Di-trans-poly-cis-decaprenylcistransferase)
(Undecaprenyl diphosphate synthase) (UDS)
Length = 247
Score = 32.0 bits (71), Expect = 3.8
Identities = 22/97 (22%), Positives = 47/97 (47%), Gaps = 1/97 (1%)
Query: 9 VNNPTRESQESEVTVEKSQMLRVANAAMQELLFLLNTSEPLWVKSSNDVVGFVLVREIYA 68
+ + +R ++ ++ +EK+Q L N + + L VK++ ++ VL RE+
Sbjct: 111 IGDISRLPEKCQIEIEKAQKLTEKNTGLVVNIALNYGGRDEIVKATQNICKKVLNRELSP 170
Query: 69 KKFPRITNSENLYVCEESSKDSLLVMASGMELVKVFL 105
+ T +++LY + D L++ SG + + FL
Sbjct: 171 EDITEDTITQHLYTASQPDPD-LIIRTSGEKRLSNFL 206
>NIA_PICAN (P49050) Nitrate reductase [NADPH] (EC 1.7.1.3) (NR)
Length = 859
Score = 31.6 bits (70), Expect = 4.9
Identities = 42/182 (23%), Positives = 73/182 (40%), Gaps = 30/182 (16%)
Query: 49 LWVKSSNDVVGFVLVREIYAKKFPRITNSENLYVCEESSKDSLLVMASGMELVKVFLDSD 108
L V + F+ + +AKK + S +L EE S + + +V LD
Sbjct: 546 LMVSGEDATDDFIAIHSSFAKK---LLPSMHLGRLEEVSSVTKVKSVEQNVKREVLLDPR 602
Query: 109 KWADIL---PTIVSKAKTIQVFDVGSIENHSGALRLMWNEVHVLSPLVPSREYFFLRYCQ 165
KW I ++S I FD+ E SG +P+ ++ FLR
Sbjct: 603 KWHKITLAEKEVISSDSRIFKFDLEHSEQLSG---------------LPTGKHLFLRL-- 645
Query: 166 QVEAGEWLIVDVSLDSFGNCASRS---VAKKFPSGCRIRELPNGCCMVTWVEQVEVGQSF 222
+ +G++++ + S + R + FP+ RE PNG M +E ++VG
Sbjct: 646 KDSSGKYVMRAYTPKSSNSLRGRLEILIKVYFPN----REYPNGGIMTNLIENLQVGNQI 701
Query: 223 QL 224
++
Sbjct: 702 EV 703
>ILVD_MYCTU (P65154) Dihydroxy-acid dehydratase (EC 4.2.1.9) (DAD)
Length = 575
Score = 31.6 bits (70), Expect = 4.9
Identities = 32/101 (31%), Positives = 47/101 (45%), Gaps = 14/101 (13%)
Query: 331 IAVSSIWLPVTPKNLFLFLKDEFRKA--EWDFLAGGRPMRQKAHISINESNHVSILQTSS 388
I V+S W +TP NL L D A E F AGG P+ +I+ S+ +S+
Sbjct: 52 IGVASSWNEITPCNLSL---DRLANAVKEGVFSAGGYPLE---FGTISVSDGISMGHE-- 103
Query: 389 IIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSS 429
G H + I D V+ A ++LD +++ GCD S
Sbjct: 104 --GMHFSLVSREVIADSVEVVMQA--ERLDGSVLLAGCDKS 140
>ILVD_MYCBO (P65155) Dihydroxy-acid dehydratase (EC 4.2.1.9) (DAD)
Length = 575
Score = 31.6 bits (70), Expect = 4.9
Identities = 32/101 (31%), Positives = 47/101 (45%), Gaps = 14/101 (13%)
Query: 331 IAVSSIWLPVTPKNLFLFLKDEFRKA--EWDFLAGGRPMRQKAHISINESNHVSILQTSS 388
I V+S W +TP NL L D A E F AGG P+ +I+ S+ +S+
Sbjct: 52 IGVASSWNEITPCNLSL---DRLANAVKEGVFSAGGYPLE---FGTISVSDGISMGHE-- 103
Query: 389 IIGEHMVIFQESSIDDLGGYVVYAPIDKLDAYMIVNGCDSS 429
G H + I D V+ A ++LD +++ GCD S
Sbjct: 104 --GMHFSLVSREVIADSVEVVMQA--ERLDGSVLLAGCDKS 140
>YA28_SCHPO (Q09699) Hypothetical protein C2F7.08c in chromosome I
Length = 632
Score = 31.2 bits (69), Expect = 6.5
Identities = 32/133 (24%), Positives = 55/133 (41%), Gaps = 26/133 (19%)
Query: 2 ETLNDNVVNNPTRE---------SQESEVTVEKSQMLRVANAAMQELLFLLNTSEPLWVK 52
ETLN+ NNP RE SQ+ ++ VE+SQ L + N ++L L N+
Sbjct: 434 ETLNEINTNNPEREHLIVRLKVDSQKLKIIVEQSQTLTLNNFQQRKLTPLPNS------- 486
Query: 53 SSNDVVGFVLVREIYAKKFPRITNSENL-----YVCEESSKDSLLVMASGMELVKVFLDS 107
+ F + E +++ T +N+ + S D+ L ++ E V +
Sbjct: 487 -----MSFDSLNEFESERNTPSTYGQNISPLPQFSSPSLSSDAFLTNSNSSESALVHMQK 541
Query: 108 DKWADILPTIVSK 120
D P+ V +
Sbjct: 542 LNLPDFTPSWVKR 554
>RPA2_ASHGO (Q75DS1) DNA-directed RNA polymerase I polypeptide 2 (EC
2.7.7.6) (RNA polymerase I subunit 2)
Length = 1198
Score = 30.8 bits (68), Expect = 8.4
Identities = 39/183 (21%), Positives = 70/183 (37%), Gaps = 20/183 (10%)
Query: 260 SYTASTEGLEGEVKLLDGRKSVINLSNRMVGSFFKVLNTQGIQNLLDPSNQLTTSIKLIA 319
S+ A TEG G ++NL R +G+ +N N+L S+ ++
Sbjct: 47 SFNALTEGPGG---------GLLNLGARDIGAKVVFDGKASDENPNYLGNKLALSVTQVS 97
Query: 320 HRNTIL-DGIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHISINES 378
+ DG+ A +++ K L + K W G ++ + +
Sbjct: 98 LTKPMSNDGVTAAAERNVFPAEARKRLTTYRGKLLLKLNWSVNDG-----EETFSEVRDC 152
Query: 379 NHVSILQTSSIIGEHMVIFQE-----SSIDDLGGYVVYAPIDKLDAYMIVNGCDSSKKLI 433
+ ++ S+ H + QE D+LGGY + I+KL +IV + +I
Sbjct: 153 GPLPVMLQSNRCHLHKMSPQELVEHKEESDELGGYFIVNGIEKLIRMLIVQRRNHPMAII 212
Query: 434 LPS 436
PS
Sbjct: 213 RPS 215
>G6PD_RHIME (Q9Z3S2) Glucose-6-phosphate 1-dehydrogenase (EC
1.1.1.49) (G6PD)
Length = 491
Score = 30.8 bits (68), Expect = 8.4
Identities = 24/94 (25%), Positives = 41/94 (43%), Gaps = 13/94 (13%)
Query: 195 PSGCRIRELPNGCCMVTWVEQVEVGQSFQLDMLVRDIVRNNIAYGAKR--------WLLE 246
P G R+R +P +++ E V + + L+ D++RNN +R W+
Sbjct: 395 PGGMRLRNVPLD---MSFAEAFAVRNADAYERLLLDVIRNNQTLFVRRDEVEAAWQWIDP 451
Query: 247 LQRTFEKKACSEFSYTASTEGLEGEVKLL--DGR 278
+ + +E YTA T G + L+ DGR
Sbjct: 452 ILKAWEATGQQVQGYTAGTWGPSQSIALIERDGR 485
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,054,009
Number of Sequences: 164201
Number of extensions: 2151647
Number of successful extensions: 4576
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 4560
Number of HSP's gapped (non-prelim): 12
length of query: 497
length of database: 59,974,054
effective HSP length: 114
effective length of query: 383
effective length of database: 41,255,140
effective search space: 15800718620
effective search space used: 15800718620
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0321.6


