

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0316.5
(318 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
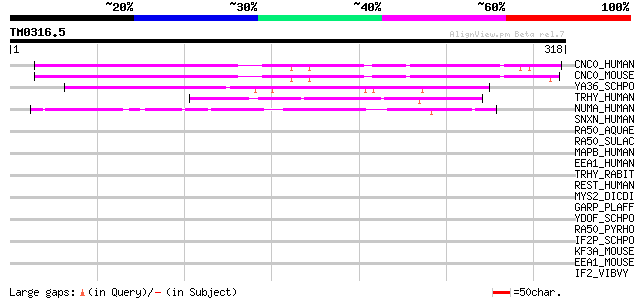
Score E
Sequences producing significant alignments: (bits) Value
CNC0_HUMAN (Q8NEJ9) Protein C14orf120 130 6e-30
CNC0_MOUSE (Q9DB96) Protein C14orf120 homolog 129 7e-30
YA36_SCHPO (Q09713) Hypothetical protein C18B11.06 in chromosome I 85 3e-16
TRHY_HUMAN (Q07283) Trichohyalin 49 2e-05
NUMA_HUMAN (Q14980) Nuclear mitotic apparatus protein 1 (NuMA pr... 45 3e-04
SNXN_HUMAN (Q96L93) Kinesin-like motor protein C20orf23 (Sorting... 41 0.003
RA50_AQUAE (O67124) Probable DNA double-strand break repair rad5... 41 0.003
RA50_SULAC (O33600) DNA double-strand break repair rad50 ATPase 41 0.005
MAPB_HUMAN (P46821) Microtubule-associated protein 1B (MAP 1B) [... 41 0.005
EEA1_HUMAN (Q15075) Early endosome antigen 1 (Endosome-associate... 41 0.005
TRHY_RABIT (P37709) Trichohyalin 40 0.008
REST_HUMAN (P30622) Restin (Cytoplasmic linker protein-170 alpha... 40 0.008
MYS2_DICDI (P08799) Myosin II heavy chain, non muscle 40 0.008
GARP_PLAFF (P13816) Glutamic acid-rich protein precursor 40 0.008
YDOF_SCHPO (O13734) Hypothetical protein C15A10.15 in chromosome I 40 0.010
RA50_PYRHO (O58687) DNA double-strand break repair rad50 ATPase 40 0.010
IF2P_SCHPO (Q10251) Eukaryotic translation initiation factor 5B ... 40 0.010
KF3A_MOUSE (P28741) Kinesin-like protein KIF3A (Microtubule plus... 39 0.013
EEA1_MOUSE (Q8BL66) Early endosome antigen 1 (Fragment) 39 0.017
IF2_VIBVY (Q7MI09) Translation initiation factor IF-2 39 0.022
>CNC0_HUMAN (Q8NEJ9) Protein C14orf120
Length = 315
Score = 130 bits (326), Expect = 6e-30
Identities = 98/329 (29%), Positives = 152/329 (45%), Gaps = 48/329 (14%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
E + LLK ++E + V +++SLT KV+ G YPT G S+LE K+ LLL Y L
Sbjct: 8 ESDLPSAVTLLKNLQEQVMAVTAQVKSLTQKVQAGAYPTEKGLSFLEVKDQLLLMYLMDL 67
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPS 134
+ +L KA G S++ H V +VEIR LEK+RP+D+K +YQI KL+K + S+
Sbjct: 68 THLILDKASGGSLQGHDAVLRLVEIRTVLEKLRPLDQKLKYQIDKLIKTAVTGSLSE--- 124
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSR---EDLAPQEGHD----------------AYRPV 175
D +++P+P ++S+ ED E D Y P
Sbjct: 125 ----------NDPLRFKPHPSNMMSKLSSEDEEEDEAEDDQSEASGKKSVKGVSKKYVPP 174
Query: 176 KFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEV 235
+ P D+ + ++++ R ++ L SS IR L + PEEIRD V
Sbjct: 175 RLVPVHYDETEAEREKKRLERAKRRALS----SSVIRELKEQYSDAPEEIRD--ARHPHV 228
Query: 236 ERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLP---- 291
R + + R EE + R+ ++K E+ R K + L LT F +I L
Sbjct: 229 TRQSQEDQHRINYEESMMVRLSVSKREKGRRKRANVMSSQLHSLTH--FSDISALTGGTV 286
Query: 292 --FEDET--GEQVMGPSKGSNRNGRLKKR 316
ED+ ++ P KG + G ++R
Sbjct: 287 HLDEDQNPIKKRKKIPQKGRKKKGFRRRR 315
>CNC0_MOUSE (Q9DB96) Protein C14orf120 homolog
Length = 315
Score = 129 bits (325), Expect = 7e-30
Identities = 96/323 (29%), Positives = 149/323 (45%), Gaps = 43/323 (13%)
Query: 15 EKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSL 74
E + S LLK ++E + V +IQ+LT KV+ G Y T G S+LE K+ LLL Y L
Sbjct: 8 ESDVSSSITLLKNLQEQVMAVTAQIQALTTKVRAGTYSTEKGLSFLEVKDQLLLMYLMDL 67
Query: 75 VYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPS 134
+ +L KA G S++ HP V +VEIR LEK+RP+D+K +YQI KL+K + S+
Sbjct: 68 SHLILDKASGASLQGHPAVLRLVEIRTVLEKLRPLDQKLKYQIDKLVKTAVTGSLSE--- 124
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSR------EDLAPQEGHD-------------AYRPV 175
D +++P+P +VS+ E+ +EG Y P
Sbjct: 125 ----------NDPLRFKPHPSNMVSKLSSEDEEESEAEEGQSEASGKKSAKGSAKKYVPP 174
Query: 176 KFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEV 235
+ P D+ + ++++ + ++ L SS IR L + PEEIRD V
Sbjct: 175 RLVPVHYDETEAEREQKRLEKAKRRALS----SSVIRELKEQYSDAPEEIRD--ARHPHV 228
Query: 236 ERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDE 295
R + + R EE + R+ ++K E+ + + L LT F +I L
Sbjct: 229 TRQSQEDQHRVNYEESMMVRLSVSKREKGLRRRASAMSSQLHSLTH--FSDISALTGGTA 286
Query: 296 TGEQVMGPSKGSN---RNGRLKK 315
++ P K + GR KK
Sbjct: 287 HLDEDQNPVKKRKKLPKKGRKKK 309
>YA36_SCHPO (Q09713) Hypothetical protein C18B11.06 in chromosome I
Length = 327
Score = 84.7 bits (208), Expect = 3e-16
Identities = 73/268 (27%), Positives = 128/268 (47%), Gaps = 26/268 (9%)
Query: 32 LDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSLVYYLLRKAKGFSIEEHP 91
+DV+ IQ+ + + ++DG S + K LLL+Y Q L + +L K S +H
Sbjct: 1 MDVLNKSIQTALEALPKTSSSSSDGVSLVSLKCQLLLSYVQKLAFLMLVKLDDESFLQHQ 60
Query: 92 -VVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPV--ASDK-SEDV 147
VV +V++R+ +EKIRP++ + QY + KL++ ++ S KEP +DK S+D
Sbjct: 61 DVVEKLVQLRIEIEKIRPLENRIQYSVDKLLRAA--GRKEEIGSIKEPENNGNDKDSQDS 118
Query: 148 SK--YRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEIL--- 202
K Y+PN D E + K + +S D+++ + ++ + R I
Sbjct: 119 LKLHYKPNLSEFADDSDGPASENNVVKEDDKSSISSEDEEEELRSAKDGIYRPPRIRAVT 178
Query: 203 ----KESKH------SSFIRTLMNDI-EEKPEEIRDYEGSSKEV---ERHIAKMEERAWQ 248
K ++H F+ + M+ + + P + E + + ER + KM ER
Sbjct: 179 MDSEKRTRHRPNHLVDEFVSSDMSSVPQSMPSVGSNLEKRGRVIHADERELQKMRERIEY 238
Query: 249 EEELFNRVP-LTKHERKREKYLKKSRNG 275
EE + R+P L+K + K+ K +KK G
Sbjct: 239 EESNYTRLPKLSKKDLKKSKRVKKHDYG 266
>TRHY_HUMAN (Q07283) Trichohyalin
Length = 1898
Score = 48.5 bits (114), Expect = 2e-05
Identities = 38/171 (22%), Positives = 79/171 (45%), Gaps = 10/171 (5%)
Query: 104 EKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDL 163
EK R ++++QY+ K ++ E + P K+ + E KYR ++ E L
Sbjct: 966 EKRRRQERERQYRKDKKLQQKEEQLLGEEPEKRR-----RQEREKKYREEEELQQEEEQL 1020
Query: 164 APQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPE 223
+E + R ++ +D+ ++E LR E+E + + R +++++ E
Sbjct: 1021 LREE-REKRRRQEWERQYRKKDELQQEEEQLLREEREKRRLQERERQYRE-EEELQQEEE 1078
Query: 224 EIRDYEGSSK---EVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKK 271
++ E ++ E+ER K EE +EE+L P + ++RE+ ++
Sbjct: 1079 QLLGEERETRRRQELERQYRKEEELQQEEEQLLREEPEKRRRQERERQCRE 1129
Score = 33.9 bits (76), Expect = 0.55
Identities = 37/207 (17%), Positives = 85/207 (40%), Gaps = 17/207 (8%)
Query: 79 LRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEP 138
L+ KG EE P + L + + ++KQQ + ++ +V + + K+E
Sbjct: 193 LQSCKGHETEEFPDEEQLRRRELLELRRKGREEKQQQRRERQDRVFQEEEEKEW-RKRET 251
Query: 139 VASDKSEDVSKYRPNPDMLVSRED------------LAPQEGHDAYRPVKFAPTSMDQDK 186
V + E + + P + E+ QE + ++ + +
Sbjct: 252 VLRKEEEKLQEEEPQRQRELQEEEEQLRKLERQELRRERQEEEQQQQRLRREQQLRRKQE 311
Query: 187 TSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERA 246
++E+ RRE++ +E + + L + EE+ E+ E + E+ + + +E
Sbjct: 312 EERREQQEERREQQERREQQEERREQQLRREQEERREQQLRREQEEERREQQLRREQEEE 371
Query: 247 WQEEELFNRVPLTKHERKREKYLKKSR 273
+E++L + E +RE+ L++ +
Sbjct: 372 RREQQLRRE----QEEERREQQLRREQ 394
Score = 32.0 bits (71), Expect = 2.1
Identities = 25/122 (20%), Positives = 50/122 (40%), Gaps = 12/122 (9%)
Query: 161 EDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMND--- 217
+ L QE +R + ++ + QE + RE+E L++ + +R D
Sbjct: 1717 QQLRRQERDRKFREEEQLRQGREEQQLRSQESDRKFREEEQLRQEREEQQLRPQQRDGKY 1776
Query: 218 --------IEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYL 269
+EE+ + +R E A E+ +E+EL+ K ++RE+ L
Sbjct: 1777 RWEEEQLQLEEQEQRLRQERDRQYRAEEQFATQEKSRREEQELWQEEE-QKRRQERERKL 1835
Query: 270 KK 271
++
Sbjct: 1836 RE 1837
Score = 31.2 bits (69), Expect = 3.6
Identities = 30/144 (20%), Positives = 63/144 (42%), Gaps = 17/144 (11%)
Query: 160 REDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIE 219
+E L QE + + ++ + +QER+ RE+E L++ + +R+ +
Sbjct: 1692 QEQLRRQERYRKILEEEQLRPEREEQQLRRQERDRKFREEEQLRQGREEQQLRS-----Q 1746
Query: 220 EKPEEIRDYEGSSKEVERHIAKMEER----AWQEEELFNRVPLTKHERKREKYLKKSRNG 275
E + R+ E +E E + ++R W+EE+L ++E+ L++ R+
Sbjct: 1747 ESDRKFREEEQLRQEREEQQLRPQQRDGKYRWEEEQL--------QLEEQEQRLRQERDR 1798
Query: 276 LDGLTESFFDEIKTLPFEDETGEQ 299
E F + K+ E E ++
Sbjct: 1799 QYRAEEQFATQEKSRREEQELWQE 1822
>NUMA_HUMAN (Q14980) Nuclear mitotic apparatus protein 1 (NuMA
protein) (SP-H antigen)
Length = 2115
Score = 44.7 bits (104), Expect = 3e-04
Identities = 62/271 (22%), Positives = 122/271 (44%), Gaps = 35/271 (12%)
Query: 13 SKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQ 72
+KEK+A LA L++ ++ +RH+++ L++ +K+ + + EA ++ Q
Sbjct: 528 AKEKQAQ-LAQTLQQQEQASQGLRHQVEQLSSSLKQKEQQLKEVAEKQEATRQ---DHAQ 583
Query: 73 SLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDL 132
L + + S+ E +E L EK ++ QQ Q+Q + ++A TS
Sbjct: 584 QLA--TAAEEREASLRERDAALKQLEA-LEKEKAAKLEILQQ-QLQVANEARDSAQTSVT 639
Query: 133 PSKKEPVA-SDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQE 191
+++E S K E+ L + + A QE H+A V + ++ E
Sbjct: 640 QAQREKAELSRKVEE----------LQACVETARQEQHEAQAQVAELELQLRSEQQKATE 689
Query: 192 RNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIA---KMEERAWQ 248
+ + +EK+ L+E ++ E ++ +GS +E +R A + ++R
Sbjct: 690 KERVAQEKDQLQE------------QLQALKESLKVTKGSLEEEKRRAADALEEQQRCIS 737
Query: 249 EEELFNRVPLTKHERKREKYLKKSRNGLDGL 279
E + R + +H+R+R K L++ R G GL
Sbjct: 738 ELKAETRSLVEQHKRER-KELEEERAGRKGL 767
Score = 30.8 bits (68), Expect = 4.7
Identities = 54/259 (20%), Positives = 102/259 (38%), Gaps = 47/259 (18%)
Query: 1 MAETVNH-LNNHDSKEKEASPL----AALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTAD 55
+ E + H L + K++E + L AA +KE++E V+ + L K KE + +
Sbjct: 1048 LQEALAHALTEKEGKDQELAKLRGLEAAQIKELEELRQTVKQLKEQLAKKEKE--HASGS 1105
Query: 56 GFSYLEAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQY 115
G QS +A G P + +R + K+ +KQQ
Sbjct: 1106 G--------------AQS-------EAAG---RTEPTGPKLEALRAEVSKLEQQCQKQQE 1141
Query: 116 QIQKLMKV--GENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSRE-DLAPQEGHDAY 172
Q L + E A+ ++ S E + E + + L S + +LA
Sbjct: 1142 QADSLERSLEAERASRAERDSALETLQGQLEEKAQELGHSQSALASAQRELAA------- 1194
Query: 173 RPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSS 232
F D K + + + R ++ + + +S I +L ++ ++ + EG S
Sbjct: 1195 ----FRTKVQDHSKAEDEWKAQVARGRQ--EAERKNSLISSLEEEVSILNRQVLEKEGES 1248
Query: 233 KEVERHIAKMEERAWQEEE 251
KE++R + E++ + EE
Sbjct: 1249 KELKRLVMAESEKSQKLEE 1267
>SNXN_HUMAN (Q96L93) Kinesin-like motor protein C20orf23 (Sorting
nexin 23)
Length = 1317
Score = 41.2 bits (95), Expect = 0.003
Identities = 48/206 (23%), Positives = 88/206 (42%), Gaps = 25/206 (12%)
Query: 87 IEEHPV-VRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPV----AS 141
+E+H V + E+ EKI+P++ + QY+ ++L + +N + L K+
Sbjct: 875 LEKHDESVTDVTEVPQDFEKIKPVEYRLQYKERQLQYLLQNHLPTLLEEKQRAFEILDRG 934
Query: 142 DKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEI 201
S D + Y+ +M E LA + +A + K T ++QE ++EKEI
Sbjct: 935 PLSLDNTLYQVEKEMEEKEEQLAQYQA-NANQLQKLQATFEFTANIARQEEKVRKKEKEI 993
Query: 202 LK-----------------ESKHSSFIR--TLMNDIEEKPEEIRDYEGSSKEVERHIAKM 242
L+ E +HS+ R TL +IEE+ +++ S+E A +
Sbjct: 994 LESREKQQREALERALARLERRHSALQRHSTLGMEIEEQRQKLASLNSGSREQSGLQASL 1053
Query: 243 EERAWQEEELFNRVPLTKHERKREKY 268
E E+ R+ + K++ Y
Sbjct: 1054 EAEQEALEKDQERLEYEIQQLKQKIY 1079
>RA50_AQUAE (O67124) Probable DNA double-strand break repair rad50
ATPase
Length = 978
Score = 41.2 bits (95), Expect = 0.003
Identities = 47/223 (21%), Positives = 104/223 (46%), Gaps = 17/223 (7%)
Query: 79 LRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEP 138
L+KA+ E+ + R + ++ L+++ ++K+ + +KL + A + + E
Sbjct: 230 LKKAE----EKDSLERELSQVVTKLKELENLEKEVEKLREKLEFSRKVAPYVPIAKRIEE 285
Query: 139 VASDKSE-DVSKYRPNPDMLVSREDL--APQEGHDAYRPVKFAPTSMDQDKTSKQERNAL 195
+ +E V K + ++ V +++L A +E + + +++K + L
Sbjct: 286 IDKKLTELKVRKNKLTKELAVLKDELSFAQEELNRIEAEKEKFKEEKEREKELEHRLKKL 345
Query: 196 RREKEILKE-SKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFN 254
+ KEILKE S+ SS ++ + E+ +E D ++ ++ +A+ EE+ + +ELF+
Sbjct: 346 QEIKEILKELSQLSSSLKEKEREYEQAKQEFEDLSERVEKGKKLVAETEEKLEKIKELFS 405
Query: 255 RVPLTKHERKRE---------KYLKKSRNGLDGLTESFFDEIK 288
T + K K LK+ L+ LT+ + ++ K
Sbjct: 406 EEEYTSLKMKERLLVELQRKLKELKEKEGQLENLTQKYKEKKK 448
Score = 38.1 bits (87), Expect = 0.029
Identities = 29/95 (30%), Positives = 52/95 (54%), Gaps = 7/95 (7%)
Query: 186 KTSKQERNALRREKEILKE---SKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKM 242
K + +R AL++E E+LK+ +K +TL N +EE+ +E+++ E +++ + + K
Sbjct: 178 KNLEGKREALKKEYELLKDYTPTKKEVLEKTLKN-LEEELKELKETE---EKLRQELKKA 233
Query: 243 EERAWQEEELFNRVPLTKHERKREKYLKKSRNGLD 277
EE+ E EL V K EK ++K R L+
Sbjct: 234 EEKDSLERELSQVVTKLKELENLEKEVEKLREKLE 268
Score = 36.6 bits (83), Expect = 0.085
Identities = 39/183 (21%), Positives = 85/183 (46%), Gaps = 20/183 (10%)
Query: 104 EKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRP----NPDMLVS 159
E+++ + + ++ Q+L K E K+ + + S+ V+K + ++
Sbjct: 214 EELKELKETEEKLRQELKKAEE----------KDSLERELSQVVTKLKELENLEKEVEKL 263
Query: 160 REDLAPQEGHDAYRPVKFAPTSMDQDKTS-KQERNALRREKEILKESKHSSFIRTLMNDI 218
RE L Y P+ +D+ T K +N L +E +LK+ SF + +N I
Sbjct: 264 REKLEFSRKVAPYVPIAKRIEEIDKKLTELKVRKNKLTKELAVLKDEL--SFAQEELNRI 321
Query: 219 EEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDG 278
E + E+ ++ + KE+E + K++E +E L L+ +++E+ ++++ +
Sbjct: 322 EAEKEKFKEEKEREKELEHRLKKLQE---IKEILKELSQLSSSLKEKEREYEQAKQEFED 378
Query: 279 LTE 281
L+E
Sbjct: 379 LSE 381
>RA50_SULAC (O33600) DNA double-strand break repair rad50 ATPase
Length = 886
Score = 40.8 bits (94), Expect = 0.005
Identities = 67/312 (21%), Positives = 134/312 (42%), Gaps = 49/312 (15%)
Query: 37 HKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSI 96
HKI T + +G + K LL +N + +LR++ G H +++ +
Sbjct: 126 HKILMSTTIIGQGSVESVFSDFPEVMKELLKINKLE-----MLRESNG---PIHSLIKVL 177
Query: 97 VEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKY------ 150
+ L+ I+ I K+++ +I +L K E ++E A +K +++++Y
Sbjct: 178 TDRIRSLQSIKDILKREEAEIDRLKKEIEEIKVKLENIERE--AKEKEDELNQYNTEFNR 235
Query: 151 ----RPNPDMLVSR--------EDLAP-----QEGHDAYRPVKFAPTSMDQDKTSKQERN 193
+ D+L E++A +E Y ++ +D+++ ++ N
Sbjct: 236 IKEIKVQYDILSGELSVVNKKIEEIALRLKDFEEKEKRYNKIETEVKELDENR---EKIN 292
Query: 194 ALRREKEILKE-SKHSSFIRTLMNDIEEKPE------EIRDYEGSSKEVERHIAKMEERA 246
+ K IL + S I + ND++ K E E+ + E +E+E+ ++EE+
Sbjct: 293 TISSFKSILVQIDSLKSQINVVENDLKRKKEKLKRKKELEEKEKQYEEIEKRKKELEEKE 352
Query: 247 WQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKG 306
Q EE+ R+ +R+K + N +D T+ ++IK + + Q+ KG
Sbjct: 353 KQYEEIEKRLTYVLKNIERQKNEIEKLNYVD--TQDLENKIKDV---SDRINQIDNELKG 407
Query: 307 -SNRNGRLKKRK 317
+R G L RK
Sbjct: 408 LLDRRGDLNGRK 419
>MAPB_HUMAN (P46821) Microtubule-associated protein 1B (MAP 1B)
[Contains: MAP1 light chain LC1]
Length = 2468
Score = 40.8 bits (94), Expect = 0.005
Identities = 43/195 (22%), Positives = 87/195 (44%), Gaps = 13/195 (6%)
Query: 86 SIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATS----DLPSKKEPVAS 141
S EE P V + + EK ++ K++ ++K V S ++PSK+EP +
Sbjct: 561 SKEETPEVTKVNHV----EKPPKVESKEKVMVKKDKPVKTETKPSVTEKEVPSKEEP-SP 615
Query: 142 DKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEI 201
K+E K + ++E +E K ++ K+++ +++E++
Sbjct: 616 VKAEVAEKQATDVKPKAAKEKTVKKETKVKPEDKKEEKEKPKKEVAKKEDKTPIKKEEKP 675
Query: 202 LKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKH 261
KE + + + +++P++ E KEV++ + K E++ ++EE + P K
Sbjct: 676 KKEEVKKEVKKEIKKEEKKEPKKEVKKETPPKEVKKEVKKEEKKEVKKEE---KEP-KKE 731
Query: 262 ERKREKYLKKSRNGL 276
+K K KKS L
Sbjct: 732 IKKLPKDAKKSSTPL 746
>EEA1_HUMAN (Q15075) Early endosome antigen 1 (Endosome-associated
protein p162) (Zinc finger FYVE domain containing
protein 2)
Length = 1411
Score = 40.8 bits (94), Expect = 0.005
Identities = 53/230 (23%), Positives = 100/230 (43%), Gaps = 14/230 (6%)
Query: 14 KEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQS 73
K+++ L ALL++ KE + ++ + + L AK++ G+ TA + L+ KN L Q
Sbjct: 521 KDQKIQNLEALLQKSKENISLLEKEREDLYAKIQAGEGETA-VLNQLQEKNHTL----QE 575
Query: 74 LVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLP 133
V L K K S E H + + ++ +K + Q ++ L + N S L
Sbjct: 576 QVTQLTEKLKNQS-ESHKQAQENLHDQVQEQKAHL--RAAQDRVLSL-ETSVNELNSQLN 631
Query: 134 SKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERN 193
KE V S+ + + ++L+S E + D + A ++ QDK + +
Sbjct: 632 ESKEKV----SQLDIQIKAKTELLLSAEAAKTAQRADLQNHLDTAQNAL-QDKQQELNKI 686
Query: 194 ALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKME 243
+ ++ K L + ++E E+ E ++E+E I K+E
Sbjct: 687 TTQLDQVTAKLQDKQEHCSQLESHLKEYKEKYLSLEQKTEELEGQIKKLE 736
>TRHY_RABIT (P37709) Trichohyalin
Length = 1407
Score = 40.0 bits (92), Expect = 0.008
Identities = 33/162 (20%), Positives = 72/162 (44%), Gaps = 8/162 (4%)
Query: 110 DKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGH 169
+++QQ + ++ K E L ++E + E K R +L E L QE
Sbjct: 1087 EREQQLRRERDRKFREE---EQLLQEREEERLRRQERARKLREEEQLLRREEQLLRQERD 1143
Query: 170 DAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYE 229
+R + ++++ +QER RE+E L + + +R +E+ ++R+ E
Sbjct: 1144 RKFREEEQLLQESEEERLRRQERERKLREEEQLLQEREEERLRR-----QERARKLREEE 1198
Query: 230 GSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKK 271
++ E+ + + R +EEE R + ++R++ ++
Sbjct: 1199 QLLRQEEQELRQERARKLREEEQLLRQEEQELRQERDRKFRE 1240
Score = 39.7 bits (91), Expect = 0.010
Identities = 37/171 (21%), Positives = 74/171 (42%), Gaps = 5/171 (2%)
Query: 104 EKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSRED- 162
E++R +++QQ + ++ K E L ++E + E K R +L RE+
Sbjct: 755 ERLRRQEREQQLRRERDRKFREE---EQLLQEREEERLRRQERERKLREEEQLLQEREEE 811
Query: 163 -LAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEK 221
L QE R + ++++ +QER RE+E L + + + E+
Sbjct: 812 RLRRQERERKLREEEQLLQEREEERLRRQERERKLREEEQLLRQEEQELRQERARKLREE 871
Query: 222 PEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKS 272
+ +R E ++ + EE+ ++EE R + R+ E+ L++S
Sbjct: 872 EQLLRQEEQELRQERDRKLREEEQLLRQEEQELRQERDRKLREEEQLLQES 922
Score = 35.4 bits (80), Expect = 0.19
Identities = 19/88 (21%), Positives = 46/88 (51%), Gaps = 3/88 (3%)
Query: 182 MDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAK 241
++Q++ +Q+ LRRE+ + +E + +R + +I E+ + + E + +E+ +
Sbjct: 273 LEQEERREQQ---LRREQRLEQEERREQQLRRELEEIREREQRLEQEERREQRLEQEERR 329
Query: 242 MEERAWQEEELFNRVPLTKHERKREKYL 269
++ + EE+ R + E +RE+ L
Sbjct: 330 EQQLKRELEEIREREQRLEQEERREQLL 357
Score = 33.1 bits (74), Expect = 0.94
Identities = 31/174 (17%), Positives = 79/174 (44%), Gaps = 11/174 (6%)
Query: 100 RLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVS 159
R + E+ + + ++++ ++++ + + L ++E + + E K+R +L
Sbjct: 1098 RKFREEEQLLQEREEERLRRQERARKLREEEQLLRREEQLL--RQERDRKFREEEQLLQE 1155
Query: 160 RED--LAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMND 217
E+ L QE R + ++++ +QER RE+E L +
Sbjct: 1156 SEEERLRRQERERKLREEEQLLQEREEERLRRQERARKLREEEQLLRQEEQEL------- 1208
Query: 218 IEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKK 271
+E+ ++R+ E ++ E+ + + +R ++EEE R + R+R++ ++
Sbjct: 1209 RQERARKLREEEQLLRQEEQELRQERDRKFREEEQLLRREEQELRRERDRKFRE 1262
Score = 30.8 bits (68), Expect = 4.7
Identities = 36/181 (19%), Positives = 75/181 (40%), Gaps = 6/181 (3%)
Query: 100 RLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVS 159
R + E+ + + ++++ ++++ + + L ++E + E K R +L
Sbjct: 772 RKFREEEQLLQEREEERLRRQERERKLREEEQLLQEREEERLRRQERERKLREEEQLLQE 831
Query: 160 RED--LAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMND 217
RE+ L QE R + +Q+ +QER RE+E L + +
Sbjct: 832 REEERLRRQERERKLREEEQLLRQEEQEL--RQERARKLREEEQLLRQEEQELRQERDRK 889
Query: 218 IEEKPEEIRDYEGSSKEVERHIAKMEERAWQ--EEELFNRVPLTKHERKREKYLKKSRNG 275
+ E+ + +R E ++ + EE+ Q EEE R + R+ E+ L++
Sbjct: 890 LREEEQLLRQEEQELRQERDRKLREEEQLLQESEEERLRRQERERKLREEEQLLRREEQE 949
Query: 276 L 276
L
Sbjct: 950 L 950
Score = 30.8 bits (68), Expect = 4.7
Identities = 42/180 (23%), Positives = 78/180 (43%), Gaps = 22/180 (12%)
Query: 143 KSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKE-- 200
+ E K+R +L E +E +R + ++++ +QER RE+E
Sbjct: 1231 RQERDRKFREEEQLLRREEQELRRERDRKFREEEQLLQEREEERLRRQERARKLREEEEQ 1290
Query: 201 ILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTK 260
+L E + +R D + EE E S+ +ER + + EE+ + E
Sbjct: 1291 LLFEEQEEQRLRQ-ERDRRYRAEEQFAREEKSRRLERELRQEEEQRRRRE---------- 1339
Query: 261 HERK-REKYLKKSRNGLDGLTESFFDEIKTLPF-EDETGEQVMGPSKGSNRNGRLKKRKA 318
ERK RE+ L++ + E +++ F ED++ QV+ P G+ + R+ R +
Sbjct: 1340 RERKFREEQLRRQQE-----EEQRRRQLRERQFREDQSRRQVLEP--GTRQFARVPVRSS 1392
Score = 30.4 bits (67), Expect = 6.1
Identities = 29/134 (21%), Positives = 54/134 (39%), Gaps = 10/134 (7%)
Query: 143 KSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEIL 202
+ E K R +L E QE R + ++++ +QER RE+E L
Sbjct: 883 RQERDRKLREEEQLLRQEEQELRQERDRKLREEEQLLQESEEERLRRQERERKLREEEQL 942
Query: 203 KESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERA---WQEEELFNRVPLT 259
+ R E+ ++R+ E +E E + +ERA +EE+L R
Sbjct: 943 LRREEQELRR-------ERARKLREEEQLLQEREEERLRRQERARKLREEEQLLRREEQE 995
Query: 260 KHERKREKYLKKSR 273
+ + K+ ++ +
Sbjct: 996 LRQERDRKFREEEQ 1009
Score = 30.0 bits (66), Expect = 7.9
Identities = 18/82 (21%), Positives = 38/82 (45%)
Query: 190 QERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQE 249
+E+ LRRE E ++E + + + + E +++ R + ++ ER +
Sbjct: 254 REQQQLRRELEEIREREQRLEQEERREQQLRREQRLEQEERREQQLRRELEEIREREQRL 313
Query: 250 EELFNRVPLTKHERKREKYLKK 271
E+ R + E +RE+ LK+
Sbjct: 314 EQEERREQRLEQEERREQQLKR 335
Score = 30.0 bits (66), Expect = 7.9
Identities = 41/190 (21%), Positives = 82/190 (42%), Gaps = 18/190 (9%)
Query: 88 EEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDV 147
EE + R E +L E+ ++Q+ + ++ K+ E L ++E + E
Sbjct: 923 EEERLRRQERERKLREEEQLLRREEQELRRERARKLREE---EQLLQEREEERLRRQERA 979
Query: 148 SKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKH 207
K R +L E QE +R + ++++ +QER+ RE+E ++ +
Sbjct: 980 RKLREEEQLLRREEQELRQERDRKFREEEQLLQEREEERLRRQERDRKFREEE--RQLRR 1037
Query: 208 SSFIRTLMNDIEEK---PEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERK 264
+ + K E+IR E K++ R + +R ++EEE R ++
Sbjct: 1038 QELEEQFRQERDRKFRLEEQIRQ-EKEEKQLRR---QERDRKFREEEQQRR------RQE 1087
Query: 265 REKYLKKSRN 274
RE+ L++ R+
Sbjct: 1088 REQQLRRERD 1097
>REST_HUMAN (P30622) Restin (Cytoplasmic linker protein-170 alpha-2)
(CLIP-170) (Reed-Sternberg intermediate filament
associated protein)
Length = 1427
Score = 40.0 bits (92), Expect = 0.008
Identities = 54/294 (18%), Positives = 127/294 (42%), Gaps = 21/294 (7%)
Query: 4 TVNHLNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAK 63
T L + D+ K +S + +K++++ L+ +I+ L + + L+ +
Sbjct: 758 TEEKLLDLDALRKASSEGKSEMKKLRQQLEAAEKQIKHLEIEKNAESSKASSITRELQGR 817
Query: 64 NLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKV 123
L L N ++L + + K +E +++ + E+ + + Q + KL +
Sbjct: 818 ELKLTNLQENLSE--VSQVKETLEKELQILKE--KFAEASEEAVSVQRSMQETVNKLHQK 873
Query: 124 GE--NAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTS 181
E N +SDL +E +A +++ K ++ ++E L +D +K + +
Sbjct: 874 EEQFNMLSSDLEKLRENLADMEAKFREKDEREEQLIKAKEKLE----NDIAEIMKMSGDN 929
Query: 182 MDQ-----DKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEE-----IRDYEGS 231
Q D+ +ER+ + ++ K ++++SF++ + D+ K E+ + +E
Sbjct: 930 SSQLTKMNDELRLKERDVEELQLKLTKANENASFLQKSIEDMTVKAEQSQQEAAKKHEEE 989
Query: 232 SKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFD 285
KE+ER ++ +E++ + ++ER + K L L ++ D
Sbjct: 990 KKELERKLSDLEKKMETSHNQCQELK-ARYERATSETKTKHEEILQNLQKTLLD 1042
>MYS2_DICDI (P08799) Myosin II heavy chain, non muscle
Length = 2116
Score = 40.0 bits (92), Expect = 0.008
Identities = 35/138 (25%), Positives = 67/138 (48%), Gaps = 7/138 (5%)
Query: 184 QDKTSKQERNALRREKEILKESKHSSFIRT--LMNDIEEKPEEIRDYEGSSKEVERHIAK 241
Q K K+ A+ K+ L+ K IR + ++++EK + + + + VE +
Sbjct: 871 QLKAEKETLKAMYDSKDALEAQKRELEIRVEDMESELDEKKLALENLQNQKRSVEEKVRD 930
Query: 242 MEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTE--SFFDEIKTLPFEDETGEQ 299
+EE +E++L N L K ++K E+ L++ + DG ++ S ++IK + E E
Sbjct: 931 LEEELQEEQKLRN--TLEKLKKKYEEELEEMKRVNDGQSDTISRLEKIKD-ELQKEVEEL 987
Query: 300 VMGPSKGSNRNGRLKKRK 317
S+ S G L+K +
Sbjct: 988 TESFSEESKDKGVLEKTR 1005
Score = 39.7 bits (91), Expect = 0.010
Identities = 58/315 (18%), Positives = 133/315 (41%), Gaps = 29/315 (9%)
Query: 10 NHDSKEKEASPLAALLKEMKEGLDVV---RHKIQSLTAKVKEGQYPTADGFSYLEAKNLL 66
N D+ EK+ L A+L+EMK+ L+ + + L K + + S L++ +
Sbjct: 1109 NRDALEKKKKALDAMLEEMKDQLESTGGEKKSLYDLKVKQESDMEALRNQISELQS-TIA 1167
Query: 67 LLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGEN 126
L +S + + + +G +E + +S VE + ++ DK Q E
Sbjct: 1168 KLEKIKSTLEGEVARLQG-ELEAEQLAKSNVEKQKKKVELDLEDKSAQL-------AEET 1219
Query: 127 AATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDK 186
AA L K+ + + SE ++ + V+ + + + F ++ +
Sbjct: 1220 AAKQALDKLKKKLEQELSEVQTQLSEANNKNVN------SDSTNKHLETSFNNLKLELEA 1273
Query: 187 TSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERA 246
K ++ ++ + E KH + +EE+ ++ E ++E+ +++++++
Sbjct: 1274 EQKAKQALEKKRLGLESELKH------VNEQLEEEKKQKESNEKRKVDLEKEVSELKDQI 1327
Query: 247 WQEEELFNRVPLTKHERKREKYL---KKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGP 303
EEE+ ++ +T+ + K+E L K+ + + +++KTL ++E
Sbjct: 1328 --EEEVASKKAVTEAKNKKESELDEIKRQYADVVSSRDKSVEQLKTLQAKNEELRNTAEE 1385
Query: 304 SKGSNRNGRLKKRKA 318
++G K+KA
Sbjct: 1386 AEGQLDRAERSKKKA 1400
Score = 30.4 bits (67), Expect = 6.1
Identities = 55/280 (19%), Positives = 114/280 (40%), Gaps = 21/280 (7%)
Query: 5 VNHLNNHDSKEKEASPLAA--LLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEA 62
VN + K+KE++ L KE+ E D + ++ S A V E + ++
Sbjct: 1295 VNEQLEEEKKQKESNEKRKVDLEKEVSELKDQIEEEVASKKA-VTEAKNKKESELDEIKR 1353
Query: 63 KNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMK 122
+ +++ V L K + + + E L++ KK ++ +++ +K
Sbjct: 1354 QYADVVSSRDKSVEQL----KTLQAKNEELRNTAEEAEGQLDRAERSKKKAEFDLEEAVK 1409
Query: 123 VGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSM 182
E + ++K A K+E + YR L ++++ ++ Y +K +
Sbjct: 1410 NLEEETAKKVKAEK---AMKKAE--TDYRSTKSELDDAKNVSSEQ----YVQIKRLNEEL 1460
Query: 183 DQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKM 242
+ ++ +E + R I + S + +L ++I+ E SKE+E +A++
Sbjct: 1461 SELRSVLEEADE-RCNSAIKAKKTAESALESLKDEIDAANNAKAKAERKSKELEVRVAEL 1519
Query: 243 EERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTES 282
EE + N + RK++ + R LD TES
Sbjct: 1520 EESLEDKSGTVN----VEFIRKKDAEIDDLRARLDRETES 1555
>GARP_PLAFF (P13816) Glutamic acid-rich protein precursor
Length = 678
Score = 40.0 bits (92), Expect = 0.008
Identities = 41/200 (20%), Positives = 86/200 (42%), Gaps = 10/200 (5%)
Query: 111 KKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHD 170
KK+ ++ L K G++ + +E + + +++ +L S Q G
Sbjct: 164 KKENSEVMSLYKTGQHKPKNATEHGEENLDEEMVSEINNNAQGGLLLSSPYQYREQGGCG 223
Query: 171 AYRPV-KFAPTSMDQDKTS-KQERNALRREKEILK-----ESKHSSFIRTLMNDIEEKPE 223
V + + + D DK + +++ +++E+LK E K IE+K +
Sbjct: 224 IISSVHETSNDTKDNDKENISEDKKEDHQQEEMLKTLDKKERKQKEKEMKEQEKIEKKKK 283
Query: 224 EIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESF 283
+ + E +E ER + +ER +E+E+ + + K +K+E+ KK + E+
Sbjct: 284 KQEEKEKKKQEKERKKQEKKERKQKEKEMKKQKKIEKERKKKEEKEKKKKKHDKENEETM 343
Query: 284 FDEIKTLPFEDETGEQVMGP 303
+T +ET ++M P
Sbjct: 344 QQPDQT---SEETNNEIMVP 360
Score = 35.8 bits (81), Expect = 0.14
Identities = 22/94 (23%), Positives = 42/94 (44%), Gaps = 3/94 (3%)
Query: 158 VSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMND 217
+ R ++ P+ + + K + + K +E + KE+ +ESK ++ +
Sbjct: 515 IGRVNVVPRRDNHKKKMAKIEEAELQKQKHVDKEEDKKEESKEVQEESKE---VQEDEEE 571
Query: 218 IEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEE 251
+EE EE + E +E E + EE +EEE
Sbjct: 572 VEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 605
>YDOF_SCHPO (O13734) Hypothetical protein C15A10.15 in chromosome I
Length = 647
Score = 39.7 bits (91), Expect = 0.010
Identities = 33/134 (24%), Positives = 61/134 (44%), Gaps = 18/134 (13%)
Query: 98 EIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSD---------LPSKKEPVASDKSEDVS 148
+IR Y E IR I K Q +I++L E + + L + E V ++ E+ +
Sbjct: 17 QIRQYKEIIR-ISKAQSIRIKELQLENERLLSENIDLRTTAINLEEQLETVQNENEENKT 75
Query: 149 K-------YRPNPDMLVSREDLAPQEGHDAYRPVKF-APTSMDQDKTSKQERNALRREKE 200
K + D +S+ L QE D ++PV+ +D D E + ++ +E
Sbjct: 76 KLAALLNRFHEETDNFLSKLSLCQQEIQDTFKPVEANLAYDVDTDSEDLDEESVVKDTEE 135
Query: 201 ILKESKHSSFIRTL 214
I+++++H +R L
Sbjct: 136 IIEQAQHDVSLRNL 149
>RA50_PYRHO (O58687) DNA double-strand break repair rad50 ATPase
Length = 879
Score = 39.7 bits (91), Expect = 0.010
Identities = 48/204 (23%), Positives = 89/204 (43%), Gaps = 16/204 (7%)
Query: 92 VVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSEDVSKYR 151
+ I+EIR L KI D K+ + +L+K N ++ S K+ V +++ Y+
Sbjct: 501 IADQIIEIRERLSKINLEDLKRDKEEYELLKSESNKLKGEVESLKKEV-----NELNDYK 555
Query: 152 PNPDMLVSREDLAPQEGHDAY-RPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSF 210
L D A +E + R ++ ++D+ S + R + + ++
Sbjct: 556 NESTKLEIEIDKAKKELSEIEDRLLRLGFKTIDE--LSGRIRELEKFHNKYIEAKNAEKE 613
Query: 211 IRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLK 270
+R ++ ++++ EE+ ++E I K+ Q EL + K+E KREK +K
Sbjct: 614 LRDILESLKDEREELDKAFEELAKIETDIEKVTS---QLNELQRKFDQKKYEEKREKMMK 670
Query: 271 KSR--NGLDGLTESF---FDEIKT 289
S GL+ E DEIK+
Sbjct: 671 LSMEIKGLETKLEELERRRDEIKS 694
>IF2P_SCHPO (Q10251) Eukaryotic translation initiation factor 5B
(eIF-5B) (Translation initiation factor IF-2)
Length = 1079
Score = 39.7 bits (91), Expect = 0.010
Identities = 40/197 (20%), Positives = 80/197 (40%), Gaps = 25/197 (12%)
Query: 125 ENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQ 184
E AA ++P + +K + + + L ++ A ++G + + S +
Sbjct: 165 EAAAPPEIPEVRVKTKKEKERE----KKEREKLRKKQQQAKKKGSTGEDTLASSEVSSEV 220
Query: 185 DKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEK------PEEIRDYEGSSKEVERH 238
D ++ E ++ + K+ + + L +EEK + IR+ E E E+
Sbjct: 221 DISTPAENDSSAKGKQAAGSKRKGPNVTALQKMLEEKRAREEEEQRIREEEARIAEEEKR 280
Query: 239 IAKMEERAWQEEELFNRVPLTKHERKRE------KYLKKSRNGLDGLTESFFDEIKTLPF 292
+A++EE +E L + K +K+E KYL K + L + ++
Sbjct: 281 LAEVEEARKEEARLKKK---EKERKKKEEMKAQGKYLSKKQKEQQALAQRRLQQML---- 333
Query: 293 EDETGEQVMGPSKGSNR 309
E+G +V G S G +
Sbjct: 334 --ESGVRVAGLSNGEKK 348
>KF3A_MOUSE (P28741) Kinesin-like protein KIF3A (Microtubule plus
end-directed kinesin motor 3A)
Length = 701
Score = 39.3 bits (90), Expect = 0.013
Identities = 38/171 (22%), Positives = 77/171 (44%), Gaps = 13/171 (7%)
Query: 111 KKQQYQIQKLMKVGENAATSDLP-SKKEPVASDKSEDVSKYRPNPDML----VSREDLAP 165
+K+ +++K ++ GE + SD+ S+++ + ED K + D VS + +
Sbjct: 364 QKEIEELKKKLEEGEEVSGSDISGSEEDDEEGELGEDGEKRKKRRDQAGKKKVSPDKMVE 423
Query: 166 QEGH-DAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEK--- 221
+ D R M++++ +K RREK++LK + + ++ +E+K
Sbjct: 424 MQAKIDEERKALETKLDMEEEERNKARAELERREKDLLKAQQEHQSLLEKLSALEKKVIV 483
Query: 222 --PEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERK--REKY 268
+ + E K +E ++EER + E+L + + ER EKY
Sbjct: 484 GGVDLLAKAEEQEKLLEESNMELEERRRRAEQLRKELEEKEQERLDIEEKY 534
>EEA1_MOUSE (Q8BL66) Early endosome antigen 1 (Fragment)
Length = 790
Score = 38.9 bits (89), Expect = 0.017
Identities = 47/233 (20%), Positives = 101/233 (43%), Gaps = 14/233 (6%)
Query: 14 KEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYLEAKNLLLLNYCQS 73
K+++ L ALL++ KE + ++ + + L AK++ G+ TA + L+ KN L
Sbjct: 365 KDQKIQNLEALLQKGKESVSLLEKEREDLYAKIQAGEGETA-VLNQLQEKNHALQQQLTQ 423
Query: 74 LVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKLMKVGENAATSDLP 133
L L +++ E + + E + +L + + + +L +S L
Sbjct: 424 LTEKLKNQSESHKQAEENLHDQVQEQKAHLRAAQDRVLSLETSVSEL--------SSQLN 475
Query: 134 SKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERN 193
KE V S+ + + ++L+S E + D + A ++ QDK + +
Sbjct: 476 ESKEKV----SQLDIQIKAKTELLLSAEAAKAAQRADLQNHLDTAQHAL-QDKQQELNKV 530
Query: 194 ALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERA 246
+++ ++ K + L + +++ E+ E +++E HI K+E A
Sbjct: 531 SVQLDQLTAKFQEKQEHCIQLESHLKDHKEKHLSLEQKVEDLEGHIKKLEADA 583
>IF2_VIBVY (Q7MI09) Translation initiation factor IF-2
Length = 907
Score = 38.5 bits (88), Expect = 0.022
Identities = 45/221 (20%), Positives = 85/221 (38%), Gaps = 6/221 (2%)
Query: 100 RLYLEKIRPIDKKQQYQIQKLMKVGENAA--TSDLPSKKEPVASDKSEDVSKYRPNPDML 157
R Y+++ D+ ++ + + E AA ++ +K+E + K E K + +
Sbjct: 95 RTYVKRSAIEDEAKREAEEAAQREAEEAAKRAAEEAAKREAEEAAKREAEEKAKREAEEA 154
Query: 158 VSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMND 217
RE + + + A +D K ++ A R+E E LK + R +
Sbjct: 155 AKREAEKSVDRDAEEKAKRDAEGKAKRDAEEKVKQEAARKEAEELKRRQEEEAKRKAEEE 214
Query: 218 IEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLD 277
+ K EE R+ +KE + EE E+ V +++ R+ E +
Sbjct: 215 SQRKLEEAREMAEKNKE---RWSAAEENKGDMEDTDYHVTTSQYAREAEDEADRKEEEAR 271
Query: 278 GLTESFFDEIKTLPFEDETGEQVMGPSKGSNRNGRLKKRKA 318
+ K ++ G +V KG R G+L K K+
Sbjct: 272 RRKKKTKSSAKASENDERGGPRVQRGGKG-GRKGKLSKPKS 311
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.130 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 37,896,008
Number of Sequences: 164201
Number of extensions: 1655527
Number of successful extensions: 6234
Number of sequences better than 10.0: 275
Number of HSP's better than 10.0 without gapping: 28
Number of HSP's successfully gapped in prelim test: 251
Number of HSP's that attempted gapping in prelim test: 5763
Number of HSP's gapped (non-prelim): 574
length of query: 318
length of database: 59,974,054
effective HSP length: 110
effective length of query: 208
effective length of database: 41,911,944
effective search space: 8717684352
effective search space used: 8717684352
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0316.5


