

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0314.4
(199 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
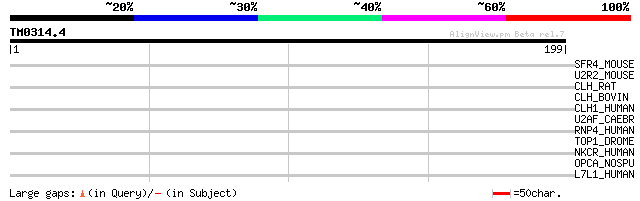
SFR4_MOUSE (Q8VE97) Splicing factor, arginine/serine-rich 4 39 0.010
U2R2_MOUSE (Q62377) U2 small nuclear ribonucleoprotein auxiliary... 33 0.56
CLH_RAT (P11442) Clathrin heavy chain 30 3.7
CLH_BOVIN (P49951) Clathrin heavy chain 30 3.7
CLH1_HUMAN (Q00610) Clathrin heavy chain 1 (CLH-17) 30 3.7
U2AF_CAEBR (P90727) Splicing factor U2AF 65 kDa subunit (U2 auxi... 30 4.8
RNP4_HUMAN (Q86U06) RNA-binding region containing protein 4 (Spl... 30 4.8
TOP1_DROME (P30189) DNA topoisomerase I (EC 5.99.1.2) 29 6.2
NKCR_HUMAN (P30414) NK-tumor recognition protein (Natural-killer... 29 6.2
OPCA_NOSPU (P48971) Putative OxPP cycle protein opcA 29 8.2
L7L1_HUMAN (Q9NQ29) Putative RNA-binding protein Luc7-like 1 (SR... 29 8.2
>SFR4_MOUSE (Q8VE97) Splicing factor, arginine/serine-rich 4
Length = 489
Score = 38.5 bits (88), Expect = 0.010
Identities = 23/59 (38%), Positives = 33/59 (54%), Gaps = 2/59 (3%)
Query: 1 RTRGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSR--ERFDHPHRVHLRAVTGRR 57
R+R RS + R R+ S +RD + SS S S + +DHSR R R H +A +G+R
Sbjct: 362 RSRSRSKSERSRKHSSKRDSKVSSSSSSSKKKKDTDHSRSPSRSVSKEREHAKAESGQR 420
Score = 29.6 bits (65), Expect = 4.8
Identities = 15/40 (37%), Positives = 21/40 (52%)
Query: 2 TRGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRER 41
+R RS + R++ SSHS SR+ SR HSR +
Sbjct: 192 SRSRSRSRHSRKSRSRSGSSKSSHSKSRSRSRSGSHSRSK 231
Score = 28.9 bits (63), Expect = 8.2
Identities = 21/57 (36%), Positives = 27/57 (46%), Gaps = 1/57 (1%)
Query: 2 TRGRSNTPRFRQASRERDREASSH-SGSRTGSRFSDHSRERFDHPHRVHLRAVTGRR 57
+R R + R R SR R R S S SR+GS S HS+ R H R+ + R
Sbjct: 179 SRRRRSYSRSRSHSRSRSRSRHSRKSRSRSGSSKSSHSKSRSRSRSGSHSRSKSRSR 235
>U2R2_MOUSE (Q62377) U2 small nuclear ribonucleoprotein auxiliary
factor 35 kDa subunit related-protein 2 (U2(RNU2) small
nuclear RNA auxillary factor 1-like 2)
Length = 462
Score = 32.7 bits (73), Expect = 0.56
Identities = 18/47 (38%), Positives = 25/47 (52%)
Query: 1 RTRGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRERFDHPHR 47
R R RS +P F + + E DR++SS + R+ SRER P R
Sbjct: 382 RRRRRSPSPDFYKRNGESDRKSSSRHRVKKSHRYGMKSRERRSSPSR 428
>CLH_RAT (P11442) Clathrin heavy chain
Length = 1675
Score = 30.0 bits (66), Expect = 3.7
Identities = 20/73 (27%), Positives = 28/73 (37%)
Query: 78 FFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPF 137
F Q G E + A+ G+A G E P +PF + F P QN F
Sbjct: 201 FAQFKMEGNAEESTLFCFAVRGQAGGKLHIIEVGTPPTGNQPFPKKAVDVFFPPEAQNDF 260
Query: 138 GPLLSIKQKGSVM 150
+ I +K V+
Sbjct: 261 PVAMQISEKHDVV 273
>CLH_BOVIN (P49951) Clathrin heavy chain
Length = 1675
Score = 30.0 bits (66), Expect = 3.7
Identities = 20/73 (27%), Positives = 28/73 (37%)
Query: 78 FFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPF 137
F Q G E + A+ G+A G E P +PF + F P QN F
Sbjct: 201 FAQFKMEGNAEESTLFCFAVRGQAGGKLHIIEVGTPPTGNQPFPKKAVDVFFPPEAQNDF 260
Query: 138 GPLLSIKQKGSVM 150
+ I +K V+
Sbjct: 261 PVAMQISEKHDVV 273
>CLH1_HUMAN (Q00610) Clathrin heavy chain 1 (CLH-17)
Length = 1674
Score = 30.0 bits (66), Expect = 3.7
Identities = 20/73 (27%), Positives = 28/73 (37%)
Query: 78 FFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPF 137
F Q G E + A+ G+A G E P +PF + F P QN F
Sbjct: 200 FAQFKMEGNAEESTLFCFAVRGQAGGKLHIIEVGTPPTGNQPFPKKAVDVFFPPEAQNDF 259
Query: 138 GPLLSIKQKGSVM 150
+ I +K V+
Sbjct: 260 PVAMQISEKHDVV 272
>U2AF_CAEBR (P90727) Splicing factor U2AF 65 kDa subunit (U2
auxiliary factor 65 kDa subunit) (U2 snRNP auxiliary
factor large subunit) (U2AF65)
Length = 488
Score = 29.6 bits (65), Expect = 4.8
Identities = 15/40 (37%), Positives = 18/40 (44%)
Query: 1 RTRGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRE 40
R+R R + R R R RDR + R G R D RE
Sbjct: 75 RSRDRRDRDRSRSRERRRDRSRDRNRDDRRGGRDDDRRRE 114
Score = 28.9 bits (63), Expect = 8.2
Identities = 18/41 (43%), Positives = 21/41 (50%), Gaps = 2/41 (4%)
Query: 1 RTRGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRER 41
R+R R R R+ SR RDR S SR R D SR+R
Sbjct: 60 RSRSRDRGDRDRKRSRSRDRRDRDRSRSR--ERRRDRSRDR 98
>RNP4_HUMAN (Q86U06) RNA-binding region containing protein 4
(Splicing factor SF2) (PP239)
Length = 439
Score = 29.6 bits (65), Expect = 4.8
Identities = 21/62 (33%), Positives = 28/62 (44%), Gaps = 3/62 (4%)
Query: 1 RTRGRSNTP---RFRQASRERDREASSHSGSRTGSRFSDHSRERFDHPHRVHLRAVTGRR 57
+ R RS+ R R SR+RDR +S SR+ R H +D H R+ RR
Sbjct: 61 KKRSRSHNKSRDRKRSRSRDRDRYRRRNSRSRSPGRQCRHRSRSWDRRHGSESRSRDHRR 120
Query: 58 VD 59
D
Sbjct: 121 ED 122
>TOP1_DROME (P30189) DNA topoisomerase I (EC 5.99.1.2)
Length = 972
Score = 29.3 bits (64), Expect = 6.2
Identities = 20/55 (36%), Positives = 30/55 (54%), Gaps = 10/55 (18%)
Query: 1 RTRGRSNTPRFRQAS--RERDREASSH-SGSRTGSRFSDHSRERF-------DHP 45
+ RG S++ R + +S R+++R +SSH S S + S S HS R DHP
Sbjct: 148 KDRGSSSSSRHKSSSSSRDKERSSSSHKSSSSSSSSKSKHSSSRHSSSSSSKDHP 202
>NKCR_HUMAN (P30414) NK-tumor recognition protein (Natural-killer
cells cyclophilin-related protein) (NK-TR protein)
Length = 1462
Score = 29.3 bits (64), Expect = 6.2
Identities = 22/46 (47%), Positives = 27/46 (57%), Gaps = 3/46 (6%)
Query: 1 RTRGRSNTPRFRQASRERDRE-ASSHSGSRTGSRFSDHSRERF-DH 44
R+R RS+ R S R R ASSHS SR+ S S HSR ++ DH
Sbjct: 691 RSRSRSSRSRSYSRSYTRSRSLASSHSRSRSPSSRS-HSRNKYSDH 735
>OPCA_NOSPU (P48971) Putative OxPP cycle protein opcA
Length = 465
Score = 28.9 bits (63), Expect = 8.2
Identities = 26/123 (21%), Positives = 48/123 (38%), Gaps = 13/123 (10%)
Query: 86 EEERLEMVMIAMEGKALGWFQW-----WEEQAPERAWEPFKHALIRRFQPTLVQ----NP 136
E + L + + G L W W+E E P + A +R + NP
Sbjct: 261 ESDLLRLEELVEAGVPLADLNWRRLASWQELTAEAYDSPKRRAALREIDRVTIDYEKGNP 320
Query: 137 FGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFLNGLQEEIRAEMKLYPAADLA 196
LL + +A R ++ + D +I + F++ Q ++ AE+ P AD+
Sbjct: 321 AQALLFL----GWLASRLEWQPVSYQKESGDYDITRIHFISQDQRQVEAELAGVPVADVG 376
Query: 197 GLM 199
++
Sbjct: 377 DII 379
>L7L1_HUMAN (Q9NQ29) Putative RNA-binding protein Luc7-like 1
(SR+89) (Putative SR protein LUC7B1)
Length = 371
Score = 28.9 bits (63), Expect = 8.2
Identities = 14/30 (46%), Positives = 17/30 (56%)
Query: 12 RQASRERDREASSHSGSRTGSRFSDHSRER 41
R+ RER+ S SGSRT R SR+R
Sbjct: 246 RREEREREERLSRRSGSRTRDRRRSRSRDR 275
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.136 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,693,224
Number of Sequences: 164201
Number of extensions: 918727
Number of successful extensions: 2263
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 2224
Number of HSP's gapped (non-prelim): 42
length of query: 199
length of database: 59,974,054
effective HSP length: 105
effective length of query: 94
effective length of database: 42,732,949
effective search space: 4016897206
effective search space used: 4016897206
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0314.4


