

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0304.15
(697 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
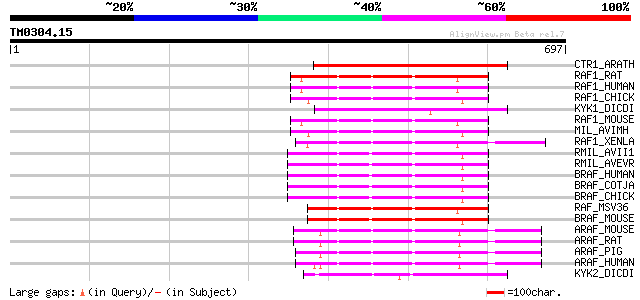
Score E
Sequences producing significant alignments: (bits) Value
CTR1_ARATH (Q05609) Serine/threonine-protein kinase CTR1 (EC 2.7... 308 2e-83
RAF1_RAT (P11345) RAF proto-oncogene serine/threonine-protein ki... 167 1e-40
RAF1_HUMAN (P04049) RAF proto-oncogene serine/threonine-protein ... 165 3e-40
RAF1_CHICK (P05625) RAF proto-oncogene serine/threonine-protein ... 164 7e-40
KYK1_DICDI (P18160) Non-receptor tyrosine kinase spore lysis A (... 163 1e-39
RAF1_MOUSE (Q99N57) RAF proto-oncogene serine/threonine-protein ... 162 2e-39
MIL_AVIMH (P00531) Serine/threonine-protein kinase transforming ... 162 2e-39
RAF1_XENLA (P09560) RAF proto-oncogene serine/threonine-protein ... 160 8e-39
RMIL_AVII1 (P10533) Serine/threonine-protein kinase transforming... 158 4e-38
RMIL_AVEVR (P27966) Serine/threonine-protein kinase transforming... 158 4e-38
BRAF_HUMAN (P15056) B-Raf proto-oncogene serine/threonine-protei... 158 4e-38
BRAF_COTJA (P34908) B-Raf proto-oncogene serine/threonine-protei... 158 4e-38
BRAF_CHICK (Q04982) B-Raf proto-oncogene serine/threonine-protei... 158 4e-38
RAF_MSV36 (P00532) Serine/threonine-protein kinase transforming ... 157 7e-38
BRAF_MOUSE (P28028) B-Raf proto-oncogene serine/threonine-protei... 155 3e-37
ARAF_MOUSE (P04627) A-Raf proto-oncogene serine/threonine-protei... 155 3e-37
ARAF_RAT (P14056) A-Raf proto-oncogene serine/threonine-protein ... 154 8e-37
ARAF_PIG (O19004) A-Raf proto-oncogene serine/threonine-protein ... 154 1e-36
ARAF_HUMAN (P10398) A-Raf proto-oncogene serine/threonine-protei... 152 3e-36
KYK2_DICDI (P18161) Tyrosine-protein kinase 2 (EC 2.7.1.112) (Fr... 149 2e-35
>CTR1_ARATH (Q05609) Serine/threonine-protein kinase CTR1 (EC
2.7.1.37)
Length = 821
Score = 308 bits (790), Expect = 2e-83
Identities = 148/246 (60%), Positives = 188/246 (76%), Gaps = 2/246 (0%)
Query: 382 DCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKR 441
D +I W DL+++++IG GS+ V+ W+GSDVA+K+ + E + ++ +E+ IMKR
Sbjct: 543 DMDIPWCDLNIKEKIGAGSFGTVHRAEWHGSDVAVKILMEQDFHAERVNEFLREVAIMKR 602
Query: 442 LRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSN--QTLDIRRRLRMALDIARGM 499
LRHPN++LFMGAV L+IVTE L RGSL++ LHKS + LD RRRL MA D+A+GM
Sbjct: 603 LRHPNIVLFMGAVTQPPNLSIVTEYLSRGSLYRLLHKSGAREQLDERRRLSMAYDVAKGM 662
Query: 500 NYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEI 559
NYLH+RNPPIVHRDLKS NLLVDK +TVKV DFGLSRLK +T L++KS GTP+WMAPE+
Sbjct: 663 NYLHNRNPPIVHRDLKSPNLLVDKKYTVKVCDFGLSRLKASTFLSSKSAAGTPEWMAPEV 722
Query: 560 LRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVAS 619
LR+EPSNEKSDVYS+GV+LWEL T PW NLN QVV VGF +RL++P L+P VA+
Sbjct: 723 LRDEPSNEKSDVYSFGVILWELATLQQPWGNLNPAQVVAAVGFKCKRLEIPRNLNPQVAA 782
Query: 620 IINDCW 625
II CW
Sbjct: 783 IIEGCW 788
>RAF1_RAT (P11345) RAF proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (Raf-1) (C-RAF) (cRaf)
Length = 648
Score = 167 bits (422), Expect = 1e-40
Identities = 99/260 (38%), Positives = 157/260 (60%), Gaps = 15/260 (5%)
Query: 353 QGRLQLPSSRESV---GSHESSS--SKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHG 407
Q + +P+ RE G+ E + +G DS EI ++ L IG GS+ VY G
Sbjct: 307 QPKTPVPAQRERAPGSGTQEKNKIRPRGQRDSSYYWEIEASEVMLSTRIGSGSFGTVYKG 366
Query: 408 IWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELL 467
W+G DVA+K+ T E LQ ++ E+ ++++ RH N+LLFMG + + + LAIVT+
Sbjct: 367 KWHG-DVAVKILKVVDPTPEQLQAFRNEVAVLRKTRHVNILLFMGYM-TKDNLAIVTQWC 424
Query: 468 PRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV 527
SL+K LH + + + +A A+GM+YLH +N I+HRD+KS+N+ + + TV
Sbjct: 425 EGSSLYKHLHVQETKFQMFQLIDIARQTAQGMDYLHAKN--IIHRDMKSNNIFLHEGLTV 482
Query: 528 KVGDFGLSRLKD--TTLLTTKSGRGTPQWMAPEILR---NEPSNEKSDVYSYGVVLWELM 582
K+GDFGL+ +K + + G+ WMAPE++R N P + +SDVYSYG+VL+ELM
Sbjct: 483 KIGDFGLATVKSRWSGSQQVEQPTGSVLWMAPEVIRMQDNNPFSFQSDVYSYGIVLYELM 542
Query: 583 TQSIPWENLNSL-QVVGVVG 601
T +P+ ++N+ Q++ +VG
Sbjct: 543 TGELPYSHINNRDQIIFMVG 562
>RAF1_HUMAN (P04049) RAF proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (Raf-1) (C-RAF) (cRaf)
Length = 648
Score = 165 bits (418), Expect = 3e-40
Identities = 98/260 (37%), Positives = 156/260 (59%), Gaps = 15/260 (5%)
Query: 353 QGRLQLPSSRESV---GSHESSS--SKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHG 407
Q + +P+ RE G+ E + +G DS EI ++ L IG GS+ VY G
Sbjct: 307 QPKTPVPAQRERAPVSGTQEKNKIRPRGQRDSSYYWEIEASEVMLSTRIGSGSFGTVYKG 366
Query: 408 IWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELL 467
W+G DVA+K+ T E Q ++ E+ ++++ RH N+LLFMG + + + LAIVT+
Sbjct: 367 KWHG-DVAVKILKVVDPTPEQFQAFRNEVAVLRKTRHVNILLFMGYM-TKDNLAIVTQWC 424
Query: 468 PRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV 527
SL+K LH + + + +A A+GM+YLH +N I+HRD+KS+N+ + + TV
Sbjct: 425 EGSSLYKHLHVQETKFQMFQLIDIARQTAQGMDYLHAKN--IIHRDMKSNNIFLHEGLTV 482
Query: 528 KVGDFGLSRLKD--TTLLTTKSGRGTPQWMAPEILR---NEPSNEKSDVYSYGVVLWELM 582
K+GDFGL+ +K + + G+ WMAPE++R N P + +SDVYSYG+VL+ELM
Sbjct: 483 KIGDFGLATVKSRWSGSQQVEQPTGSVLWMAPEVIRMQDNNPFSFQSDVYSYGIVLYELM 542
Query: 583 TQSIPWENLNSL-QVVGVVG 601
T +P+ ++N+ Q++ +VG
Sbjct: 543 TGELPYSHINNRDQIIFMVG 562
>RAF1_CHICK (P05625) RAF proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (RAF-1) (C-RAF) (MIL proto-oncogene
serine/threonine-protein kinase) (C-MIL)
Length = 647
Score = 164 bits (415), Expect = 7e-40
Identities = 98/260 (37%), Positives = 156/260 (59%), Gaps = 15/260 (5%)
Query: 353 QGRLQLPSSRESV-GSHESSSSK----GDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHG 407
Q + +P+ RE G++ +K G DS EI ++ L IG GS+ VY G
Sbjct: 307 QPKTPVPAQRERAPGTNTQEKNKIRPRGQRDSSYYWEIEASEVMLSTRIGSGSFGTVYKG 366
Query: 408 IWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELL 467
W+G DVA+K+ T E Q ++ E+ ++++ RH N+LLFMG + + + LAIVT+
Sbjct: 367 KWHG-DVAVKILKVVDPTPEQFQAFRNEVAVLRKTRHVNILLFMGYM-TKDNLAIVTQWC 424
Query: 468 PRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV 527
SL+K LH + + + +A A+GM+YLH +N I+HRD+KS+N+ + + TV
Sbjct: 425 EGSSLYKHLHVQETKFQMFQLIDIARQTAQGMDYLHAKN--IIHRDMKSNNIFLHEGLTV 482
Query: 528 KVGDFGLSRLKD--TTLLTTKSGRGTPQWMAPEILRNEPSNE---KSDVYSYGVVLWELM 582
K+GDFGL+ +K + + G+ WMAPE++R + SN +SDVYSYG+VL+ELM
Sbjct: 483 KIGDFGLATVKSRWSGSQQVEQPTGSILWMAPEVIRMQDSNPFSFQSDVYSYGIVLYELM 542
Query: 583 TQSIPWENLNSL-QVVGVVG 601
T +P+ ++N+ Q++ +VG
Sbjct: 543 TGELPYSHINNRDQIIFMVG 562
>KYK1_DICDI (P18160) Non-receptor tyrosine kinase spore lysis A (EC
2.7.1.112) (Tyrosine-protein kinase 1)
Length = 1584
Score = 163 bits (413), Expect = 1e-39
Identities = 96/260 (36%), Positives = 145/260 (54%), Gaps = 18/260 (6%)
Query: 384 EIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGY-TEETLQDYKKEIDIMKRL 442
EI + +L IG+G + V G W +DVAIK+ + + + T+ +L ++ E+ I+ +L
Sbjct: 1283 EIDFNELEFGQTIGKGFFGEVKRGYWRETDVAIKIIYRDQFKTKSSLVMFQNEVGILSKL 1342
Query: 443 RHPNVLLFMGAVYS--LERLAIVTELLPRGSL--FKTLHKSNQTLDIRRRLRMALDIARG 498
RHPNV+ F+GA + + IVTE + GSL F T H + + RL++ALDIA+G
Sbjct: 1343 RHPNVVQFLGACTAGGEDHHCIVTEWMGGGSLRQFLTDHFNLLEQNPHIRLKLALDIAKG 1402
Query: 499 MNYLHHRNPPIVHRDLKSSNLLVDKNWT-------------VKVGDFGLSRLKDTTLLTT 545
MNYLH PPI+HRDL S N+L+D N K+ DFGLSRLK
Sbjct: 1403 MNYLHGWTPPILHRDLSSRNILLDHNIDPKNPVVSSRQDIKCKISDFGLSRLKKEQASQM 1462
Query: 546 KSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDR 605
G +MAPE+ + + ++EKSDVYSYG+VL+EL+T P +++ +++ + +
Sbjct: 1463 TQSVGCIPYMAPEVFKGDSNSEKSDVYSYGMVLFELLTSDEPQQDMKPMKMAHLAAYESY 1522
Query: 606 RLDLPEGLDPHVASIINDCW 625
R +P I+ CW
Sbjct: 1523 RPPIPLTTSSKWKEILTQCW 1542
>RAF1_MOUSE (Q99N57) RAF proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (Raf-1) (C-RAF) (cRaf)
Length = 648
Score = 162 bits (411), Expect = 2e-39
Identities = 97/260 (37%), Positives = 156/260 (59%), Gaps = 15/260 (5%)
Query: 353 QGRLQLPSSRESV---GSHESSS--SKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHG 407
Q + +P+ RE G+ E + +G DS EI ++ L IG GS+ VY G
Sbjct: 307 QPKTPVPAQRERAPGSGTQEKNKIRPRGQRDSSYYWEIEASEVMLSTRIGSGSFGTVYKG 366
Query: 408 IWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELL 467
W+G DVA+K+ T E LQ ++ E+ ++++ RH N+LLFMG + + + LAIVT+
Sbjct: 367 KWHG-DVAVKILKVVDPTPEQLQAFRNEVAVLRKTRHVNILLFMGYM-TKDNLAIVTQWC 424
Query: 468 PRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV 527
SL+K LH + + + +A A+GM+YLH +N I+HRD+KS+N+ + + TV
Sbjct: 425 EGSSLYKHLHVQETKFQMFQLIDIARQTAQGMDYLHAKN--IIHRDMKSNNIFLHEGLTV 482
Query: 528 KVGDFGLSRLKD--TTLLTTKSGRGTPQWMAPEILR---NEPSNEKSDVYSYGVVLWELM 582
K+GDFGL+ +K + + G+ WMAPE++R + P + +SDVYSYG+VL+ELM
Sbjct: 483 KIGDFGLATVKSRWSGSQQVEQPTGSVLWMAPEVIRMQDDNPFSFQSDVYSYGIVLYELM 542
Query: 583 TQSIPWENLNSL-QVVGVVG 601
+P+ ++N+ Q++ +VG
Sbjct: 543 AGELPYAHINNRDQIIFMVG 562
>MIL_AVIMH (P00531) Serine/threonine-protein kinase transforming
protein mil (EC 2.7.1.37)
Length = 380
Score = 162 bits (411), Expect = 2e-39
Identities = 98/260 (37%), Positives = 155/260 (58%), Gaps = 15/260 (5%)
Query: 353 QGRLQLPSSRESV-GSHESSSSK----GDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHG 407
Q + +P+ RE G++ +K G DS EI ++ L IG GS+ VY G
Sbjct: 40 QPKTPVPAQRERAPGTNTQEKNKIRPRGQRDSSYYWEIEASEVLLSTRIGSGSFGTVYKG 99
Query: 408 IWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELL 467
W+G DVA+K+ T E Q ++ E+ ++++ RH N+LLFMG + + + LAIVT+
Sbjct: 100 KWHG-DVAVKILKVVDPTPEQFQAFRNEVAVLRKTRHVNILLFMGYM-TKDNLAIVTQWC 157
Query: 468 PRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV 527
SL+K LH + + + +A A+GM+YLH +N I+HRD+KS+N+ + TV
Sbjct: 158 EGSSLYKHLHVQETKFQMFQLIDIARQTAQGMDYLHAKN--IIHRDMKSNNIFLHGGLTV 215
Query: 528 KVGDFGLSRLKD--TTLLTTKSGRGTPQWMAPEILRNEPSNE---KSDVYSYGVVLWELM 582
K+GDFGL+ +K + + G+ WMAPE++R + SN +SDVYSYG+VL+ELM
Sbjct: 216 KIGDFGLATVKSRWSGSQQVEQPTGSILWMAPEVIRMQDSNPFSFQSDVYSYGIVLYELM 275
Query: 583 TQSIPWENLNSL-QVVGVVG 601
T +P+ ++N+ Q++ +VG
Sbjct: 276 TGELPYSHINNRDQIIFMVG 295
>RAF1_XENLA (P09560) RAF proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (C-RAF)
Length = 638
Score = 160 bits (406), Expect = 8e-39
Identities = 109/327 (33%), Positives = 178/327 (54%), Gaps = 25/327 (7%)
Query: 359 PSSRESVGSHESSS-----SKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSD 413
P+ RE S ++G DS EI ++ L IG GS+ VY G W+G D
Sbjct: 304 PTHREKAASSTGQEKNKIRARGQRDSSYYWEIIASEVMLSSRIGSGSFGTVYKGKWHG-D 362
Query: 414 VAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLF 473
VA+K+ T E LQ ++ E+ ++++ RH N+LLFMG + + + LAIVT+ SL+
Sbjct: 363 VAVKILKVTDPTPEQLQAFRNEVAVLRKTRHVNILLFMGYM-TKDNLAIVTQWCEGSSLY 421
Query: 474 KTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFG 533
LH + + + + +A A+GM+YLH +N I+HRD+KS+N+ + + TVK+GDFG
Sbjct: 422 YHLHVLDTKFQMFQLIDIARQTAQGMDYLHAKN--IIHRDMKSNNIFLHEGLTVKIGDFG 479
Query: 534 LSRLKD--TTLLTTKSGRGTPQWMAPEILR---NEPSNEKSDVYSYGVVLWELMTQSIPW 588
L+ +K + + G+ WMAPE++R N P + +SDVYSYG+VL+ELMT +P+
Sbjct: 480 LATVKTRWSGSQQVEQLTGSILWMAPEVIRMQDNNPFSFQSDVYSYGIVLYELMTGELPY 539
Query: 589 ENLNSL-QVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYS-SLQLYYGRNHLLP 646
++ Q++ +VG G+ P ++ + +C + + + S++ L P
Sbjct: 540 SHIRDRDQIIFLVG--------RGGVVPDLSKLYKNCPKAMKRLVADSIKKLRDERPLFP 591
Query: 647 QTESLLVLNMSSDPE-QRPSFEELIQR 672
Q S + L S P+ R + E + R
Sbjct: 592 QILSSIELLQHSLPKINRSALEPSLHR 618
>RMIL_AVII1 (P10533) Serine/threonine-protein kinase transforming
protein Rmil (EC 2.7.1.37)
Length = 367
Score = 158 bits (400), Expect = 4e-38
Identities = 97/260 (37%), Positives = 155/260 (59%), Gaps = 12/260 (4%)
Query: 349 VQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGI 408
+QK G + S S + G DS D EI + + IG GS+ VY G
Sbjct: 26 LQKSPGPQRERKSSSSSEDRNRMKTLGRRDSSDDWEIPDGQITVGQRIGSGSFGTVYKGK 85
Query: 409 WNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLE-RLAIVTELL 467
W+G DVA+K+ T + LQ +K E+ ++++ RH N+LLFMG YS + +LAIVT+
Sbjct: 86 WHG-DVAVKMLNVTAPTPQQLQAFKNEVGVLRKTRHVNILLFMG--YSTKPQLAIVTQWC 142
Query: 468 PRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV 527
SL+ LH ++ + + +A A+GM+YLH ++ I+HRDLKS+N+ + ++ TV
Sbjct: 143 EGSSLYHHLHIIETKFEMIKLIDIARQTAQGMDYLHAKS--IIHRDLKSNNIFLHEDLTV 200
Query: 528 KVGDFGLSRLKDTTLLTTKSGR--GTPQWMAPEILRNEPSNE---KSDVYSYGVVLWELM 582
K+GDFGL+ +K + + + G+ WMAPE++R + N +SDVY++G+VL+ELM
Sbjct: 201 KIGDFGLATVKSRWSGSHQFEQLSGSILWMAPEVIRMQDKNPYSFQSDVYAFGIVLYELM 260
Query: 583 TQSIPWENLNSL-QVVGVVG 601
T +P+ N+N+ Q++ +VG
Sbjct: 261 TGQLPYSNINNRDQIIFMVG 280
>RMIL_AVEVR (P27966) Serine/threonine-protein kinase transforming
protein Rmil (EC 2.7.1.37)
Length = 450
Score = 158 bits (400), Expect = 4e-38
Identities = 97/260 (37%), Positives = 155/260 (59%), Gaps = 12/260 (4%)
Query: 349 VQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGI 408
+QK G + S S + G DS D EI + + IG GS+ VY G
Sbjct: 42 LQKSPGPQRERKSSSSSEDRNRMKTLGRRDSSDDWEIPDGQITVGQRIGSGSFGTVYKGK 101
Query: 409 WNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLE-RLAIVTELL 467
W+G DVA+K+ T + LQ +K E+ ++++ RH N+LLFMG YS + +LAIVT+
Sbjct: 102 WHG-DVAVKMLNVTAPTPQQLQAFKNEVGVLRKTRHVNILLFMG--YSTKPQLAIVTQWC 158
Query: 468 PRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV 527
SL+ LH ++ + + +A A+GM+YLH ++ I+HRDLKS+N+ + ++ TV
Sbjct: 159 EGSSLYHHLHIIETKFEMIKLIDIARQTAQGMDYLHAKS--IIHRDLKSNNIFLHEDLTV 216
Query: 528 KVGDFGLSRLKDTTLLTTKSGR--GTPQWMAPEILRNEPSNE---KSDVYSYGVVLWELM 582
K+GDFGL+ +K + + + G+ WMAPE++R + N +SDVY++G+VL+ELM
Sbjct: 217 KIGDFGLATVKSRWSGSHQFEQLSGSILWMAPEVIRMQDKNPYSFQSDVYAFGIVLYELM 276
Query: 583 TQSIPWENLNSL-QVVGVVG 601
T +P+ N+N+ Q++ +VG
Sbjct: 277 TGQLPYSNINNRDQIIFMVG 296
>BRAF_HUMAN (P15056) B-Raf proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (p94) (v-Raf murine sarcoma viral
oncogene homolog B1)
Length = 766
Score = 158 bits (400), Expect = 4e-38
Identities = 97/260 (37%), Positives = 155/260 (59%), Gaps = 12/260 (4%)
Query: 349 VQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGI 408
+QK G + S S + G DS D EI + + IG GS+ VY G
Sbjct: 416 LQKSPGPQRERKSSSSSEDRNRMKTLGRRDSSDDWEIPDGQITVGQRIGSGSFGTVYKGK 475
Query: 409 WNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLE-RLAIVTELL 467
W+G DVA+K+ T + LQ +K E+ ++++ RH N+LLFMG YS + +LAIVT+
Sbjct: 476 WHG-DVAVKMLNVTAPTPQQLQAFKNEVGVLRKTRHVNILLFMG--YSTKPQLAIVTQWC 532
Query: 468 PRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV 527
SL+ LH ++ + + +A A+GM+YLH ++ I+HRDLKS+N+ + ++ TV
Sbjct: 533 EGSSLYHHLHIIETKFEMIKLIDIARQTAQGMDYLHAKS--IIHRDLKSNNIFLHEDLTV 590
Query: 528 KVGDFGLSRLKDTTLLTTKSGR--GTPQWMAPEILRNEPSNE---KSDVYSYGVVLWELM 582
K+GDFGL+ +K + + + G+ WMAPE++R + N +SDVY++G+VL+ELM
Sbjct: 591 KIGDFGLATVKSRWSGSHQFEQLSGSILWMAPEVIRMQDKNPYSFQSDVYAFGIVLYELM 650
Query: 583 TQSIPWENLNSL-QVVGVVG 601
T +P+ N+N+ Q++ +VG
Sbjct: 651 TGQLPYSNINNRDQIIFMVG 670
>BRAF_COTJA (P34908) B-Raf proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (RMIL serine/threonine-protein
kinase) (c-RMIL)
Length = 807
Score = 158 bits (400), Expect = 4e-38
Identities = 97/260 (37%), Positives = 155/260 (59%), Gaps = 12/260 (4%)
Query: 349 VQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGI 408
+QK G + S S + G DS D EI + + IG GS+ VY G
Sbjct: 456 LQKSPGPQRERKSSSSSEDRNRMKTLGRRDSSDDWEIPDGQITVGQRIGSGSFGTVYKGK 515
Query: 409 WNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLE-RLAIVTELL 467
W+G DVA+K+ T + LQ +K E+ ++++ RH N+LLFMG YS + +LAIVT+
Sbjct: 516 WHG-DVAVKMLNVTAPTPQQLQAFKNEVGVLRKTRHVNILLFMG--YSTKPQLAIVTQWC 572
Query: 468 PRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV 527
SL+ LH ++ + + +A A+GM+YLH ++ I+HRDLKS+N+ + ++ TV
Sbjct: 573 EGSSLYHHLHIIETKFEMIKLIDIARQTAQGMDYLHAKS--IIHRDLKSNNIFLHEDLTV 630
Query: 528 KVGDFGLSRLKDTTLLTTKSGR--GTPQWMAPEILRNEPSNE---KSDVYSYGVVLWELM 582
K+GDFGL+ +K + + + G+ WMAPE++R + N +SDVY++G+VL+ELM
Sbjct: 631 KIGDFGLATVKSRWSGSHQFEQLSGSILWMAPEVIRMQDKNPYSFQSDVYAFGIVLYELM 690
Query: 583 TQSIPWENLNSL-QVVGVVG 601
T +P+ N+N+ Q++ +VG
Sbjct: 691 TGQLPYSNINNRDQIIFMVG 710
>BRAF_CHICK (Q04982) B-Raf proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (RMIL serine/threonine-protein
kinase) (c-RMIL)
Length = 806
Score = 158 bits (400), Expect = 4e-38
Identities = 97/260 (37%), Positives = 155/260 (59%), Gaps = 12/260 (4%)
Query: 349 VQKQQGRLQLPSSRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGI 408
+QK G + S S + G DS D EI + + IG GS+ VY G
Sbjct: 456 LQKSPGPQRERKSSSSSEDRNRMKTLGRRDSSDDWEIPDGQITVGQRIGSGSFGTVYKGK 515
Query: 409 WNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLE-RLAIVTELL 467
W+G DVA+K+ T + LQ +K E+ ++++ RH N+LLFMG YS + +LAIVT+
Sbjct: 516 WHG-DVAVKMLNVTAPTPQQLQAFKNEVGVLRKTRHVNILLFMG--YSTKPQLAIVTQWC 572
Query: 468 PRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV 527
SL+ LH ++ + + +A A+GM+YLH ++ I+HRDLKS+N+ + ++ TV
Sbjct: 573 EGSSLYHHLHIIETKFEMIKLIDIARQTAQGMDYLHAKS--IIHRDLKSNNIFLHEDLTV 630
Query: 528 KVGDFGLSRLKDTTLLTTKSGR--GTPQWMAPEILRNEPSNE---KSDVYSYGVVLWELM 582
K+GDFGL+ +K + + + G+ WMAPE++R + N +SDVY++G+VL+ELM
Sbjct: 631 KIGDFGLATVKSRWSGSHQFEQLSGSILWMAPEVIRMQDKNPYSFQSDVYAFGIVLYELM 690
Query: 583 TQSIPWENLNSL-QVVGVVG 601
T +P+ N+N+ Q++ +VG
Sbjct: 691 TGQLPYSNINNRDQIIFMVG 710
>RAF_MSV36 (P00532) Serine/threonine-protein kinase transforming
protein raf (EC 2.7.1.37)
Length = 323
Score = 157 bits (398), Expect = 7e-38
Identities = 89/234 (38%), Positives = 145/234 (61%), Gaps = 10/234 (4%)
Query: 374 KGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYK 433
+G DS ++ ++ L IG GS+ VY G W+G DVA+K+ T E LQ ++
Sbjct: 8 RGQRDSSYYWKMEASEVMLSTRIGSGSFGTVYKGKWHG-DVAVKILKVVDPTPEQLQAFR 66
Query: 434 KEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMAL 493
E+ ++++ RH N+LLFMG + + + LAIVT+ SL+K LH + + + +A
Sbjct: 67 NEVAVLRKTRHVNILLFMGYM-TKDNLAIVTQWCEGSSLYKHLHVQETKFQMFQLIDIAR 125
Query: 494 DIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKD--TTLLTTKSGRGT 551
A+GM+YLH +N I+HRD+KS+N+ + + TVK+GDFGL+ +K + + G+
Sbjct: 126 QTAQGMDYLHAKN--IIHRDMKSNNIFLHEGLTVKIGDFGLATVKSRWSGSQQVEQPTGS 183
Query: 552 PQWMAPEILR---NEPSNEKSDVYSYGVVLWELMTQSIPWENLNSL-QVVGVVG 601
WMAPE++R + P + +SDVYSYG+VL+ELM +P+ ++N+ Q++ +VG
Sbjct: 184 VLWMAPEVIRMQDDNPFSFQSDVYSYGIVLYELMAGELPYAHINNRDQIIFMVG 237
>BRAF_MOUSE (P28028) B-Raf proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (Fragment)
Length = 328
Score = 155 bits (392), Expect = 3e-37
Identities = 92/234 (39%), Positives = 147/234 (62%), Gaps = 12/234 (5%)
Query: 375 GDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKK 434
G DS D EI + + IG GS+ VY G W+G DVA+K+ T + LQ +K
Sbjct: 4 GRRDSSDDWEIPDGQITVGQRIGSGSFGTVYKGKWHG-DVAVKMLNVTAPTPQQLQAFKN 62
Query: 435 EIDIMKRLRHPNVLLFMGAVYSLE-RLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMAL 493
E+ ++++ RH N+LLFMG YS + +LAIVT+ SL+ LH ++ + + +A
Sbjct: 63 EVGVLRKTRHVNILLFMG--YSTKPQLAIVTQWCEGSSLYHHLHIIETKFEMIKLIDIAR 120
Query: 494 DIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGR--GT 551
A+GM+YLH ++ I+HRDLKS+N+ + ++ TVK+GDFGL+ +K + + + G+
Sbjct: 121 QTAQGMDYLHAKS--IIHRDLKSNNIFLHEDLTVKIGDFGLATVKSRWSGSHQFEQLSGS 178
Query: 552 PQWMAPEILRNEPSNE---KSDVYSYGVVLWELMTQSIPWENLNSL-QVVGVVG 601
WMAPE++R + N +SDVY++G+VL+ELMT +P+ N+N+ Q++ +VG
Sbjct: 179 ILWMAPEVIRMQDKNPYSFQSDVYAFGIVLYELMTGQLPYSNINNRDQIIFMVG 232
>ARAF_MOUSE (P04627) A-Raf proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37)
Length = 604
Score = 155 bits (392), Expect = 3e-37
Identities = 108/323 (33%), Positives = 174/323 (53%), Gaps = 23/323 (7%)
Query: 357 QLPSSRESVGSHESSSSKGDNDSVVDCEIHWE----DLHLRDEIGQGSYAVVYHGIWNGS 412
+LPS + S K N D +WE ++ L IG GS+ V+ G W+G
Sbjct: 271 KLPSEQRERKSLADEKKKVKNLGYRDSGYYWEVPPSEVQLLKRIGTGSFGTVFRGRWHG- 329
Query: 413 DVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSL 472
DVA+KV T E Q +K E+ ++++ RH N+LLFMG + + AI+T+ SL
Sbjct: 330 DVAVKVLKVAQPTAEQAQAFKNEMQVLRKTRHVNILLFMGFM-TRPGFAIITQWCEGSSL 388
Query: 473 FKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDF 532
+ LH ++ D+ + + +A A+GM+YLH +N I+HRDLKS+N+ + + TVK+GDF
Sbjct: 389 YHHLHVADTRFDMVQLIDVARQTAQGMDYLHAKN--IIHRDLKSNNIFLHEGLTVKIGDF 446
Query: 533 GLSRLKD--TTLLTTKSGRGTPQWMAPEILRNE---PSNEKSDVYSYGVVLWELMTQSIP 587
GL+ +K + + G+ WMA E++R + P + +SDVY+YGVVL+ELMT S+P
Sbjct: 447 GLATVKTRWSGAQPLEQPSGSVLWMAAEVIRMQDPNPYSFQSDVYAYGVVLYELMTGSLP 506
Query: 588 WENLNSL-QVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSS-LQLYYGRNHLL 645
+ ++ S Q++ +VG L P ++ I ++C + + + L+ L
Sbjct: 507 YSHIGSRDQIIFMVG--------RGYLSPDLSKIFSNCPKAMRRLLTDCLKFQREERPLF 558
Query: 646 PQTESLLVLNMSSDPEQRPSFEE 668
PQ + + L S P+ S E
Sbjct: 559 PQILATIELLQRSLPKIERSASE 581
>ARAF_RAT (P14056) A-Raf proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37)
Length = 604
Score = 154 bits (389), Expect = 8e-37
Identities = 107/323 (33%), Positives = 174/323 (53%), Gaps = 23/323 (7%)
Query: 357 QLPSSRESVGSHESSSSKGDNDSVVDCEIHWE----DLHLRDEIGQGSYAVVYHGIWNGS 412
+LP+ + S K N D +WE ++ L IG GS+ V+ G W+G
Sbjct: 271 KLPAEQRERKSLADEKKKVKNLGYRDSGYYWEVPPSEVQLLKRIGTGSFGTVFRGRWHG- 329
Query: 413 DVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSL 472
DVA+KV T E Q +K E+ ++++ RH N+LLFMG + + AI+T+ SL
Sbjct: 330 DVAVKVLKVAQPTAEQAQAFKNEMQVLRKTRHVNILLFMGFM-TRPGFAIITQWCEGSSL 388
Query: 473 FKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDF 532
+ LH ++ D+ + + +A A+GM+YLH +N I+HRDLKS+N+ + + TVK+GDF
Sbjct: 389 YHHLHVADTRFDMVQLIDVARQTAQGMDYLHAKN--IIHRDLKSNNIFLHEGLTVKIGDF 446
Query: 533 GLSRLKD--TTLLTTKSGRGTPQWMAPEILRNE---PSNEKSDVYSYGVVLWELMTQSIP 587
GL+ +K + + G+ WMA E++R + P + +SDVY+YGVVL+ELMT S+P
Sbjct: 447 GLATVKTRWSGAQPLEQPSGSVLWMAAEVIRMQDPNPYSFQSDVYAYGVVLYELMTGSLP 506
Query: 588 WENLNSL-QVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSS-LQLYYGRNHLL 645
+ ++ S Q++ +VG L P ++ I ++C + + + L+ L
Sbjct: 507 YSHIGSRDQIIFMVG--------RGYLSPDLSKIFSNCPKAMRRLLTDCLKFQREERPLF 558
Query: 646 PQTESLLVLNMSSDPEQRPSFEE 668
PQ + + L S P+ S E
Sbjct: 559 PQILATIELLQRSLPKIERSASE 581
>ARAF_PIG (O19004) A-Raf proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (A-Raf-1)
Length = 606
Score = 154 bits (388), Expect = 1e-36
Identities = 108/321 (33%), Positives = 172/321 (52%), Gaps = 23/321 (7%)
Query: 359 PSSRESVGSHESSSSKGDNDSVVDCEIHWE----DLHLRDEIGQGSYAVVYHGIWNGSDV 414
PS + S K N D +WE ++ L IG GS+ V+ G W+G DV
Sbjct: 275 PSEQRERKSLADDKKKVKNLGYRDSGYYWEVPPSEVQLLKRIGTGSFGTVFRGRWHG-DV 333
Query: 415 AIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFK 474
A+KV T E Q +K E+ ++++ RH N+LLFMG + + AI+T+ SL+
Sbjct: 334 AVKVLKVAQPTAEQAQAFKNEMQVLRKTRHVNILLFMGFM-TRPGFAIITQWCEGSSLYH 392
Query: 475 TLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGL 534
LH ++ D+ + + +A A+GM+YLH +N I+HRDLKS+N+ + + TVK+GDFGL
Sbjct: 393 HLHVADTRFDMVQLIDVARQTAQGMDYLHAKN--IIHRDLKSNNIFLHEGLTVKIGDFGL 450
Query: 535 SRLKD--TTLLTTKSGRGTPQWMAPEILRNE---PSNEKSDVYSYGVVLWELMTQSIPWE 589
+ +K + + G+ WMA E++R + P + +SDVY+YGVVL+ELMT S+P+
Sbjct: 451 ATVKTRWSGAQPLEQPSGSVLWMAAEVIRMQDPNPYSFQSDVYAYGVVLYELMTGSLPYS 510
Query: 590 NLNSL-QVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSS-LQLYYGRNHLLPQ 647
++ S Q++ +VG L P ++ I ++C + + S L+ L PQ
Sbjct: 511 HIGSRDQIIFMVG--------RGYLSPDLSKISSNCPKAMRRLLSDCLKFQREERPLFPQ 562
Query: 648 TESLLVLNMSSDPEQRPSFEE 668
+ + L S P+ S E
Sbjct: 563 ILATIELLQRSLPKIERSASE 583
>ARAF_HUMAN (P10398) A-Raf proto-oncogene serine/threonine-protein
kinase (EC 2.7.1.37) (A-raf-1) (Proto-oncogene Pks)
Length = 606
Score = 152 bits (384), Expect = 3e-36
Identities = 108/326 (33%), Positives = 173/326 (52%), Gaps = 28/326 (8%)
Query: 359 PSSRESVGSHESSSSKGDNDSVV-----DCEIHWE----DLHLRDEIGQGSYAVVYHGIW 409
P S+ E S D V D +WE ++ L IG GS+ V+ G W
Sbjct: 270 PHSKSPAEQRERKSLADDKKKVKNLGYRDSGYYWEVPPSEVQLLKRIGTGSFGTVFRGRW 329
Query: 410 NGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPR 469
+G DVA+KV + T E Q +K E+ ++++ RH N+LLFMG + + AI+T+
Sbjct: 330 HG-DVAVKVLKVSQPTAEQAQAFKNEMQVLRKTRHVNILLFMGFM-TRPGFAIITQWCEG 387
Query: 470 GSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKV 529
SL+ LH ++ D+ + + +A A+GM+YLH +N I+HRDLKS+N+ + + TVK+
Sbjct: 388 SSLYHHLHVADTRFDMVQLIDVARQTAQGMDYLHAKN--IIHRDLKSNNIFLHEGLTVKI 445
Query: 530 GDFGLSRLKD--TTLLTTKSGRGTPQWMAPEILRNE---PSNEKSDVYSYGVVLWELMTQ 584
GDFGL+ +K + + G+ WMA E++R + P + +SDVY+YGVVL+ELMT
Sbjct: 446 GDFGLATVKTRWSGAQPLEQPSGSVLWMAAEVIRMQDPNPYSFQSDVYAYGVVLYELMTG 505
Query: 585 SIPWENLNSL-QVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSS-LQLYYGRN 642
S+P+ ++ Q++ +VG L P ++ I ++C + + S L+
Sbjct: 506 SLPYSHIGCRDQIIFMVG--------RGYLSPDLSKISSNCPKAMRRLLSDCLKFQREER 557
Query: 643 HLLPQTESLLVLNMSSDPEQRPSFEE 668
L PQ + + L S P+ S E
Sbjct: 558 PLFPQILATIELLQRSLPKIERSASE 583
>KYK2_DICDI (P18161) Tyrosine-protein kinase 2 (EC 2.7.1.112)
(Fragment)
Length = 410
Score = 149 bits (376), Expect = 2e-35
Identities = 86/265 (32%), Positives = 147/265 (55%), Gaps = 16/265 (6%)
Query: 369 ESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEET 428
E S G+ + ++D D+ ++G+G+++ V+ G W G VAIK G E+
Sbjct: 91 ELKSILGEREYIIDIN----DIQFIQKVGEGAFSEVWEGWWKGIHVAIKKLKIIGDEEQF 146
Query: 429 LQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRR 488
+ + +E+ +K+ H N+++F+GA Y + I+TE + GSL+ LH N + +
Sbjct: 147 KERFIREVQNLKKGNHQNIVMFIGACY--KPACIITEYMAGGSLYNILHNPNSSTPKVKY 204
Query: 489 -----LRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLS--RLKDTT 541
L+MA D+A G+ +LH + IVHRDL S N+L+D+ +K+ DFGLS + ++ +
Sbjct: 205 SFPLVLKMATDMALGLLHLH--SITIVHRDLTSQNILLDELGNIKISDFGLSAEKSREGS 262
Query: 542 LLTTKSGRGTPQWMAPEILRNEPS-NEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVV 600
+ T G P+W PE+ +N +EK DVY + +V+WE++T IP+ +L+ Q V
Sbjct: 263 MTMTNGGICNPRWRPPELTKNLGHYSEKVDVYCFSLVVWEILTGEIPFSDLDGSQRSAQV 322
Query: 601 GFMDRRLDLPEGLDPHVASIINDCW 625
+ R +PE DP + ++ CW
Sbjct: 323 AYAGLRPPIPEYCDPELKLLLTQCW 347
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.132 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 83,060,265
Number of Sequences: 164201
Number of extensions: 3633449
Number of successful extensions: 13756
Number of sequences better than 10.0: 1656
Number of HSP's better than 10.0 without gapping: 1315
Number of HSP's successfully gapped in prelim test: 341
Number of HSP's that attempted gapping in prelim test: 9361
Number of HSP's gapped (non-prelim): 2124
length of query: 697
length of database: 59,974,054
effective HSP length: 117
effective length of query: 580
effective length of database: 40,762,537
effective search space: 23642271460
effective search space used: 23642271460
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0304.15


