

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
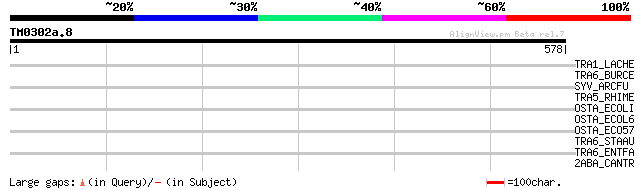
Query= TM0302a.8
(578 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TRA1_LACHE (P35880) Transposase for insertion sequence element I... 41 0.007
TRA6_BURCE (P24575) Transposase for insertion sequence element I... 34 0.91
SYV_ARCFU (O28059) Valyl-tRNA synthetase (EC 6.1.1.9) (Valine--t... 34 0.91
TRA5_RHIME (Q52873) Transposase for insertion sequence element I... 33 2.0
OSTA_ECOLI (P31554) Organic solvent tolerance protein precursor 33 2.7
OSTA_ECOL6 (Q8CWE6) Organic solvent tolerance protein precursor 33 2.7
OSTA_ECO57 (Q8XA13) Organic solvent tolerance protein precursor 33 2.7
TRA6_STAAU (P19775) Transposase for insertion sequence element I... 32 5.9
TRA6_ENTFA (P59787) Transposase for insertion sequence element I... 32 5.9
2ABA_CANTR (P53031) Protein phosphatase PP2A regulatory subunit ... 31 7.7
>TRA1_LACHE (P35880) Transposase for insertion sequence element
IS1201
Length = 369
Score = 41.2 bits (95), Expect = 0.007
Identities = 27/105 (25%), Positives = 45/105 (42%), Gaps = 5/105 (4%)
Query: 245 VAFDATYGKNRYKC----PFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMK 300
V DATY R + + G+ + + + NE IE + LL+
Sbjct: 160 VYLDATYVPLRRETFEREAVYIAIGIKPNGHKEVIDYCIAPNENIEVWTELLQSMKSRGL 219
Query: 301 GKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHVHI 345
+ LF ++DG MK A+ + +P AH + C H++ N V +
Sbjct: 220 EQVELF-LSDGVVGMKTALAKTYPQAHFQRCLVHVMRNICAKVRV 263
>TRA6_BURCE (P24575) Transposase for insertion sequence element
IS406
Length = 388
Score = 34.3 bits (77), Expect = 0.91
Identities = 25/100 (25%), Positives = 43/100 (43%), Gaps = 12/100 (12%)
Query: 248 DATYGKNRYK-----CPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKG- 301
DA Y K R + C ++ G+ K + + ++E + RFLE++
Sbjct: 166 DARYEKVRLEGRIVDCAVLIAVGIEASGKRRVLGCEVATSEAEINW----RRFLESLLAR 221
Query: 302 --KAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNA 339
K +I D +KAA + V P+ + C +H+ NA
Sbjct: 222 GLKGVTLIIADDHAGLKAARRAVLPSVPWQRCQFHLQQNA 261
>SYV_ARCFU (O28059) Valyl-tRNA synthetase (EC 6.1.1.9) (Valine--tRNA
ligase) (ValRS)
Length = 863
Score = 34.3 bits (77), Expect = 0.91
Identities = 47/210 (22%), Positives = 83/210 (39%), Gaps = 25/210 (11%)
Query: 127 IAAHRKMNESDIMHMNNMSRSGISTP-QIHNTFASQTGGYQK-------VRFSKGNMYNV 178
IA +++MN ++M G+ T ++ + + G + V F++ N+ +
Sbjct: 63 IARYKRMNGYEVMFPQGWDCHGLPTEVKVEEKYGIKKGDIPRDEFRRLCVEFTEENIAKM 122
Query: 179 QGRQRRMK-----KKE-----PE---KTDAEFAISYIRGLGSKD--PLLYCRHVANDDGN 223
+ RRM KE PE KT F Y +GL +D P+++C
Sbjct: 123 RETARRMGYSIDWSKEYITMYPEYYSKTQLSFVRMYNKGLIYRDYHPVVFCPRCETTIA- 181
Query: 224 LDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNE 283
L + + G ++LNY F D V T + C + + + NK I V
Sbjct: 182 LAEIEYRQGKTKLNYIKFDDDVIIATTRPELIPACVAIAVHPDDERNKHLIGKKVRVPTT 241
Query: 284 KIETYVWLLERFLEAMKGKAPLFVITDGDK 313
E V + + ++ G + + T GD+
Sbjct: 242 PYEVEV-IADEEVDPEFGTGVVMICTFGDR 270
>TRA5_RHIME (Q52873) Transposase for insertion sequence element
ISRM5
Length = 398
Score = 33.1 bits (74), Expect = 2.0
Identities = 23/86 (26%), Positives = 38/86 (43%), Gaps = 7/86 (8%)
Query: 261 VVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKG---KAPLFVITDGDKAMKA 317
++ G++ + I S + E + + FL +KG K V++D + A
Sbjct: 182 LIAVGIDWDGRRQILSVEMAGRESRSAW----KDFLVRLKGRGLKGVELVVSDDHAGLVA 237
Query: 318 AIKQVFPNAHHRLCAWHILDNARKHV 343
AI +V P A + C H L NA H+
Sbjct: 238 AIGEVIPEAAWQRCYVHFLRNALDHL 263
>OSTA_ECOLI (P31554) Organic solvent tolerance protein precursor
Length = 784
Score = 32.7 bits (73), Expect = 2.7
Identities = 33/134 (24%), Positives = 51/134 (37%), Gaps = 14/134 (10%)
Query: 108 KLFSDEHNHELEDDKMCGMIAAHRKMNESDIMHMNNMSRSGISTPQIHNTF----ASQTG 163
K++ DEH + DD + D + N+ + +S P N F S T
Sbjct: 296 KVYEDEHPN---DDSSRRWLFYWNHSGVMDQVWRFNVDYTKVSDPSYFNDFDNKYGSSTD 352
Query: 164 GYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDGN 223
GY +FS G Y VQ + K+ + + SY S +P L + ND G
Sbjct: 353 GYATQKFSVG--YAVQNFNATVSTKQFQVFSEQNTSSY-----SAEPQLDVNYYQNDVGP 405
Query: 224 LDRLFWSDGISQLN 237
D + + +N
Sbjct: 406 FDTRIYGQAVHFVN 419
>OSTA_ECOL6 (Q8CWE6) Organic solvent tolerance protein precursor
Length = 784
Score = 32.7 bits (73), Expect = 2.7
Identities = 33/134 (24%), Positives = 51/134 (37%), Gaps = 14/134 (10%)
Query: 108 KLFSDEHNHELEDDKMCGMIAAHRKMNESDIMHMNNMSRSGISTPQIHNTF----ASQTG 163
K++ DEH + DD + D + N+ + +S P N F S T
Sbjct: 296 KVYKDEHPN---DDSSRRWLFYWNHSGVMDQVWRFNVDYTKVSDPSYFNDFDNKYGSSTD 352
Query: 164 GYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDGN 223
GY +FS G Y VQ + K+ + + SY S +P L + ND G
Sbjct: 353 GYATQKFSVG--YAVQNFNATVSTKQFQVFSEQNTSSY-----SAEPQLDVNYYQNDVGP 405
Query: 224 LDRLFWSDGISQLN 237
D + + +N
Sbjct: 406 FDTRIYGQAVHFVN 419
>OSTA_ECO57 (Q8XA13) Organic solvent tolerance protein precursor
Length = 784
Score = 32.7 bits (73), Expect = 2.7
Identities = 33/134 (24%), Positives = 51/134 (37%), Gaps = 14/134 (10%)
Query: 108 KLFSDEHNHELEDDKMCGMIAAHRKMNESDIMHMNNMSRSGISTPQIHNTF----ASQTG 163
K++ DEH + DD + D + N+ + +S P N F S T
Sbjct: 296 KVYEDEHPN---DDSSRRWLFYWNHSGVMDQVWRFNVDYTKVSDPSYFNDFDNKYGSSTD 352
Query: 164 GYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAEFAISYIRGLGSKDPLLYCRHVANDDGN 223
GY +FS G Y VQ + K+ + + SY S +P L + ND G
Sbjct: 353 GYATQKFSVG--YAVQNFNATVSTKQFQVFSEQNTSSY-----SAEPQLDVNYYQNDVGP 405
Query: 224 LDRLFWSDGISQLN 237
D + + +N
Sbjct: 406 FDTRIYGQAVHFVN 419
>TRA6_STAAU (P19775) Transposase for insertion sequence element
IS256 in transposon Tn4001
Length = 390
Score = 31.6 bits (70), Expect = 5.9
Identities = 27/109 (24%), Positives = 44/109 (39%), Gaps = 6/109 (5%)
Query: 233 ISQLNYQVFGDVVAFDATYGKNRY---KCPFVVFYGVNHHNKSTIFSTALVSNEKIETYV 289
+S+ NY V + +NR C + G+ I + S E ET+
Sbjct: 156 LSEKNYPYLMTDVLYIKVREENRVLSKSCHIAI--GITKDGDREIIGFMIQSGESEETWT 213
Query: 290 WLLERFLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDN 338
E +L+ + VI+D K + +AI++ F N + C H L N
Sbjct: 214 TFFE-YLKERGLQGTELVISDAHKGLVSAIRKSFTNVSWQRCQVHFLRN 261
>TRA6_ENTFA (P59787) Transposase for insertion sequence element
IS256 in transposon Tn4001
Length = 390
Score = 31.6 bits (70), Expect = 5.9
Identities = 27/109 (24%), Positives = 44/109 (39%), Gaps = 6/109 (5%)
Query: 233 ISQLNYQVFGDVVAFDATYGKNRY---KCPFVVFYGVNHHNKSTIFSTALVSNEKIETYV 289
+S+ NY V + +NR C + G+ I + S E ET+
Sbjct: 156 LSEKNYPYLMTDVLYIKVREENRVLSKSCHIAI--GITKDGDREIIGFMIQSGESEETWT 213
Query: 290 WLLERFLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDN 338
E +L+ + VI+D K + +AI++ F N + C H L N
Sbjct: 214 TFFE-YLKERGLQGTELVISDAHKGLVSAIRKSFTNVSWQRCQVHFLRN 261
>2ABA_CANTR (P53031) Protein phosphatase PP2A regulatory subunit B
(PR55) (Cell division control protein 55)
Length = 508
Score = 31.2 bits (69), Expect = 7.7
Identities = 20/82 (24%), Positives = 33/82 (39%), Gaps = 10/82 (12%)
Query: 220 DDGNLDRLFWSDGISQLNYQVFGD----------VVAFDATYGKNRYKCPFVVFYGVNHH 269
D G+++ + +D IS + + GD VV F+ K + C + F H
Sbjct: 11 DKGDIENITEADIISTVEFDHTGDFLATGDKGGRVVLFERNQSKKKQSCEYKFFTEFQSH 70
Query: 270 NKSTIFSTALVSNEKIETYVWL 291
+ + +L EKI WL
Sbjct: 71 DAEFDYLKSLEIEEKINKIKWL 92
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.340 0.149 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,426,147
Number of Sequences: 164201
Number of extensions: 2629545
Number of successful extensions: 7384
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 7381
Number of HSP's gapped (non-prelim): 11
length of query: 578
length of database: 59,974,054
effective HSP length: 116
effective length of query: 462
effective length of database: 40,926,738
effective search space: 18908152956
effective search space used: 18908152956
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0302a.8


