

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0301a.4
(147 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
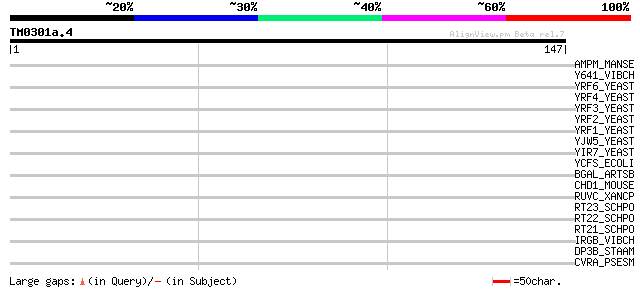
Score E
Sequences producing significant alignments: (bits) Value
AMPM_MANSE (Q11001) Membrane alanyl aminopeptidase precursor (EC... 31 1.1
Y641_VIBCH (Q9KU82) Hypothetical UPF0090 protein VC0641 30 1.9
YRF6_YEAST (P53819) Y'helicase protein 1 copy 6 29 3.2
YRF4_YEAST (O13559) Y'helicase protein 1 copy 4 29 3.2
YRF3_YEAST (P53345) Y'helicase protein 1 copies 3/7 29 3.2
YRF2_YEAST (P40105) Y'helicase protein 1 copy 2 29 3.2
YRF1_YEAST (P24088) Y'helicase protein 1 copies 1/5/8 29 3.2
YJW5_YEAST (P40889) Hypothetical 197.6 kDa protein in FSP2 5'region 29 3.2
YIR7_YEAST (P40434) Hypothetical 197.5 kDa protein in SDL1 5'region 29 3.2
YCFS_ECOLI (P75954) Hypothetical protein ycfS precursor 29 3.2
BGAL_ARTSB (Q59140) Beta-galactosidase (EC 3.2.1.23) (Lactase) 29 3.2
CHD1_MOUSE (P40201) Chromodomain-helicase-DNA-binding protein 1 ... 29 4.2
RUVC_XANCP (Q8P6E4) Crossover junction endodeoxyribonuclease ruv... 28 9.3
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 28 9.3
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 28 9.3
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 28 9.3
IRGB_VIBCH (P25543) Iron-regulated virulence regulatory protein ... 28 9.3
DP3B_STAAM (P50029) DNA polymerase III, beta chain (EC 2.7.7.7) 28 9.3
CVRA_PSESM (Q87VB6) Cell volume regulation protein A homolog 28 9.3
>AMPM_MANSE (Q11001) Membrane alanyl aminopeptidase precursor (EC
3.4.11.-) (Aminopeptidase N-like protein) (CryIA(C)
receptor) (Fragment)
Length = 990
Score = 30.8 bits (68), Expect = 1.1
Identities = 19/66 (28%), Positives = 28/66 (41%), Gaps = 8/66 (12%)
Query: 50 VHPVFHASLLKEAVGNNSVELQL--------LDHLTGEEVASVHPFSVITSRFTTRQEST 101
V P H+S L A+ +N V +D +T + P + ITSR T + T
Sbjct: 823 VRPQDHSSALSSAITSNDVNTMRAFDWLTKNVDQITRTLGSITSPLNTITSRLLTEAQMT 882
Query: 102 LPQVWI 107
Q W+
Sbjct: 883 QVQTWL 888
>Y641_VIBCH (Q9KU82) Hypothetical UPF0090 protein VC0641
Length = 151
Score = 30.0 bits (66), Expect = 1.9
Identities = 22/92 (23%), Positives = 39/92 (41%), Gaps = 13/92 (14%)
Query: 3 IKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHAS----- 57
+++ EN + DC +++ + I +V Y L++ G P+F A+
Sbjct: 37 LRIFIDHENGITVEDCAEVSRQVSAVLDVEDPI-SVVYNLEVSSPGLERPLFKAAHYEQF 95
Query: 58 -------LLKEAVGNNSVELQLLDHLTGEEVA 82
+LK AVGN ++ + GE VA
Sbjct: 96 IGHEVSIVLKMAVGNRRKWKGVIQSIDGETVA 127
>YRF6_YEAST (P53819) Y'helicase protein 1 copy 6
Length = 1859
Score = 29.3 bits (64), Expect = 3.2
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 9/71 (12%)
Query: 47 GGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQ--ESTLPQ 104
G R + S E + N+SV+ TG + V FS+ + TT + +
Sbjct: 728 GSRFETDLYESATSELMANHSVQ-------TGRNIYGVDSFSLTSVSGTTATLLQERASE 780
Query: 105 VWIQWQGKPAD 115
WIQW G +D
Sbjct: 781 RWIQWLGLESD 791
>YRF4_YEAST (O13559) Y'helicase protein 1 copy 4
Length = 1382
Score = 29.3 bits (64), Expect = 3.2
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 9/71 (12%)
Query: 47 GGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQ--ESTLPQ 104
G R + S E + N+SV+ TG + V FS+ + TT + +
Sbjct: 250 GSRFETDLYESATSELMANHSVQ-------TGRNIYGVDSFSLTSVSGTTATLLQERASE 302
Query: 105 VWIQWQGKPAD 115
WIQW G +D
Sbjct: 303 RWIQWLGLESD 313
>YRF3_YEAST (P53345) Y'helicase protein 1 copies 3/7
Length = 1859
Score = 29.3 bits (64), Expect = 3.2
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 9/71 (12%)
Query: 47 GGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQ--ESTLPQ 104
G R + S E + N+SV+ TG + V FS+ + TT + +
Sbjct: 728 GSRFETDLYESATSELMANHSVQ-------TGRNIYGVDSFSLTSVSGTTATLLQERASE 780
Query: 105 VWIQWQGKPAD 115
WIQW G +D
Sbjct: 781 RWIQWLGLESD 791
>YRF2_YEAST (P40105) Y'helicase protein 1 copy 2
Length = 1681
Score = 29.3 bits (64), Expect = 3.2
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 9/71 (12%)
Query: 47 GGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQ--ESTLPQ 104
G R + S E + N+SV+ TG + V FS+ + TT + +
Sbjct: 550 GSRFETDLYESATSELMANHSVQ-------TGRNIYGVDSFSLTSVSGTTATLLQERASE 602
Query: 105 VWIQWQGKPAD 115
WIQW G +D
Sbjct: 603 RWIQWLGLESD 613
>YRF1_YEAST (P24088) Y'helicase protein 1 copies 1/5/8
Length = 1796
Score = 29.3 bits (64), Expect = 3.2
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 9/71 (12%)
Query: 47 GGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQ--ESTLPQ 104
G R + S E + N+SV+ TG + V FS+ + TT + +
Sbjct: 664 GSRFETDLYESATSELMANHSVQ-------TGRNIYGVDSFSLTSVSGTTATLLQERASE 716
Query: 105 VWIQWQGKPAD 115
WIQW G +D
Sbjct: 717 RWIQWLGLESD 727
>YJW5_YEAST (P40889) Hypothetical 197.6 kDa protein in FSP2 5'region
Length = 1758
Score = 29.3 bits (64), Expect = 3.2
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 9/71 (12%)
Query: 47 GGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQ--ESTLPQ 104
G R + S E + N+SV+ TG + V FS+ + TT + +
Sbjct: 535 GSRFETDLYESATSELMANHSVQ-------TGRNIYGVDSFSLTSVSGTTATLLQERASE 587
Query: 105 VWIQWQGKPAD 115
WIQW G +D
Sbjct: 588 RWIQWLGLESD 598
>YIR7_YEAST (P40434) Hypothetical 197.5 kDa protein in SDL1 5'region
Length = 1758
Score = 29.3 bits (64), Expect = 3.2
Identities = 20/71 (28%), Positives = 30/71 (42%), Gaps = 9/71 (12%)
Query: 47 GGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQ--ESTLPQ 104
G R + S E + N+SV+ TG + V FS+ + TT + +
Sbjct: 535 GSRFETDLYESATSELMANHSVQ-------TGRNIYGVDSFSLTSVSGTTATLLQERASE 587
Query: 105 VWIQWQGKPAD 115
WIQW G +D
Sbjct: 588 RWIQWLGLESD 598
>YCFS_ECOLI (P75954) Hypothetical protein ycfS precursor
Length = 320
Score = 29.3 bits (64), Expect = 3.2
Identities = 25/104 (24%), Positives = 42/104 (40%), Gaps = 9/104 (8%)
Query: 44 LPEGGRVHPVFHASLL----KEAVGNNSVELQLLDHLTGEEVASVHPFSVITSRFTTRQE 99
+P G V + +LL +E + N EL+L + G+ +V+P +
Sbjct: 81 VPRAGSVLTIPLQTLLPDAPREGIVINIAELRLYYYPPGKNSVTVYPIGI-----GQLGG 135
Query: 100 STLPQVWIQWQGKPADEPTWKDTLNIRSQFPVFNLEDKVDLSAG 143
TL + PTW T NIR+++ +E + AG
Sbjct: 136 DTLTPTMVTTVSDKRANPTWTPTANIRARYKAQGIELPAVVPAG 179
>BGAL_ARTSB (Q59140) Beta-galactosidase (EC 3.2.1.23) (Lactase)
Length = 1015
Score = 29.3 bits (64), Expect = 3.2
Identities = 8/17 (47%), Positives = 13/17 (76%)
Query: 107 IQWQGKPADEPTWKDTL 123
++W G P+D+P W+D L
Sbjct: 393 LKWVGNPSDDPAWRDAL 409
>CHD1_MOUSE (P40201) Chromodomain-helicase-DNA-binding protein 1
(CHD-1)
Length = 1711
Score = 28.9 bits (63), Expect = 4.2
Identities = 12/40 (30%), Positives = 20/40 (50%)
Query: 90 ITSRFTTRQESTLPQVWIQWQGKPADEPTWKDTLNIRSQF 129
I + + + LP + +WQG P E +W+D I +F
Sbjct: 392 IIAHSNQKSAAGLPDYYCKWQGLPYSECSWEDGALISKKF 431
>RUVC_XANCP (Q8P6E4) Crossover junction endodeoxyribonuclease ruvC
(EC 3.1.22.4) (Holliday junction nuclease ruvC)
(Holliday juction resolvase ruvC)
Length = 174
Score = 27.7 bits (60), Expect = 9.3
Identities = 16/48 (33%), Positives = 24/48 (49%)
Query: 42 LQLPEGGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHPFSV 89
+ + E GR VFHA L+ G+ + L+ L H GE + + P V
Sbjct: 19 IDVDESGRSRHVFHAPLVLLGEGDFAQRLKRLLHGLGELIETYQPQEV 66
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 27.7 bits (60), Expect = 9.3
Identities = 15/57 (26%), Positives = 28/57 (48%), Gaps = 2/57 (3%)
Query: 20 QLTAPYYGPYPIIQRIGAVAYRLQLPEGGR--VHPVFHASLLKEAVGNNSVELQLLD 74
+L + GP+ ++Q+ G Y L LP+ + FH S L++ N+ + +D
Sbjct: 1213 KLAPSFAGPFYVLQKSGPNNYELDLPDSIKHMFSSTFHVSHLEKYRHNSELNYATID 1269
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 27.7 bits (60), Expect = 9.3
Identities = 15/57 (26%), Positives = 28/57 (48%), Gaps = 2/57 (3%)
Query: 20 QLTAPYYGPYPIIQRIGAVAYRLQLPEGGR--VHPVFHASLLKEAVGNNSVELQLLD 74
+L + GP+ ++Q+ G Y L LP+ + FH S L++ N+ + +D
Sbjct: 1213 KLAPSFAGPFYVLQKSGPNNYELDLPDSIKHMFSSTFHVSHLEKYRHNSELNYATID 1269
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 27.7 bits (60), Expect = 9.3
Identities = 15/57 (26%), Positives = 28/57 (48%), Gaps = 2/57 (3%)
Query: 20 QLTAPYYGPYPIIQRIGAVAYRLQLPEGGR--VHPVFHASLLKEAVGNNSVELQLLD 74
+L + GP+ ++Q+ G Y L LP+ + FH S L++ N+ + +D
Sbjct: 1213 KLAPSFAGPFYVLQKSGPNNYELDLPDSIKHMFSSTFHVSHLEKYRHNSELNYATID 1269
>IRGB_VIBCH (P25543) Iron-regulated virulence regulatory protein
irgB
Length = 298
Score = 27.7 bits (60), Expect = 9.3
Identities = 16/46 (34%), Positives = 23/46 (49%)
Query: 41 RLQLPEGGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASVHP 86
RL L + G V V+ LL+ A + +L + +TGE VHP
Sbjct: 55 RLTLTKAGEVFAVYSEQLLELANKSQEALQELNNQVTGELTLVVHP 100
>DP3B_STAAM (P50029) DNA polymerase III, beta chain (EC 2.7.7.7)
Length = 377
Score = 27.7 bits (60), Expect = 9.3
Identities = 20/62 (32%), Positives = 29/62 (46%), Gaps = 4/62 (6%)
Query: 27 GPYPIIQRIGAVAY--RLQLPEGGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVASV 84
G YP R+ Y +L + G H + ASLL GNN ++L D + E++S
Sbjct: 247 GHYPDTTRLFPENYEIKLSIDNGEFYHAIDRASLLAREGGNNVIKLSTGDDVV--ELSST 304
Query: 85 HP 86
P
Sbjct: 305 SP 306
>CVRA_PSESM (Q87VB6) Cell volume regulation protein A homolog
Length = 580
Score = 27.7 bits (60), Expect = 9.3
Identities = 15/46 (32%), Positives = 24/46 (51%)
Query: 35 IGAVAYRLQLPEGGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEE 80
IGA L++PEG R+ +F L G+ ++E+ L + G E
Sbjct: 427 IGAALRELKMPEGTRIAALFRGQQLLHPSGSTTLEVGDLLCVIGHE 472
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.137 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,918,034
Number of Sequences: 164201
Number of extensions: 704033
Number of successful extensions: 1512
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 1507
Number of HSP's gapped (non-prelim): 19
length of query: 147
length of database: 59,974,054
effective HSP length: 100
effective length of query: 47
effective length of database: 43,553,954
effective search space: 2047035838
effective search space used: 2047035838
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0301a.4


