

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0301a.2
(80 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
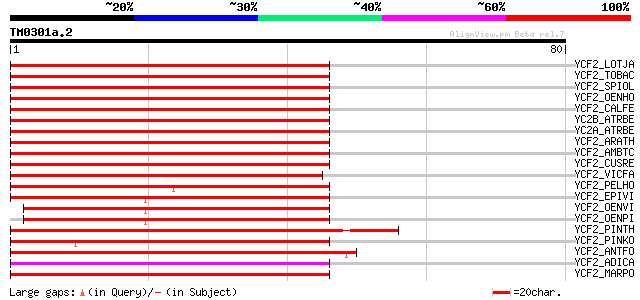
Score E
Sequences producing significant alignments: (bits) Value
YCF2_LOTJA (Q9B1K6) Protein ycf2 91 4e-19
YCF2_TOBAC (P09976) Protein ycf2 82 2e-16
YCF2_SPIOL (P08973) Protein ycf2 82 2e-16
YCF2_OENHO (Q9MEF2) Protein ycf2 82 2e-16
YCF2_CALFE (Q7Y667) Protein ycf2 82 2e-16
YC2B_ATRBE (Q8S8U1) Protein ycf2 82 2e-16
YC2A_ATRBE (Q8S8V2) Protein ycf2 82 2e-16
YCF2_ARATH (P56786) Protein ycf2 82 2e-16
YCF2_AMBTC (P61241) Protein ycf2 80 9e-16
YCF2_CUSRE (P32033) Protein ycf2 75 2e-14
YCF2_VICFA (P15821) Protein ycf2 (Fragment) 75 4e-14
YCF2_PELHO (Q32836) Protein ycf2 70 1e-12
YCF2_EPIVI (P30072) Protein ycf2 69 2e-12
YCF2_OENVI (P31569) Protein ycf2 (Fragment) 64 7e-11
YCF2_OENPI (P31568) Protein ycf2 (Fragment) 64 7e-11
YCF2_PINTH (P41653) Protein ycf2 63 2e-10
YCF2_PINKO (Q85WV5) Protein ycf2 58 4e-09
YCF2_ANTFO (Q859W7) Protein ycf2 49 2e-06
YCF2_ADICA (Q85B60) Protein ycf2 44 7e-05
YCF2_MARPO (P09975) Protein ycf2 44 1e-04
>YCF2_LOTJA (Q9B1K6) Protein ycf2
Length = 2298
Score = 91.3 bits (225), Expect = 4e-19
Identities = 44/46 (95%), Positives = 44/46 (95%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLIHRQRWLR NNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD
Sbjct: 2224 FELLIHRQRWLRTNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 2269
>YCF2_TOBAC (P09976) Protein ycf2
Length = 2280
Score = 82.4 bits (202), Expect = 2e-16
Identities = 40/46 (86%), Positives = 43/46 (92%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLI+RQRWLR N+SLSNG FRSNTLSESYQYLSN+FLSNGTLLD
Sbjct: 2213 FELLINRQRWLRTNSSLSNGSFRSNTLSESYQYLSNLFLSNGTLLD 2258
>YCF2_SPIOL (P08973) Protein ycf2
Length = 2131
Score = 82.4 bits (202), Expect = 2e-16
Identities = 40/46 (86%), Positives = 43/46 (92%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLI+RQRWLR N+SLSNG FRSNTLSESYQYLSN+FLSNGTLLD
Sbjct: 2048 FELLINRQRWLRTNSSLSNGSFRSNTLSESYQYLSNLFLSNGTLLD 2093
>YCF2_OENHO (Q9MEF2) Protein ycf2
Length = 2280
Score = 82.4 bits (202), Expect = 2e-16
Identities = 40/46 (86%), Positives = 43/46 (92%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLI+RQRWLR N+SLSNG FRSNTLSESYQYLSN+FLSNGTLLD
Sbjct: 2213 FELLINRQRWLRTNSSLSNGSFRSNTLSESYQYLSNLFLSNGTLLD 2258
>YCF2_CALFE (Q7Y667) Protein ycf2
Length = 2287
Score = 82.4 bits (202), Expect = 2e-16
Identities = 40/46 (86%), Positives = 42/46 (90%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLIHRQRWLR N+SLSNG FRSNT SESYQYLSN+FLSNGTLLD
Sbjct: 2211 FELLIHRQRWLRTNSSLSNGSFRSNTPSESYQYLSNLFLSNGTLLD 2256
>YC2B_ATRBE (Q8S8U1) Protein ycf2
Length = 2291
Score = 82.4 bits (202), Expect = 2e-16
Identities = 40/46 (86%), Positives = 43/46 (92%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLI+RQRWLR N+SLSNG FRSNTLSESYQYLSN+FLSNGTLLD
Sbjct: 2224 FELLINRQRWLRTNSSLSNGSFRSNTLSESYQYLSNLFLSNGTLLD 2269
>YC2A_ATRBE (Q8S8V2) Protein ycf2
Length = 2291
Score = 82.4 bits (202), Expect = 2e-16
Identities = 40/46 (86%), Positives = 43/46 (92%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLI+RQRWLR N+SLSNG FRSNTLSESYQYLSN+FLSNGTLLD
Sbjct: 2224 FELLINRQRWLRTNSSLSNGSFRSNTLSESYQYLSNLFLSNGTLLD 2269
>YCF2_ARATH (P56786) Protein ycf2
Length = 2294
Score = 82.0 bits (201), Expect = 2e-16
Identities = 39/46 (84%), Positives = 42/46 (90%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLI RQRWLR N+SLSNGFFRSNT SESYQYLSN+F+SNGTLLD
Sbjct: 2227 FELLIQRQRWLRTNSSLSNGFFRSNTRSESYQYLSNLFISNGTLLD 2272
>YCF2_AMBTC (P61241) Protein ycf2
Length = 2304
Score = 80.1 bits (196), Expect = 9e-16
Identities = 39/46 (84%), Positives = 41/46 (88%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLIHRQRWL N+SLSNG FRSNT SESYQYLSN+FLSNGTLLD
Sbjct: 2228 FELLIHRQRWLSTNSSLSNGSFRSNTPSESYQYLSNLFLSNGTLLD 2273
>YCF2_CUSRE (P32033) Protein ycf2
Length = 740
Score = 75.5 bits (184), Expect = 2e-14
Identities = 38/46 (82%), Positives = 40/46 (86%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LIHRQR LR N+SLSN FRSNTLSESYQYLSN+FLSNGTLLD
Sbjct: 673 FEFLIHRQRCLRTNSSLSNRSFRSNTLSESYQYLSNLFLSNGTLLD 718
>YCF2_VICFA (P15821) Protein ycf2 (Fragment)
Length = 121
Score = 74.7 bits (182), Expect = 4e-14
Identities = 36/45 (80%), Positives = 39/45 (86%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLL 45
F LLI RQRW R N+S+SN FFRSNTL ESYQYLSN+FLSNGTLL
Sbjct: 49 FKLLIDRQRWFRTNSSVSNEFFRSNTLYESYQYLSNLFLSNGTLL 93
>YCF2_PELHO (Q32836) Protein ycf2
Length = 2109
Score = 70.1 bits (170), Expect = 1e-12
Identities = 34/48 (70%), Positives = 40/48 (82%), Gaps = 2/48 (4%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFF--RSNTLSESYQYLSNMFLSNGTLLD 46
F LIHR+RWLR N+SLSNGFF ++ TL ESYQYLSN+FLSNG L+D
Sbjct: 2020 FEFLIHRERWLRTNSSLSNGFFCSKTQTLFESYQYLSNLFLSNGRLVD 2067
>YCF2_EPIVI (P30072) Protein ycf2
Length = 2216
Score = 68.9 bits (167), Expect = 2e-12
Identities = 36/48 (75%), Positives = 39/48 (81%), Gaps = 2/48 (4%)
Query: 1 FLLLIHRQRWLRPNNSLS--NGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLI+RQRWLR +SLS NG FRSNTL ESYQYLS +FLSNGTL D
Sbjct: 2150 FELLINRQRWLRTKSSLSKSNGSFRSNTLFESYQYLSTLFLSNGTLFD 2197
>YCF2_OENVI (P31569) Protein ycf2 (Fragment)
Length = 630
Score = 63.9 bits (154), Expect = 7e-11
Identities = 34/46 (73%), Positives = 36/46 (77%), Gaps = 2/46 (4%)
Query: 3 LLIHRQRWLRPNNSLS--NGFFRSNTLSESYQYLSNMFLSNGTLLD 46
LL+ RQRWL SLS NGFFRSNT SESYQYLSN+FLSN LLD
Sbjct: 571 LLLDRQRWLITKRSLSKSNGFFRSNTPSESYQYLSNLFLSNRRLLD 616
>YCF2_OENPI (P31568) Protein ycf2 (Fragment)
Length = 721
Score = 63.9 bits (154), Expect = 7e-11
Identities = 34/46 (73%), Positives = 36/46 (77%), Gaps = 2/46 (4%)
Query: 3 LLIHRQRWLRPNNSLS--NGFFRSNTLSESYQYLSNMFLSNGTLLD 46
LL+ RQRWL SLS NGFFRSNT SESYQYLSN+FLSN LLD
Sbjct: 662 LLLDRQRWLITKRSLSKSNGFFRSNTPSESYQYLSNLFLSNRRLLD 707
>YCF2_PINTH (P41653) Protein ycf2
Length = 2054
Score = 62.8 bits (151), Expect = 2e-10
Identities = 35/56 (62%), Positives = 38/56 (67%), Gaps = 1/56 (1%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLDNTKMK*IIEN 56
F LIHRQRWL N LSN F NTL ESYQYLSN+FLSN LLD K ++EN
Sbjct: 1973 FQSLIHRQRWLGTNRLLSNESFLYNTLFESYQYLSNLFLSNRMLLDQI-TKTLLEN 2027
>YCF2_PINKO (Q85WV5) Protein ycf2
Length = 1320
Score = 58.2 bits (139), Expect = 4e-09
Identities = 32/47 (68%), Positives = 34/47 (72%), Gaps = 1/47 (2%)
Query: 1 FLLLIHRQ-RWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LIHRQ RWL N+ LSN F NTL ESYQYLSN+FLSN LLD
Sbjct: 1238 FQSLIHRQKRWLGTNSLLSNESFLYNTLFESYQYLSNLFLSNRVLLD 1284
>YCF2_ANTFO (Q859W7) Protein ycf2
Length = 2392
Score = 49.3 bits (116), Expect = 2e-06
Identities = 27/51 (52%), Positives = 33/51 (63%), Gaps = 1/51 (1%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLDN-TKM 50
F LL HR+RW++ NN + F +TL ESY YL N F+SN LLD TKM
Sbjct: 2321 FDLLNHRERWIKLNNKSFHYFSIHSTLLESYNYLLNFFISNRILLDEMTKM 2371
>YCF2_ADICA (Q85B60) Protein ycf2
Length = 2104
Score = 43.9 bits (102), Expect = 7e-05
Identities = 22/46 (47%), Positives = 26/46 (55%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LL+HR +WLR N S F N L E+Y+YLSN F LD
Sbjct: 2059 FRLLVHRNKWLRRNRSPFRDFLIYNMLLETYEYLSNPFRFGKASLD 2104
>YCF2_MARPO (P09975) Protein ycf2
Length = 2136
Score = 43.5 bits (101), Expect = 1e-04
Identities = 21/46 (45%), Positives = 30/46 (64%)
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LL HR++WL+ N+ L N F TL E Y+YL + F++N LL+
Sbjct: 2065 FELLTHRKKWLKINSLLLNDSFIYTTLLEIYEYLLHFFIANKKLLN 2110
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.350 0.154 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,240,109
Number of Sequences: 164201
Number of extensions: 206212
Number of successful extensions: 963
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 23
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 932
Number of HSP's gapped (non-prelim): 28
length of query: 80
length of database: 59,974,054
effective HSP length: 56
effective length of query: 24
effective length of database: 50,778,798
effective search space: 1218691152
effective search space used: 1218691152
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0301a.2


