

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0299.14
(88 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
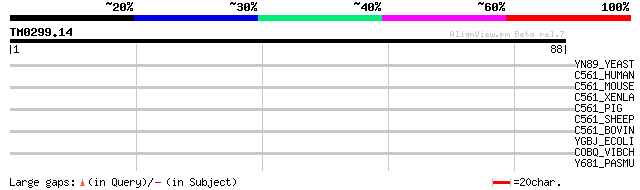
Score E
Sequences producing significant alignments: (bits) Value
YN89_YEAST (P53721) Hypothetical 25.3 kDa protein in TIM23-ARE2 ... 34 0.057
C561_HUMAN (P49447) Cytochrome b561 (Cytochrome b-561) 32 0.28
C561_MOUSE (Q60720) Cytochrome b561 (Cytochrome b-561) 31 0.49
C561_XENLA (Q91577) Cytochrome b561 (Cytochrome b-561) 31 0.63
C561_PIG (Q95245) Cytochrome b561 (Cytochrome b-561) 31 0.63
C561_SHEEP (Q95204) Cytochrome b561 (Cytochrome b-561) 30 1.1
C561_BOVIN (P10897) Cytochrome b561 (Cytochrome b-561) 30 1.4
YGBJ_ECOLI (Q46888) Hypothetical oxidoreductase ygbJ (EC 1.1.-.-) 28 3.1
COBQ_VIBCH (Q9KLL6) Cobyric acid synthase 28 4.1
Y681_PASMU (P57864) Hypothetical protein PM0681 27 7.0
>YN89_YEAST (P53721) Hypothetical 25.3 kDa protein in TIM23-ARE2
intergenic region
Length = 224
Score = 34.3 bits (77), Expect = 0.057
Identities = 17/71 (23%), Positives = 36/71 (49%)
Query: 17 DHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDH 76
++K + + W + + GS P M + KI+ AR++AQ +T+ L + + Y++
Sbjct: 114 NNKYKIITGAWAASLYGSWVIVNKDPIMTKAQKIVQARMYAQFITVGLLLASVGLSMYEN 173
Query: 77 RAEAKRAKNSE 87
+ + K +E
Sbjct: 174 KLHPNKQKVNE 184
>C561_HUMAN (P49447) Cytochrome b561 (Cytochrome b-561)
Length = 251
Score = 32.0 bits (71), Expect = 0.28
Identities = 17/58 (29%), Positives = 29/58 (49%), Gaps = 1/58 (1%)
Query: 23 VGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDHRAEA 80
+G ++L G + Y R K + K++H LH AL +A + AV +Y+ + A
Sbjct: 59 IGLIFLQG-NALLVYRVFRNEAKRTTKVLHGLLHIFALVIALVGLVAVFDYHRKKGYA 115
>C561_MOUSE (Q60720) Cytochrome b561 (Cytochrome b-561)
Length = 250
Score = 31.2 bits (69), Expect = 0.49
Identities = 20/72 (27%), Positives = 33/72 (45%), Gaps = 1/72 (1%)
Query: 9 ESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGA 68
ES ++ V +G ++L G + Y R K + KI+H LH A +A +
Sbjct: 44 ESSLQFNVHPLCMVIGMIFLQG-DALLVYRVFRREAKRTTKILHGLLHVFAFIIALVGLV 102
Query: 69 AVVEYYDHRAEA 80
AV +Y+ + A
Sbjct: 103 AVFDYHKKKGYA 114
>C561_XENLA (Q91577) Cytochrome b561 (Cytochrome b-561)
Length = 247
Score = 30.8 bits (68), Expect = 0.63
Identities = 17/52 (32%), Positives = 27/52 (51%), Gaps = 1/52 (1%)
Query: 23 VGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYY 74
+G ++L G + Y R K S KI+H LH AL ++ + AV +Y+
Sbjct: 59 LGMVFLCG-EALLVYRVFRHETKRSTKILHGVLHIMALVISLVGVIAVFQYH 109
>C561_PIG (Q95245) Cytochrome b561 (Cytochrome b-561)
Length = 252
Score = 30.8 bits (68), Expect = 0.63
Identities = 19/66 (28%), Positives = 31/66 (46%), Gaps = 1/66 (1%)
Query: 9 ESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGA 68
ES ++ V +G ++L G + Y R K + KI+H LH A +A +
Sbjct: 46 ESALQFNVHPLCMIIGLVFLQG-DALLVYRVFRNEAKRTTKILHGLLHVLAFVIALVGLV 104
Query: 69 AVVEYY 74
AV +Y+
Sbjct: 105 AVFDYH 110
>C561_SHEEP (Q95204) Cytochrome b561 (Cytochrome b-561)
Length = 252
Score = 30.0 bits (66), Expect = 1.1
Identities = 19/72 (26%), Positives = 33/72 (45%), Gaps = 1/72 (1%)
Query: 9 ESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGA 68
ES ++ V +G ++L G + Y R K + K++H LH A +A +
Sbjct: 46 ESALQFNVHPLCMVIGLVFLQG-DALLVYRVFRNEAKRTTKVLHGLLHVFAFVIALVGLV 104
Query: 69 AVVEYYDHRAEA 80
AV E++ + A
Sbjct: 105 AVFEHHRKKGYA 116
>C561_BOVIN (P10897) Cytochrome b561 (Cytochrome b-561)
Length = 252
Score = 29.6 bits (65), Expect = 1.4
Identities = 19/72 (26%), Positives = 33/72 (45%), Gaps = 1/72 (1%)
Query: 9 ESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGA 68
ES ++ V +G ++L G + Y R K + K++H LH A +A +
Sbjct: 46 ESALQFNVHPLCMIIGLVFLQG-DALLVYRVFRNEAKRTTKVLHGLLHVFAFVIALVGLV 104
Query: 69 AVVEYYDHRAEA 80
AV E++ + A
Sbjct: 105 AVFEHHRKKGYA 116
>YGBJ_ECOLI (Q46888) Hypothetical oxidoreductase ygbJ (EC 1.1.-.-)
Length = 302
Score = 28.5 bits (62), Expect = 3.1
Identities = 19/55 (34%), Positives = 29/55 (52%), Gaps = 7/55 (12%)
Query: 28 LSGITGSIAYNWSRPNMKTSVKIIH---ARLH----AQALTLAALAGAAVVEYYD 75
L + G + + P + ++VKIIH A +H A+A+ LAA AG + YD
Sbjct: 157 LEAVAGKVYRIGAEPGLGSTVKIIHQLLAGVHIAAGAEAMALAARAGIPLDVMYD 211
>COBQ_VIBCH (Q9KLL6) Cobyric acid synthase
Length = 484
Score = 28.1 bits (61), Expect = 4.1
Identities = 13/25 (52%), Positives = 16/25 (64%)
Query: 59 ALTLAALAGAAVVEYYDHRAEAKRA 83
AL + AGA +E YDHRA +RA
Sbjct: 439 ALRICQWAGAKEIEAYDHRAAQERA 463
>Y681_PASMU (P57864) Hypothetical protein PM0681
Length = 433
Score = 27.3 bits (59), Expect = 7.0
Identities = 16/60 (26%), Positives = 36/60 (59%), Gaps = 7/60 (11%)
Query: 23 VGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQ------ALTLAALAGAAVVEYYDH 76
+ + + G T ++ +SR N++ S+++I + ++ A+T AALAG A++ Y+++
Sbjct: 309 IAFMCMFGTTITVIDGYSRTNVE-SLRLILGKKESRPSYLNVAITFAALAGLAIIFYFNN 367
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.127 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,918,892
Number of Sequences: 164201
Number of extensions: 253678
Number of successful extensions: 664
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 654
Number of HSP's gapped (non-prelim): 11
length of query: 88
length of database: 59,974,054
effective HSP length: 64
effective length of query: 24
effective length of database: 49,465,190
effective search space: 1187164560
effective search space used: 1187164560
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0299.14


