

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0296b.2
(1291 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
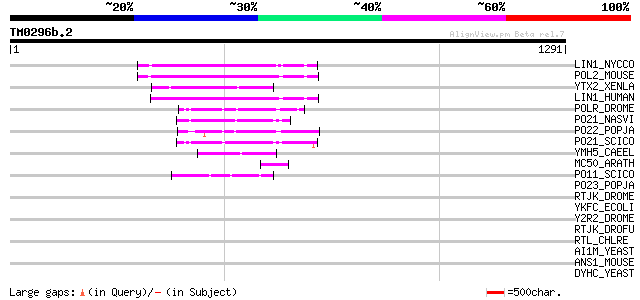
Sequences producing significant alignments: (bits) Value
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 136 4e-31
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 128 9e-29
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 128 1e-28
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 122 6e-27
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 101 2e-20
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 82 9e-15
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 66 5e-10
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 60 5e-08
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 59 1e-07
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 59 1e-07
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 50 4e-05
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 42 0.014
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 39 0.092
YKFC_ECOLI (Q47688) Hypothetical protein ykfC 36 0.59
Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I retro... 35 1.0
RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile elem... 35 1.0
RTL_CHLRE (P11660) Reverse transcriptase-like protein 34 2.3
AI1M_YEAST (P03875) COX1/OXI3 intron 1 protein 34 2.9
ANS1_MOUSE (P59672) Ankyrin repeat and SAM domain protein 1 33 3.8
DYHC_YEAST (P36022) Dynein heavy chain, cytosolic (DYHC) 33 5.0
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 136 bits (342), Expect = 4e-31
Identities = 110/424 (25%), Positives = 196/424 (45%), Gaps = 19/424 (4%)
Query: 298 NRRNSYGTRKLERTGSDWGTLTNTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYF 357
N+ S+ K+ + LT + V ++ + ++ + +D S++Q+ Y+
Sbjct: 359 NKSKSWFFEKINKIDKPLANLTRKKRV----KSLISSIRNGNDEITTDPSEIQKILNEYY 414
Query: 358 QALFAG---NQEASQTPFPGDNVPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDG 414
+ L++ N + ++P+L + +L +P+S E+ + ++ +PGPDG
Sbjct: 415 KKLYSHKYENLKEIDQYLEACHLPRLSQKEVEMLNRPISSSEIASTIQNLPKKKSPGPDG 474
Query: 415 FQPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLLEILVVLIPKVNH-PTSIKEFRPISL 473
F F++ + E++ + + + G + E + LIPK PT + +RPISL
Sbjct: 475 FTSEFYQTFKEELVPILLNLFQNIEKEGILPNTFYEANITLIPKPGKDPTRKENYRPISL 534
Query: 474 CNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGN 533
N+ K++ K+L NRI+ + I+ Q F+PG N + +I +++ + K +
Sbjct: 535 MNIDAKILNKILTNRIQQHIKKIIHHDQVGFIPGSQGWFNIRKSINVIQHINK-LKNKDH 593
Query: 534 LAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPG 593
+ ID EKA+D+I F+ TL+ + LI S +++ NG L SF
Sbjct: 594 MILSIDAEKAFDNIQHPFMIRTLKKIGIEGTFLKLIEAIYSKPTANIILNGVKLKSFPLR 653
Query: 594 RGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCK 653
G RQG P SP F + ME L+I I+ E K +H G I FA D++++ +
Sbjct: 654 SGTRQGCPLSPLLFNIVMEVLAIAIR---EEKAIKGIH--IGSEEIKLSLFADDMIVYLE 708
Query: 654 ASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLRALIPF--VNDLG 711
+ T L++ ++ S +IN KS A + E ++ IPF V
Sbjct: 709 NTRDSTTKLLEVIKEYSNVSGYKINTHKSVAFI---YTNNNQAEKTVKDSIPFTVVPKKM 765
Query: 712 KYLG 715
KYLG
Sbjct: 766 KYLG 769
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 128 bits (322), Expect = 9e-29
Identities = 108/427 (25%), Positives = 188/427 (43%), Gaps = 19/427 (4%)
Query: 298 NRRNSYGTRKLERTGSDWGTLTNTQAVIIRKRNSVHRLKLSDGSWCSDTSKLQEEALGYF 357
N+ S+ K+ + LT R + +++++ G +D ++Q ++
Sbjct: 386 NQTRSWFFEKINKIDKPLARLTKGH----RDKILINKIRNEKGDITTDPEEIQNTIRSFY 441
Query: 358 QALFAG---NQEASQTPFPGDNVPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDG 414
+ L++ N + VPKL + L P+S +E+ + S+ + +PGPDG
Sbjct: 442 KRLYSTKLENLDEMDKFLDRYQVPKLNQDQVDHLNSPISPKEIEAVINSLPTKKSPGPDG 501
Query: 415 FQPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLLEILVVLIPKVNH-PTSIKEFRPISL 473
F F++ + E + + K+ + G + E + LIPK PT I+ FRPISL
Sbjct: 502 FSAEFYQTFKEDLIPILHKLFHKIEVEGTLPNSFYEATITLIPKPQKDPTKIENFRPISL 561
Query: 474 CNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGN 533
N+ K++ K+L NRI+ + I+ P Q F+PG N + +IH +++ + K +
Sbjct: 562 MNIDAKILNKILANRIQEHIKAIIHPDQVGFIPGMQGWFNIRKSINVIHYINK-LKDKNH 620
Query: 534 LAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPG 593
+ +D EKA+D I F+ + LE +++I S ++ NG L +
Sbjct: 621 MIISLDAEKAFDKIQHPFMIKVLERSGIQGPYLNMIKAIYSKPVANIKVNGEKLEAIPLK 680
Query: 594 RGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCK 653
G RQG P SPY F + +E L+ I+ E + G + A D++++
Sbjct: 681 SGTRQGCPLSPYLFNIVLEVLARAIRQQKEIKGIQ-----IGKEEVKISLLADDMIVYIS 735
Query: 654 ASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLRALIPF--VNDLG 711
L++ + E +IN KS A + E +R PF V +
Sbjct: 736 DPKNSTRELLNLINSFGEVVGYKINSNKSMAFL---YTKNKQAEKEIRETTPFSIVTNNI 792
Query: 712 KYLGFPL 718
KYLG L
Sbjct: 793 KYLGVTL 799
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 128 bits (321), Expect = 1e-28
Identities = 83/288 (28%), Positives = 149/288 (50%), Gaps = 9/288 (3%)
Query: 329 RNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGNQ---EASQTPFPGDNVPKLKEVVR 385
R + L DG+ D +++ A ++Q LF+ + +A + + D +P + E +
Sbjct: 385 RKQITCLFAEDGTPLEDPEAIRDRARSFYQNLFSPDPISPDACEELW--DGLPVVSERRK 442
Query: 386 FLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVD 445
L P++ +E+ QAL M +PG DG FF+ +W+ +G +V++EAF+ G++
Sbjct: 443 ERLETPITLDELSQALRLMPHNKSPGLDGLTIEFFQFFWDTLGPDFHRVLTEAFKKGELP 502
Query: 446 ERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFL 505
++ L+PK IK +RP+SL + YK++ K + R++ L +++ P Q+ +
Sbjct: 503 LSCRRAVLSLLPKKGDLRLIKNWRPVSLLSTDYKIVAKAISLRLKSVLAEVIHPDQSYTV 562
Query: 506 PGRGTMDNAFLA*EIIHQMHRSMARKGNLAF-KIDLEKAYDSISWSFLRETLELYEFPLV 564
PGR DN FL +++H R+ +LAF +D EKA+D + +L TL+ Y F
Sbjct: 563 PGRTIFDNVFLIRDLLHFARRTGL---SLAFLSLDQEKAFDRVDHQYLIGTLQAYSFGPQ 619
Query: 565 IIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCME 612
+ + +S++ + N + GRG+RQG P S + L +E
Sbjct: 620 FVGYLKTMYASAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSLAIE 667
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 122 bits (306), Expect = 6e-27
Identities = 100/398 (25%), Positives = 182/398 (45%), Gaps = 15/398 (3%)
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAG---NQEASQTPFPGDNVPKLKEV 383
R++N + +K G +D +++Q Y++ L+A N E +P+L +
Sbjct: 384 REKNQIDTIKNDRGDITTDPTEIQTTIREYYKHLYANKLENLEEMDKFLDTYTLPRLNQE 443
Query: 384 VRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGK 443
L +P++ E+ + S+ + +PGP+GF F+++Y E++ + K+ + G
Sbjct: 444 EVESLNRPITSSEIEAIINSLPNKKSPGPEGFTAEFYQRYKEELVPFLLKLFQSIEKEGI 503
Query: 444 VDERLLEILVVLIPKVNHPTSIKE-FRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQN 502
+ E ++LIPK T+ KE FRPISL N+ K++ K+L N+I+ + ++ Q
Sbjct: 504 LPNSFYEASIILIPKPGRDTTKKENFRPISLMNIDAKILNKILANQIQQHIKKLIHHDQV 563
Query: 503 SFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFP 562
F+P N + II ++R+ ++ ID EKA+D I F+ + L
Sbjct: 564 GFIPAMQGWFNIRKSINIIQHINRT-KDTNHMIISIDAEKAFDKIQQPFMLKPLNKLGID 622
Query: 563 LVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLV 622
+ +I +++ NG L + G RQG P SP + +E L+ I+
Sbjct: 623 GTYLKIIRAIYDKPTANIILNGQKLEAPPLKTGTRQGCPLSPLLPNIVLEVLARAIRQEK 682
Query: 623 EPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKS 682
E + G + FA D++++ + + L+ + + S +INVQKS
Sbjct: 683 EIKGIQ-----LGKEEVKLSLFADDMIVYLENPIVSAQNLLKLISNFSKVSGYKINVQKS 737
Query: 683 KAITSIGVAPEVRTEIGLRALIPF--VNDLGKYLGFPL 718
+A +TE + + +PF + KYLG L
Sbjct: 738 QAFL---YTNNRQTESQIMSELPFTIASKRIKYLGIQL 772
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 101 bits (251), Expect = 2e-20
Identities = 77/295 (26%), Positives = 134/295 (45%), Gaps = 20/295 (6%)
Query: 392 VSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLLEI 451
++++++R + +S+ S +PGPDG P K E + ++++ G + +
Sbjct: 344 ITEQDLRASRVSLSS--SPGPDGITP---KSAREVPSGIMLRIMNLILWCGNLPHSIRLA 398
Query: 452 LVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPGRGTM 511
V IPK ++FRPIS+ +V + + +L R+ +N P Q FLP G
Sbjct: 399 RTVFIPKTVTAKRPQDFRPISVPSVLVRQLNAILATRLNSSIN--WDPRQRGFLPTDGCA 456
Query: 512 DNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMC 571
DNA + ++ H+ + +D+ KA+DS+S + + +TL Y P +D +
Sbjct: 457 DNATIVDLVLRHSHKHF--RSCYIANLDVSKAFDSLSHASIYDTLRAYGAPKGFVDYVQN 514
Query: 572 YVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLH 631
SL +G F P RG++QGDP SP F L M+RL + + +
Sbjct: 515 TYEGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILFNLVMDRLLRTLPS--------EIG 566
Query: 632 RTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAIT 686
G + FA D++LF + + +L L FL S++ + + K T
Sbjct: 567 AKVGNAITNAAAFADDLVLFAETRMGLQVLLDKTLDFL---SIVGLKLNADKCFT 618
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 82.0 bits (201), Expect = 9e-15
Identities = 70/266 (26%), Positives = 120/266 (44%), Gaps = 18/266 (6%)
Query: 388 LVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDER 447
L +P+S +E+++ + TA GPDG W + + + + + G+ R
Sbjct: 315 LWRPISNDEIKEVEACKR--TAAGPDGMTT----TAWNSIDECIKSLFNMIMYHGQCPRR 368
Query: 448 LLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPG 507
L+ VLIPK FRP+S+ +V + ++L NRI + ++ Q +F+
Sbjct: 369 YLDSRTVLIPKEPGTMDPACFRPLSIASVALRHFHRILANRIGE--HGLLDTRQRAFIVA 426
Query: 508 RGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVIID 567
G +N L +I + M KG +D++KA+DS+ + + L + PL + +
Sbjct: 427 DGVAENTSLLSAMIKEAR--MKIKGLYIAILDVKKAFDSVEHRSILDALRRKKLPLEMRN 484
Query: 568 LIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRW 627
IM +S+ L +P RG+RQGDP SP F M+ + ++ L E +
Sbjct: 485 YIMWVYRNSKTRLEVVKTKGRWIRPARGVRQGDPLSPLLFNCVMDAV---LRRLPENTGF 541
Query: 628 KPLHRTAGGTGISHLFFAGDVLLFCK 653
G I L FA D++L +
Sbjct: 542 -----LMGAEKIGALVFADDLVLLAE 562
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 66.2 bits (160), Expect = 5e-10
Identities = 88/349 (25%), Positives = 143/349 (40%), Gaps = 43/349 (12%)
Query: 390 QPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLL 449
+P+++EE++ A+ K +APG DG TV + V LL
Sbjct: 4 RPIAREEIQCAIKGWKP-SAPGSDGL--------------TVQAITRTRLPRNFVQLHLL 48
Query: 450 E---------ILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPM 500
+ LIPK + +RPI++ + +L+ ++L R+ + + P
Sbjct: 49 RGHVPTPWTAMRTTLIPKDGDLENPSNWRPITIASALQRLLHRILAKRLEAAVE--LHPA 106
Query: 501 QNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYE 560
Q + GT+ N+ L I R RK +D+ KA+D++S S + L+
Sbjct: 107 QKGYARIDGTLVNSLLLDTYISS--RREQRKTYNVVSLDVRKAFDTVSHSSICRALQRLG 164
Query: 561 FPLVIIDLIMCYVSSSQISL-LWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQ 619
+ I +S S ++ + G+ RG++QGDP SP+ F ++ L +Q
Sbjct: 165 IDEGTSNYITGSLSDSTTTIRVGPGSQTRKICIRRGVKQGDPLSPFLFNAVLDELLCSLQ 224
Query: 620 NLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFL-CEGSVLRIN 678
+ PG + T G I L FA D+LL V T L F G L
Sbjct: 225 S--TPG----IGGTIGEEKIPVLAFADDLLLLEDNDVLLPTTLATVANFFRLRGMSLNAK 278
Query: 679 VQKSKAITSIGVAPEVRTEIGLR---ALIPFVNDLG--KYLG--FPLQG 720
S ++ + G RT+ LR L+P + + +YLG F L G
Sbjct: 279 KSVSISVAASGGVCIPRTKPFLRVDNVLLPVTDRMQTFRYLGHFFGLSG 327
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 59.7 bits (143), Expect = 5e-08
Identities = 86/340 (25%), Positives = 146/340 (42%), Gaps = 34/340 (10%)
Query: 388 LVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFES-GKVDE 446
L PVS E++ A S + GPDG P + W + D +++ F G+V
Sbjct: 155 LWDPVSLIEIKSARASNEK--GAGPDGVTP----RSWNALDDRYKRLLYNIFVFYGRVPS 208
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNR-IRPFLNDIVGPMQNSFL 505
+ V PK+ FRP+S+C+V + K+L R + + D Q ++L
Sbjct: 209 PIKGSRTVFTPKIEGGPDPGVFRPLSICSVILREFNKILARRFVSCYTYD---ERQTAYL 265
Query: 506 PGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVI 565
P G N + II + R + ++ ++A +DL KA++S+ S L + + P +
Sbjct: 266 PIDGVCINVSMLTAIIAEAKR-LRKELHIAI-LDLVKAFNSVYHSALIDAITEAGCPPGV 323
Query: 566 IDLIMCYVSSSQISLLWNG-APLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEP 624
+D I ++ + + G L S G+ QGDP S F L E+ ++ L
Sbjct: 324 VDYIADMYNNVITEMQFEGKCELASIL--AGVYQGDPLSGPLFTLAYEKA---LRALNNE 378
Query: 625 GRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKA 684
GR+ ++ ++ D LL +V + +D LRIN +KSK
Sbjct: 379 GRF-----DIADVRVNASAYSDDGLLLA-MTVIGLQHNLDKFGETLAKIGLRINSRKSKT 432
Query: 685 ITSIGVAPEVRTEI--GLRALIP-------FVNDLGKYLG 715
++ + E + +I R L+ ++DL KYLG
Sbjct: 433 VSLVPSGREKKMKIVSNRRLLLKASELKPLTISDLWKYLG 472
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 58.5 bits (140), Expect = 1e-07
Identities = 42/184 (22%), Positives = 83/184 (44%), Gaps = 3/184 (1%)
Query: 437 EAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDI 496
++F + R +++ IPK +P+S +RPISL + +++ +++ +RIR + +
Sbjct: 654 QSFSDSAIPNRWKHAVIIPIPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYSHL 713
Query: 497 VGPMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETL 556
+ P Q+ FL R + + + H + ++ L F D KA+D +S L + L
Sbjct: 714 LSPHQHGFLNFRSCPSSLVRSISLYHSILKNEKSLDILFF--DFAKAFDKVSHPILLKKL 771
Query: 557 ELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKP-GRGLRQGDPKSPYFFVLCMERLS 615
L+ + ++ S+ N + P G+ QG P F+L + L
Sbjct: 772 ALFGLDKLTCSWFKEFLHLRTFSVKINKFVSSNAYPISSGVPQGSVSGPLLFILFINDLL 831
Query: 616 IHIQ 619
I ++
Sbjct: 832 IDLE 835
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 58.5 bits (140), Expect = 1e-07
Identities = 31/65 (47%), Positives = 36/65 (54%)
Query: 583 NGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHL 642
NGAP P RGLRQGDP SPY F+LC E LS + E GR + + I+HL
Sbjct: 15 NGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSPRINHL 74
Query: 643 FFAGD 647
FA D
Sbjct: 75 LFADD 79
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 50.1 bits (118), Expect = 4e-05
Identities = 57/243 (23%), Positives = 101/243 (41%), Gaps = 15/243 (6%)
Query: 377 VPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVW---- 432
+ +LK + R L + +EV ++ K +PGPDG + W + + ++
Sbjct: 391 IDRLKAIAR-PLPPDLEMDEVSDSVRRCKVRKSPGPDGIVGEMVRAVWGAIPEYMFCLYK 449
Query: 433 KVVSEAFESGKVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPF 492
+ + E++ K L IL+ L+ ++ +RPI L + K++ ++V R+
Sbjct: 450 QCLLESYFPQKWKIASLVILLKLLDRIRSDPG--SYRPICLLDNLGKVLEGIMVKRLDQK 507
Query: 493 LNDI-VGPMQNSFLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSF 551
L D+ V P Q +F G+ T D A + + K + ID + A+D++ W
Sbjct: 508 LMDVEVSPYQFAFTYGKSTED----AWRCVQRHVECSEMKYVIGLNIDFQGAFDNLGW-- 561
Query: 552 LRETLELYEFPLVIIDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCM 611
L L+L E L M Y ++ + + RG QG P + L M
Sbjct: 562 LSMLLKLDEAQSNEFGLWMSYFGGRKVYYV-GKTGIVRKDVTRGCPQGSKSGPAMWKLVM 620
Query: 612 ERL 614
L
Sbjct: 621 NEL 623
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 41.6 bits (96), Expect = 0.014
Identities = 34/142 (23%), Positives = 64/142 (44%), Gaps = 8/142 (5%)
Query: 509 GTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDL 568
GT+ N + I ++ R + N+ +D+ KA+D++S + + + + D
Sbjct: 1 GTLANIIMLEHYI-KLRRLKGKTYNVV-SLDIRKAFDTVSHPAILRAMRAFGIDDGMQDF 58
Query: 569 IMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWK 628
IM ++ + +++ G G++QGDP SP F + ++ L + N +PG
Sbjct: 59 IMSTITDAYTNIVVGGRTTNKIYIRNGVKQGDPLSPVLFNIVLDELVTRL-NDEQPGA-- 115
Query: 629 PLHRTAGGTGISHLFFAGDVLL 650
I+ L FA D+LL
Sbjct: 116 ---SMTPACKIASLAFADDLLL 134
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 38.9 bits (89), Expect = 0.092
Identities = 35/164 (21%), Positives = 66/164 (39%), Gaps = 17/164 (10%)
Query: 335 LKLSDGSWCSDTSKLQEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRFLL------ 388
+K G WC + + E +FA N E TP+ ++V +L
Sbjct: 381 IKNPSGGWCRTSLEKTE--------VFANNLEQRFTPYNYAPESLCRQVEEYLESPFQMS 432
Query: 389 --VQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
+ V+ EEV+ + + APG D + +Q + + + + G +
Sbjct: 433 LPLSAVTLEEVKNLIAKLPLKKAPGEDLLDNRTIRLLPDQALQFLALIFNSVLDVGYFPK 492
Query: 447 RLLEILVVLIPKVNH-PTSIKEFRPISLCNVTYKLITKLLVNRI 489
+++I K PT + +RP SL K++ +L++NR+
Sbjct: 493 AWKSASIIMIHKTGKTPTDVDSYRPTSLLPSLGKIMERLILNRL 536
>YKFC_ECOLI (Q47688) Hypothetical protein ykfC
Length = 376
Score = 36.2 bits (82), Expect = 0.59
Identities = 74/371 (19%), Positives = 135/371 (35%), Gaps = 81/371 (21%)
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
L+ QP E + +S K PG DG K + +++ + SG
Sbjct: 23 LITQPEWLAEAARITLSSKGAHTPGVDGVN----KTMLQARLAVELQILRDELLSGHYQP 78
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
L V IPK N + RP+ + + +++ + ++ + P + F P
Sbjct: 79 --LPARRVYIPKSNG-----KLRPLGIPALRDRIVQRAMLMAMEPIWESDFHTLSYGFRP 131
Query: 507 GRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVII 566
R ++ +A ++ Q+ +G + DL +D++ L + + +
Sbjct: 132 ER-SVHHAIRTVKL--QLTDCGETRGRWVIEGDLSSYFDTVHHRLLMKAVRRRISDARFM 188
Query: 567 DLIMCYVSSSQISL-LWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQ------ 619
L+ + + I + L+ A G+ QG SP + + ++
Sbjct: 189 TLLWKTIKAGHIDVGLFRAA-------SEGVPQGGVISPLLSNIMLNEFDQYLHERYLSG 241
Query: 620 ----------NLVEPGR---------WKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVT 660
N ++ GR WKP +++ +A D +L K + AQV
Sbjct: 242 KARKDRWYWNNSIQRGRSTAVRENWQWKP--------AVAYCRYADDFVLIVKGTKAQVE 293
Query: 661 VLVDALQFLCEGSV-LRINVQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYLGFPLQ 719
+ + + + EGS+ LR+N+ K+K IP VND GF
Sbjct: 294 AIREECRGVLEGSLKLRLNMDKTK--------------------IPHVND-----GFIFL 328
Query: 720 GRRLNSERCEY 730
G RL +R Y
Sbjct: 329 GHRLIRKRSRY 339
>Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I
retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 35.4 bits (80), Expect = 1.0
Identities = 36/157 (22%), Positives = 63/157 (39%), Gaps = 7/157 (4%)
Query: 396 EVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESG--KVDERLLEILV 453
EV + +KS +PG DG K W + + + + S G + + ++
Sbjct: 438 EVDTCVARLKSRRSPGLDGINGTICKAVWRAIPEHLASLFSRCIRLGYFPAEWKCPRVVS 497
Query: 454 VLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPGRGTMDN 513
+L +R I L V K++ ++VNR+R L + Q F GR D
Sbjct: 498 LLKGPDKDKCEPSSYRGICLLPVFGKVLEAIMVNRVREVLPEGC-RWQFGFRQGRCVED- 555
Query: 514 AFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWS 550
A + + A + L +D + A+D++ WS
Sbjct: 556 ---AWRHVKSSVGASAAQYVLGTFVDFKGAFDNVEWS 589
>RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 35.4 bits (80), Expect = 1.0
Identities = 38/162 (23%), Positives = 66/162 (40%), Gaps = 23/162 (14%)
Query: 340 GSWCSDTSKLQEEALGY----FQAL-FAGNQ------EASQTPFPGDNVPKLKEVVRFLL 388
G WC + + E + F+AL FA E+ QTPF L
Sbjct: 386 GGWCRTSREKTEVFANHLEQRFKALAFAPESHSLMVAESLQTPFQ-----------MALP 434
Query: 389 VQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE-R 447
PV+ EEV++ + +K APG D + +Q + + + G + R
Sbjct: 435 ADPVTLEEVKELVSKLKPKKAPGEDLLDNRTIRLLPDQALLYLVLIFNSILRVGYFPKAR 494
Query: 448 LLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRI 489
+++++ P + +RP SL K++ +L++NRI
Sbjct: 495 PTASIIMILKPGKQPLDVDSYRPTSLLPSLGKMLERLILNRI 536
>RTL_CHLRE (P11660) Reverse transcriptase-like protein
Length = 368
Score = 34.3 bits (77), Expect = 2.3
Identities = 32/112 (28%), Positives = 45/112 (39%), Gaps = 13/112 (11%)
Query: 504 FLPGRGTMDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPL 563
F PG GT + QM + + F DL+ AY+S+ L +TL+L P
Sbjct: 152 FRPGFGTHHAVLHLAQKAQQMLNKQQQAVLVTF--DLQAAYNSVDIKHLMQTLQLQYLPN 209
Query: 564 VIIDLIMCYVSSSQISLLWNGAPLPSFKPG-RGLRQGDPKSPYFFVLCMERL 614
+ LI +W PL G GL QG SP F +++L
Sbjct: 210 DMKKLIW----------MWQHLPLAQLNAGINGLAQGYAYSPTLFAWYVDQL 251
>AI1M_YEAST (P03875) COX1/OXI3 intron 1 protein
Length = 834
Score = 33.9 bits (76), Expect = 2.9
Identities = 45/179 (25%), Positives = 73/179 (40%), Gaps = 18/179 (10%)
Query: 459 VNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPG-RGTMDNAFLA 517
VN P RP+S+ N K++ +++ R L+ I ++ G R M
Sbjct: 313 VNIPKPKGGMRPLSVGNPRDKIVQEVM----RMILDTIFDKKMSTHSHGFRKNMSCQTAI 368
Query: 518 *EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMCYVSSSQ 577
E+ R+M N ++DL+K +D+IS + + L+ Y IDL+ + +
Sbjct: 369 WEV-----RNMFGGSNWFIEVDLKKCFDTISHDLIIKELKRYISDKGFIDLVYKLLRAGY 423
Query: 578 ISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQ---NLVEPGRWKPLHRT 633
I KP GL QG SP + M + ++ NL G+ K H T
Sbjct: 424 ID-----EKGTYHKPMLGLPQGSLISPILCNIVMTLVDNWLEDYINLYNKGKVKKQHPT 477
>ANS1_MOUSE (P59672) Ankyrin repeat and SAM domain protein 1
Length = 1150
Score = 33.5 bits (75), Expect = 3.8
Identities = 39/173 (22%), Positives = 71/173 (40%), Gaps = 21/173 (12%)
Query: 302 SYGTRKLERT---GSDWGTLTNTQAVI---------IRKRNSVHRLKLSDGSWCSDTSKL 349
S G ++LE++ S+W + + I +K + L+ S G W
Sbjct: 668 SIGKKRLEKSPSFASEWDEIEKIMSSIGEGIDFSQEQQKISGSRTLEQSVGEWLESIGLQ 727
Query: 350 QEEALGYFQALFAGNQEASQTPFPGDNVPKLKEVVRFLLVQPVSKEEVRQALMSMKSFTA 409
Q E+ + L G + F G NV + +++ + P + ++ QA S+ A
Sbjct: 728 QYES----KLLLNGFDDVR---FLGSNVMEEQDLREIGISDPQHRRKLLQAARSLPKVKA 780
Query: 410 PGPDGFQPFFFKQYWEQVG--DTVWKVVSEAFESGKVDERLLEILVVLIPKVN 460
G DG P + + +G D V +S + S + L E+ +V + KV+
Sbjct: 781 LGYDGVSPTSVPSWLDSLGLQDYVHSFLSSGYSSIDTVKNLWELELVNVLKVH 833
>DYHC_YEAST (P36022) Dynein heavy chain, cytosolic (DYHC)
Length = 4092
Score = 33.1 bits (74), Expect = 5.0
Identities = 21/66 (31%), Positives = 33/66 (49%), Gaps = 1/66 (1%)
Query: 443 KVDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQN 502
K D R LE + L+PK+ H ++ PIS ++Y L + ++ I +L + G N
Sbjct: 555 KKDIRQLETGLELLPKILHVEALNNIPPIS-ARISYFLNVQSRIDNIVQYLEALFGSNWN 613
Query: 503 SFLPGR 508
L GR
Sbjct: 614 DTLEGR 619
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.340 0.149 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 141,851,767
Number of Sequences: 164201
Number of extensions: 5794982
Number of successful extensions: 14485
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 14446
Number of HSP's gapped (non-prelim): 26
length of query: 1291
length of database: 59,974,054
effective HSP length: 122
effective length of query: 1169
effective length of database: 39,941,532
effective search space: 46691650908
effective search space used: 46691650908
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0296b.2


