

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0296a.5
(754 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
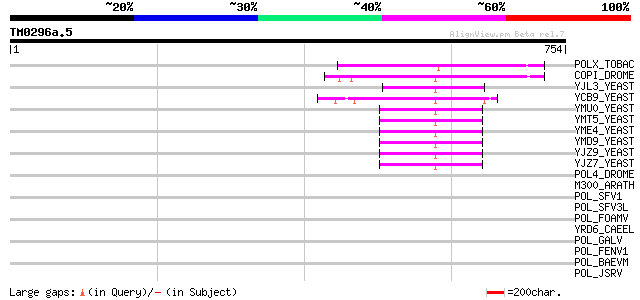
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 154 8e-37
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 123 2e-27
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 64 1e-09
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 62 6e-09
YMU0_YEAST (Q04670) Transposon Ty1 protein B 49 5e-05
YMT5_YEAST (Q04214) Transposon Ty1 protein B 49 5e-05
YME4_YEAST (Q04711) Transposon Ty1 protein B 49 5e-05
YMD9_YEAST (Q03434) Transposon Ty1 protein B 49 5e-05
YJZ9_YEAST (P47100) Transposon Ty1 protein B 49 5e-05
YJZ7_YEAST (P47098) Transposon Ty1 protein B 49 5e-05
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 40 0.017
M300_ARATH (P93293) Hypothetical mitochondrial protein AtMg00300... 39 0.051
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 39 0.066
POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.2... 37 0.15
POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse transcript... 37 0.15
YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II 37 0.25
POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC 3.4.23... 35 0.73
POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse transcript... 35 0.73
POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.2... 35 0.73
POL_JSRV (P31623) Pol polyprotein [Contains: Reverse transcripta... 33 2.1
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 154 bits (389), Expect = 8e-37
Identities = 91/286 (31%), Positives = 149/286 (51%), Gaps = 6/286 (2%)
Query: 446 NIWHHRLGHPSQPCY--ISLQQMYPPITTSPCKNCDTCHFAKQKRLSFSLRNTIYGTVLE 503
++WH R+GH S+ ++ + + + K CD C F KQ R+SF + +L+
Sbjct: 423 DLWHKRMGHMSEKGLQILAKKSLISYAKGTTVKPCDYCLFGKQHRVSFQTSSERKLNILD 482
Query: 504 LLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARAALQTFILYSQRQYGS 563
L++ D+ GP + S+ G KYF + D SR W+ ++K K + Q F +R+ G
Sbjct: 483 LVYSDVCGPMEIESMGGNKYFVTFIDDASRKLWVYILKTKDQVFQVFQKFHALVERETGR 542
Query: 564 LVKIVRLDNGVEFAMTDF---YSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARALMLQ 620
+K +R DNG E+ +F S HG++H+ + TPQ NGV ER ++ I+ R+++
Sbjct: 543 KLKRLRSDNGGEYTSREFEEYCSSHGIRHEKTVPGTPQHNGVAERMNRTIVEKVRSMLRM 602
Query: 621 SNLTLEFWSFAVIHAVCLMNLLPTKVLSYSSPFIILHNLKPDIEHLKVFGSLCYATSLTA 680
+ L FW AV A L+N P+ L++ P + N + HLKVFG +A
Sbjct: 603 AKLPKSFWGEAVQTACYLINRSPSVPLAFEIPERVWTNKEVSYSHLKVFGCRAFAHVPKE 662
Query: 681 HRTKFPLRARKCIVLGQKEGGVKGYLMFDLKTREVFLSRNVIFYET 726
RTK ++ CI +G + GY ++D ++V SR+V+F E+
Sbjct: 663 QRTKLDDKSIPCIFIGYGDEEF-GYRLWDPVKKKVIRSRDVVFRES 707
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 123 bits (309), Expect = 2e-27
Identities = 92/317 (29%), Positives = 152/317 (47%), Gaps = 23/317 (7%)
Query: 428 NHFHSVNMSTVSSSNKLPN---IWHHRLGHPSQPCYISL--------QQMYPPITTSPCK 476
N+ +N S + K N +WH R GH S + + Q + + S C+
Sbjct: 395 NNVPVINFQAYSINAKHKNNFRLWHERFGHISDGKLLEIKRKNMFSDQSLLNNLELS-CE 453
Query: 477 NCDTCHFAKQKRLSFSLRN--TIYGTVLELLHVDIWGPYSVASITGAKYFHIIVYDKSRF 534
C+ C KQ RL F T L ++H D+ GP + ++ YF I V + +
Sbjct: 454 ICEPCLNGKQARLPFKQLKDKTHIKRPLFVVHSDVCGPITPVTLDDKNYFVIFVDQFTHY 513
Query: 535 TWIKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFA---MTDFYSQHGVQHQL 591
L+K KS+ + Q F+ S+ + V + +DNG E+ M F + G+ + L
Sbjct: 514 CVTYLIKYKSDVFSMFQDFVAKSEAHFNLKVVYLYIDNGREYLSNEMRQFCVKKGISYHL 573
Query: 592 SCVKTPQQNGVVERKHQHILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVL--SY 649
+ TPQ NGV ER + I AR ++ + L FW AV+ A L+N +P++ L S
Sbjct: 574 TVPHTPQLNGVSERMIRTITEKARTMVSGAKLDKSFWGEAVLTATYLINRIPSRALVDSS 633
Query: 650 SSPFIILHNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLMFD 709
+P+ + HN KP ++HL+VFG+ Y + + KF ++ K I +G + G K ++D
Sbjct: 634 KTPYEMWHNKKPYLKHLRVFGATVY-VHIKNKQGKFDDKSFKSIFVGYEPNGFK---LWD 689
Query: 710 LKTREVFLSRNVIFYET 726
+ ++R+V+ ET
Sbjct: 690 AVNEKFIVARDVVVDET 706
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 64.3 bits (155), Expect = 1e-09
Identities = 42/144 (29%), Positives = 71/144 (49%), Gaps = 5/144 (3%)
Query: 507 VDIWGPYSVASITGAKYFHIIVYDKSRF--TWIKLMKAKSEARAALQTFILYSQRQYGSL 564
+DI+GP S ++ +Y I+V + +R+ T K A ++ I Y + Q+
Sbjct: 630 MDIFGPVSSSNADTKRYMLIMVDNNTRYCMTSTHFNKNAETILAQVRKNIQYVETQFDRK 689
Query: 565 VKIVRLDNGVEFA---MTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARALMLQS 621
V+ + D G EF + +++ G+ H L+ + NG ER + I+ A L+ QS
Sbjct: 690 VREINSDRGTEFTNDQIEEYFISKGIHHILTSTQDHAANGRAERYIRTIITDATTLLRQS 749
Query: 622 NLTLEFWSFAVIHAVCLMNLLPTK 645
NL ++FW +AV A + N L K
Sbjct: 750 NLRVKFWEYAVTSATNIRNYLEHK 773
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 62.0 bits (149), Expect = 6e-09
Identities = 66/288 (22%), Positives = 119/288 (40%), Gaps = 48/288 (16%)
Query: 419 WSFHLSLVSNHFHSVNMSTVSSS---NKLPN-IWHHRLGHPSQPCYISLQQM-------- 466
W L+ +H + ++ V+ S NK P + H LGH + + S+Q+
Sbjct: 561 WLSKKYLIPSHISKLTINNVNKSKSVNKYPYPLIHRMLGHAN---FRSIQKSLKKNAVTY 617
Query: 467 -------YPPITTSPCKNCDTCHFAKQKRLSFS-LRNTIYGTVLELLHVDIWGPYSVASI 518
+ +T C +C K + + S L+ + LH DI+GP
Sbjct: 618 LKESDIEWSNASTYQCPDCLIGKSTKHRHVKGSRLKYQESYEPFQYLHTDIFGPVHHLPK 677
Query: 519 TGAKYFHIIVYDKSRFTWI-KLMKAKSEARAALQTFIL-YSQRQYGSLVKIVRLDNGVEF 576
+ YF +K+RF W+ L + E+ + T IL + + Q+ + V ++++D G E+
Sbjct: 678 SAPSYFISFTDEKTRFQWVYPLHDRREESILNVFTSILAFIKNQFNARVLVIQMDRGSEY 737
Query: 577 A---MTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARALMLQSNLTLEFWSFAVI 633
+ F++ G+ + + +GV ER ++ +LN R L+ S L W AV
Sbjct: 738 TNKTLHKFFTNRGITACYTTTADSRAHGVAERLNRTLLNDCRTLLHCSGLPNHLWFSAVE 797
Query: 634 HAVCLMNLLP-------------------TKVLSYSSPFIILHNLKPD 662
+ + N L T +L + P +I++N PD
Sbjct: 798 FSTIIRNSLVSPKNDKSARQHAGLAGLDITTILPFGQP-VIVNNHNPD 844
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 48.9 bits (115), Expect = 5e-05
Identities = 31/145 (21%), Positives = 65/145 (44%), Gaps = 5/145 (3%)
Query: 503 ELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARA--ALQTFILYSQRQ 560
+ LH DI+GP + YF + ++F W+ + + E T + + + Q
Sbjct: 239 QYLHTDIFGPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 298
Query: 561 YGSLVKIVRLDNGVEF---AMTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARAL 617
+ + V ++++D G E+ + F ++G+ + + +GV ER ++ +L+ R
Sbjct: 299 FQASVLVIQMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQ 358
Query: 618 MLQSNLTLEFWSFAVIHAVCLMNLL 642
+ S L W A+ + + N L
Sbjct: 359 LQCSGLPNHLWFSAIEFSTIVRNSL 383
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 48.9 bits (115), Expect = 5e-05
Identities = 31/145 (21%), Positives = 65/145 (44%), Gaps = 5/145 (3%)
Query: 503 ELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARA--ALQTFILYSQRQ 560
+ LH DI+GP + YF + ++F W+ + + E T + + + Q
Sbjct: 239 QYLHTDIFGPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 298
Query: 561 YGSLVKIVRLDNGVEF---AMTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARAL 617
+ + V ++++D G E+ + F ++G+ + + +GV ER ++ +L+ R
Sbjct: 299 FQASVLVIQMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQ 358
Query: 618 MLQSNLTLEFWSFAVIHAVCLMNLL 642
+ S L W A+ + + N L
Sbjct: 359 LQCSGLPNHLWFSAIEFSTIVRNSL 383
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 48.9 bits (115), Expect = 5e-05
Identities = 31/145 (21%), Positives = 65/145 (44%), Gaps = 5/145 (3%)
Query: 503 ELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARA--ALQTFILYSQRQ 560
+ LH DI+GP + YF + ++F W+ + + E T + + + Q
Sbjct: 239 QYLHTDIFGPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 298
Query: 561 YGSLVKIVRLDNGVEF---AMTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARAL 617
+ + V ++++D G E+ + F ++G+ + + +GV ER ++ +L+ R
Sbjct: 299 FQASVLVIQMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQ 358
Query: 618 MLQSNLTLEFWSFAVIHAVCLMNLL 642
+ S L W A+ + + N L
Sbjct: 359 LQCSGLPNHLWFSAIEFSTIVRNSL 383
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 48.9 bits (115), Expect = 5e-05
Identities = 31/145 (21%), Positives = 65/145 (44%), Gaps = 5/145 (3%)
Query: 503 ELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARA--ALQTFILYSQRQ 560
+ LH DI+GP + YF + ++F W+ + + E T + + + Q
Sbjct: 239 QYLHTDIFGPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 298
Query: 561 YGSLVKIVRLDNGVEF---AMTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARAL 617
+ + V ++++D G E+ + F ++G+ + + +GV ER ++ +L+ R
Sbjct: 299 FQASVLVIQMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQ 358
Query: 618 MLQSNLTLEFWSFAVIHAVCLMNLL 642
+ S L W A+ + + N L
Sbjct: 359 LQCSGLPNHLWFSAIEFSTIVRNSL 383
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 48.9 bits (115), Expect = 5e-05
Identities = 31/145 (21%), Positives = 65/145 (44%), Gaps = 5/145 (3%)
Query: 503 ELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARA--ALQTFILYSQRQ 560
+ LH DI+GP + YF + ++F W+ + + E T + + + Q
Sbjct: 666 QYLHTDIFGPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 725
Query: 561 YGSLVKIVRLDNGVEF---AMTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARAL 617
+ + V ++++D G E+ + F ++G+ + + +GV ER ++ +L+ R
Sbjct: 726 FQASVLVIQMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQ 785
Query: 618 MLQSNLTLEFWSFAVIHAVCLMNLL 642
+ S L W A+ + + N L
Sbjct: 786 LQCSGLPNHLWFSAIEFSTIVRNSL 810
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 48.9 bits (115), Expect = 5e-05
Identities = 31/145 (21%), Positives = 65/145 (44%), Gaps = 5/145 (3%)
Query: 503 ELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARA--ALQTFILYSQRQ 560
+ LH DI+GP + YF + ++F W+ + + E T + + + Q
Sbjct: 666 QYLHTDIFGPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 725
Query: 561 YGSLVKIVRLDNGVEF---AMTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARAL 617
+ + V ++++D G E+ + F ++G+ + + +GV ER ++ +L+ R
Sbjct: 726 FQASVLVIQMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQ 785
Query: 618 MLQSNLTLEFWSFAVIHAVCLMNLL 642
+ S L W A+ + + N L
Sbjct: 786 LQCSGLPNHLWFSAIEFSTIVRNSL 810
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-); Reverse
transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 40.4 bits (93), Expect = 0.017
Identities = 61/285 (21%), Positives = 113/285 (39%), Gaps = 40/285 (14%)
Query: 475 CKNCDTCHFAKQKRLSFSLRNTIYGTVLELLHVDIWGPYSVASITGAKYFHIIVYDKSRF 534
C+ C K + ++ T + + VD GP S G +Y ++ D +++
Sbjct: 939 CQKCQKAKTTKHTKTPMTITETPEHA-FDRVVVDTIGPLP-KSENGNEYAVTLICDLTKY 996
Query: 535 TW---IKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFA---MTDFYSQHGVQ 588
I AK+ A+A ++FIL +YG + + D G E+ +TD ++
Sbjct: 997 LVAIPIANKSAKTVAKAIFESFIL----KYGPMKTFIT-DMGTEYKNSIITDLCKYLKIK 1051
Query: 589 HQLSCVKTPQQNGVVERKHQHILNIARALMLQSNLTLEFW----------SFAVIHAVCL 638
+ S Q GVVER H+ + R+ + + W + +++H C
Sbjct: 1052 NITSTAHHHQTVGVVERSHRTLNEYIRSYISTDKTDWDVWLQYFVYCFNTTQSMVHNYCP 1111
Query: 639 MNLLPTKVLSYSSPFIILHNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQK 698
L+ + + F LH+++P ++ YA RARK ++ K
Sbjct: 1112 YELVFGRTSNLPKHFNKLHSIEP------IYNIDDYAKESKYRLEVAYARARK-LLEAHK 1164
Query: 699 EGGVKGYLMFDLKTREV-------FLSRNVIFYETIFPFQATDKV 736
E + Y DLK +++ L RN + ++ F + K+
Sbjct: 1165 EKNKENY---DLKVKDIELEVGDKVLLRNEVGHKLDFKYTGPYKI 1206
>M300_ARATH (P93293) Hypothetical mitochondrial protein AtMg00300
(ORF145a) (ORF1451)
Length = 145
Score = 38.9 bits (89), Expect = 0.051
Identities = 23/71 (32%), Positives = 32/71 (44%), Gaps = 2/71 (2%)
Query: 447 IWHHRLGHPSQPCYISLQQ--MYPPITTSPCKNCDTCHFAKQKRLSFSLRNTIYGTVLEL 504
+WH RL H SQ L + S K C+ C + K R++FS L+
Sbjct: 71 LWHSRLAHMSQRGMELLVKKGFLDSSKVSSLKFCEDCIYGKTHRVNFSTGQHTTKNPLDY 130
Query: 505 LHVDIWGPYSV 515
+H D+WG SV
Sbjct: 131 VHSDLWGAPSV 141
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 38.5 bits (88), Expect = 0.066
Identities = 31/119 (26%), Positives = 54/119 (45%), Gaps = 13/119 (10%)
Query: 506 HVDIWGPYSVASITGAKYFHIIVYDKSR--FTWIKLMKAKSEARAALQTFILYSQRQYGS 563
++D GP ++ Y H++V S F W+ KA S + +L S +
Sbjct: 888 YIDYIGPLPPSN----GYLHVLVVVDSMTGFVWLYPTKAPSTSATVKALNMLTSI----A 939
Query: 564 LVKIVRLDNGVEFAMT---DFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARALML 619
+ K++ D G F + D+ + G+Q + S PQ +G VERK+ I + L++
Sbjct: 940 IPKVLHSDQGAAFTSSTFADWAKEKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTKLLI 998
>POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1157
Score = 37.4 bits (85), Expect = 0.15
Identities = 28/102 (27%), Positives = 45/102 (43%), Gaps = 9/102 (8%)
Query: 523 YFHIIVYDKSR--FTWIKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFA--- 577
Y H++V S F W+ KA S + +L S ++ K++ D G F
Sbjct: 903 YLHVLVVVDSMTGFVWLYPTKAPSTSATVKALNMLTSI----AVPKVIHSDQGAAFTSAT 958
Query: 578 MTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARALML 619
D+ G+Q + S PQ +G VERK+ I + L++
Sbjct: 959 FADWAKNKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTKLLV 1000
>POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 886
Score = 37.4 bits (85), Expect = 0.15
Identities = 29/116 (25%), Positives = 49/116 (42%), Gaps = 9/116 (7%)
Query: 507 VDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARAALQTFILYSQRQYGSLVK 566
+D GP + G Y ++V + FTW+ KA S + +L S ++ K
Sbjct: 681 IDYIGPLPPSQ--GYLYVLVVVDGMTGFTWLYPTKAPSTSATVKSLNVLTSI----AIPK 734
Query: 567 IVRLDNGVEFAMTDFYS---QHGVQHQLSCVKTPQQNGVVERKHQHILNIARALML 619
++ D G F + F + G+ + S PQ VERK+ I + L++
Sbjct: 735 VIHSDQGAAFTSSTFAEWAKERGIHLEFSTPYHPQSGSKVERKNSDIKRLLTKLLV 790
>YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II
Length = 1268
Score = 36.6 bits (83), Expect = 0.25
Identities = 25/104 (24%), Positives = 45/104 (43%), Gaps = 12/104 (11%)
Query: 505 LHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARAALQTFILYSQRQYGSL 564
+H+D GP + Y ++V K+++ +KL ++ S ++S Y
Sbjct: 852 IHIDFAGPLNGC------YLLVVVDAKTKYAEVKLTRSISAVTTIDLLEEIFSIHGYP-- 903
Query: 565 VKIVRLDNGVEFA---MTDFYSQHGVQHQLSCVKTPQQNGVVER 605
+ + DNG + HG++H+ S V P+ NG ER
Sbjct: 904 -ETIISDNGTQLTSHLFAQMCQSHGIEHKTSAVYYPRSNGAAER 946
>POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1165
Score = 35.0 bits (79), Expect = 0.73
Identities = 27/105 (25%), Positives = 46/105 (43%), Gaps = 5/105 (4%)
Query: 520 GAKYFHIIVYDKSRFTWIKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFAMT 579
G KY ++V+ + W++ K+E + IL + K++ DNG F
Sbjct: 893 GNKY--LLVFIDTFSGWVEAFPTKTETALIVCKKILEEILPRFGIPKVLGSDNGPAFVAQ 950
Query: 580 ---DFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARALMLQS 621
+Q G+ +L C PQ +G VER ++ I L L++
Sbjct: 951 VSQGLATQLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLALET 995
>POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 1046
Score = 35.0 bits (79), Expect = 0.73
Identities = 26/106 (24%), Positives = 44/106 (40%), Gaps = 3/106 (2%)
Query: 521 AKYFHIIVYDKSRFTWIKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFAMT- 579
A Y +++V+ + W++ + E + IL L K++ DNG F
Sbjct: 777 AGYKYLLVFVDTFSGWVEAYPTRQETAHMVAKKILEEIFPRFGLPKVIGSDNGPAFVSQV 836
Query: 580 --DFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARALMLQSNL 623
G+ +L C PQ +G VER ++ I L L++ L
Sbjct: 837 SQGLARTLGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTLETGL 882
>POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1189
Score = 35.0 bits (79), Expect = 0.73
Identities = 28/108 (25%), Positives = 46/108 (41%), Gaps = 7/108 (6%)
Query: 521 AKYFHIIVYDKSRFTWIKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFAMTD 580
A Y +++V+ + W++ + E + IL L K++ DNG F
Sbjct: 920 AGYKYLLVFVDTFSGWVEAFPTRQETAHIVAKKILEEIFPRFGLPKVIGSDNGPAFVSQ- 978
Query: 581 FYSQH-----GVQHQLSCVKTPQQNGVVERKHQHILNIARALMLQSNL 623
SQ G+ +L C PQ +G VER ++ I L L++ L
Sbjct: 979 -VSQGLARILGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTLETGL 1025
>POL_JSRV (P31623) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 870
Score = 33.5 bits (75), Expect = 2.1
Identities = 30/128 (23%), Positives = 50/128 (38%), Gaps = 14/128 (10%)
Query: 522 KYFHIIVYDKSRFTWIKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFAMTDF 581
KY H+ + S F L +S +S + + ++ DNG + F
Sbjct: 666 KYVHVSIDTFSNFLMASLHTGESTRHCIQHLLFCFST---SGIPQTLKTDNGPGYTSRSF 722
Query: 582 YS---QHGVQHQLSCVKTPQQNGVVERKHQHILNIARALMLQSNLTLEFWS----FAVIH 634
+ H+ PQ G+VER HQ I + +L+ E +S A+ H
Sbjct: 723 QRFCLSFQIHHKTGIPYNPQGQGIVERAHQRI----KHQLLKQKKGNELYSPSPHNALNH 778
Query: 635 AVCLMNLL 642
A+ ++N L
Sbjct: 779 ALYVLNFL 786
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.337 0.145 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 80,573,113
Number of Sequences: 164201
Number of extensions: 3192281
Number of successful extensions: 12668
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 12633
Number of HSP's gapped (non-prelim): 37
length of query: 754
length of database: 59,974,054
effective HSP length: 118
effective length of query: 636
effective length of database: 40,598,336
effective search space: 25820541696
effective search space used: 25820541696
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0296a.5


