

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0290.7
(47 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
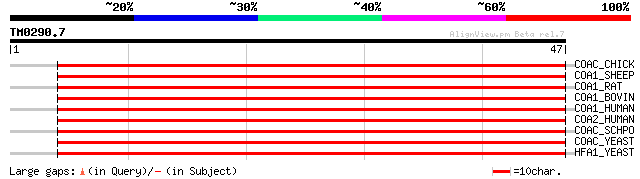
COAC_CHICK (P11029) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC) [I... 63 1e-10
COA1_SHEEP (Q28559) Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-a... 63 1e-10
COA1_RAT (P11497) Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-alp... 63 1e-10
COA1_BOVIN (Q9TTS3) Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-a... 63 1e-10
COA1_HUMAN (Q13085) Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-a... 60 8e-10
COA2_HUMAN (O00763) Acetyl-CoA carboxylase 2 (EC 6.4.1.2) (ACC-b... 58 6e-09
COAC_SCHPO (P78820) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC) [I... 55 3e-08
COAC_YEAST (Q00955) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC) [I... 54 8e-08
HFA1_YEAST (P32874) HFA1 protein 53 2e-07
>COAC_CHICK (P11029) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC)
[Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2324
Score = 63.2 bits (152), Expect = 1e-10
Identities = 27/43 (62%), Positives = 31/43 (71%)
Query: 5 NPDYGFNPTGWEVQELGFKSKPNVWAYFSVKSGGGIHEFLDSQ 47
NPD GF P+ VQEL F+S NVW YFSV + GG+HEF DSQ
Sbjct: 514 NPDEGFKPSSGTVQELNFRSNKNVWGYFSVAAAGGLHEFADSQ 556
>COA1_SHEEP (Q28559) Acetyl-CoA carboxylase 1 (EC 6.4.1.2)
(ACC-alpha) [Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2346
Score = 63.2 bits (152), Expect = 1e-10
Identities = 27/43 (62%), Positives = 31/43 (71%)
Query: 5 NPDYGFNPTGWEVQELGFKSKPNVWAYFSVKSGGGIHEFLDSQ 47
NPD GF P+ VQEL F+S NVW YFSV + GG+HEF DSQ
Sbjct: 514 NPDEGFKPSSGTVQELNFRSNKNVWGYFSVAAAGGLHEFADSQ 556
>COA1_RAT (P11497) Acetyl-CoA carboxylase 1 (EC 6.4.1.2) (ACC-alpha)
[Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2345
Score = 63.2 bits (152), Expect = 1e-10
Identities = 27/43 (62%), Positives = 31/43 (71%)
Query: 5 NPDYGFNPTGWEVQELGFKSKPNVWAYFSVKSGGGIHEFLDSQ 47
NPD GF P+ VQEL F+S NVW YFSV + GG+HEF DSQ
Sbjct: 513 NPDEGFKPSSGTVQELNFRSNKNVWGYFSVAAAGGLHEFADSQ 555
>COA1_BOVIN (Q9TTS3) Acetyl-CoA carboxylase 1 (EC 6.4.1.2)
(ACC-alpha) [Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2346
Score = 63.2 bits (152), Expect = 1e-10
Identities = 27/43 (62%), Positives = 31/43 (71%)
Query: 5 NPDYGFNPTGWEVQELGFKSKPNVWAYFSVKSGGGIHEFLDSQ 47
NPD GF P+ VQEL F+S NVW YFSV + GG+HEF DSQ
Sbjct: 514 NPDEGFKPSSGTVQELNFRSNKNVWGYFSVAAAGGLHEFADSQ 556
>COA1_HUMAN (Q13085) Acetyl-CoA carboxylase 1 (EC 6.4.1.2)
(ACC-alpha) [Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2346
Score = 60.5 bits (145), Expect = 8e-10
Identities = 26/43 (60%), Positives = 30/43 (69%)
Query: 5 NPDYGFNPTGWEVQELGFKSKPNVWAYFSVKSGGGIHEFLDSQ 47
NPD GF P+ VQEL F+S NVW YFSV + GG+HEF SQ
Sbjct: 514 NPDEGFKPSSGTVQELNFRSNKNVWGYFSVAAAGGLHEFAGSQ 556
>COA2_HUMAN (O00763) Acetyl-CoA carboxylase 2 (EC 6.4.1.2)
(ACC-beta) [Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2483
Score = 57.8 bits (138), Expect = 6e-09
Identities = 25/43 (58%), Positives = 30/43 (69%)
Query: 5 NPDYGFNPTGWEVQELGFKSKPNVWAYFSVKSGGGIHEFLDSQ 47
NPD GF P+ VQEL F+S NVW YF+V + GG+HEF SQ
Sbjct: 657 NPDEGFKPSSGTVQELNFRSSKNVWGYFTVAATGGLHEFAISQ 699
>COAC_SCHPO (P78820) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC)
[Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2280
Score = 55.5 bits (132), Expect = 3e-08
Identities = 23/43 (53%), Positives = 30/43 (69%)
Query: 5 NPDYGFNPTGWEVQELGFKSKPNVWAYFSVKSGGGIHEFLDSQ 47
+P GF P+ +++L F+S NVW YFSV + GGIHEF DSQ
Sbjct: 473 DPGEGFKPSSGMIKDLNFRSSSNVWGYFSVGTAGGIHEFADSQ 515
>COAC_YEAST (Q00955) Acetyl-CoA carboxylase (EC 6.4.1.2) (ACC)
[Includes: Biotin carboxylase (EC 6.3.4.14)]
Length = 2233
Score = 53.9 bits (128), Expect = 8e-08
Identities = 23/43 (53%), Positives = 29/43 (66%)
Query: 5 NPDYGFNPTGWEVQELGFKSKPNVWAYFSVKSGGGIHEFLDSQ 47
+P+ GF P+G + EL F+S NVW YFSV + G IH F DSQ
Sbjct: 463 DPNDGFKPSGGTLHELNFRSSSNVWGYFSVGNNGNIHSFSDSQ 505
>HFA1_YEAST (P32874) HFA1 protein
Length = 2273
Score = 52.8 bits (125), Expect = 2e-07
Identities = 22/43 (51%), Positives = 29/43 (67%)
Query: 5 NPDYGFNPTGWEVQELGFKSKPNVWAYFSVKSGGGIHEFLDSQ 47
+P+ GF P+ ++ EL F+S NVW YFSV + G IH F DSQ
Sbjct: 531 DPNEGFKPSTGKIHELNFRSSSNVWGYFSVGNNGAIHSFSDSQ 573
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.142 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,846,929
Number of Sequences: 164201
Number of extensions: 200105
Number of successful extensions: 296
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 287
Number of HSP's gapped (non-prelim): 9
length of query: 47
length of database: 59,974,054
effective HSP length: 23
effective length of query: 24
effective length of database: 56,197,431
effective search space: 1348738344
effective search space used: 1348738344
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0290.7


