

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0288.8
(846 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
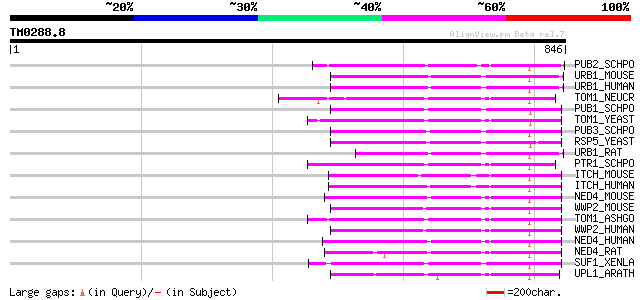
Score E
Sequences producing significant alignments: (bits) Value
PUB2_SCHPO (Q9UTG2) E3 ubiquitin--protein ligase pub2 (EC 6.3.2.-) 199 3e-50
URB1_MOUSE (Q7TMY8) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2... 195 5e-49
URB1_HUMAN (Q7Z6Z7) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2... 195 5e-49
TOM1_NEUCR (Q9P4Z1) E3 ubiquitin protein ligase TOM1-like protei... 193 2e-48
PUB1_SCHPO (Q92462) E3 ubiquitin--protein ligase pub1 (EC 6.3.2.-) 191 7e-48
TOM1_YEAST (Q03280) E3 ubiquitin protein ligase TOM1 (EC 6.3.2.-... 190 1e-47
PUB3_SCHPO (O14326) E3 ubiquitin--protein ligase pub3 (EC 6.3.2.-) 190 2e-47
RSP5_YEAST (P39940) E3 ubiquitin--protein ligase RSP5 (EC 6.3.2.-) 189 3e-47
URB1_RAT (P51593) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2.-... 187 1e-46
PTR1_SCHPO (O13834) E3 ubiquitin protein ligase ptr1 (EC 6.3.2.-... 185 5e-46
ITCH_MOUSE (Q8C863) Itchy E3 ubiquitin protein ligase (EC 6.3.2.-) 182 3e-45
ITCH_HUMAN (Q96J02) Itchy homolog E3 ubiquitin protein ligase (E... 182 4e-45
NED4_MOUSE (P46935) E3 ubiquitin-protein ligase Nedd-4 (EC 6.3.2.-) 179 3e-44
WWP2_MOUSE (Q9DBH0) Nedd-4-like E3 ubiquitin-protein ligase WWP2... 178 6e-44
TOM1_ASHGO (Q756G2) Probable E3 ubiquitin protein ligase TOM1 (E... 178 6e-44
WWP2_HUMAN (O00308) Nedd-4-like E3 ubiquitin-protein ligase WWP2... 177 1e-43
NED4_HUMAN (P46934) E3 ubiquitin-protein ligase Nedd-4 (EC 6.3.2.-) 176 2e-43
NED4_RAT (Q62940) E3 ubiquitin-protein ligase Nedd-4 (EC 6.3.2.-) 175 4e-43
SUF1_XENLA (Q9PUN2) Smad ubiquitination regulatory factor 1 (EC ... 173 2e-42
UPL1_ARATH (Q8GY23) E3 ubiquitin protein ligase UPL1 (EC 6.3.2.-... 173 2e-42
>PUB2_SCHPO (Q9UTG2) E3 ubiquitin--protein ligase pub2 (EC 6.3.2.-)
Length = 671
Score = 199 bits (506), Expect = 3e-50
Identities = 124/390 (31%), Positives = 202/390 (51%), Gaps = 19/390 (4%)
Query: 462 NFESRRHLAMMMF-PEVKEDYEELHEMLIDRSQLLSESFEYIARADTASLHAGLFMEFKN 520
N E +R +A M PE+ + +L ++ + R+ ++++ I++ + + L + F+N
Sbjct: 294 NDEYQRKIAYMYDRPEMAVNDAQL-QLKVSRATTFEDAYDIISKLSVSDMKKKLLIRFRN 352
Query: 521 EEATG-PGVLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGR 579
E+ GV RE+F ++ A+FNP +LF +D + S V+P YF F GR
Sbjct: 353 EDGLDYGGVSREFFYILSHAIFNPGYSLFEYATDDNYGLQISPLSSVNPDFRSYFRFVGR 412
Query: 580 VIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDA 639
V+ LA+ H+ + + F+ ++ K + LEDVKD D Y S K I D D ++
Sbjct: 413 VMGLAIYHRRYLDVQFVLPFYKRILQKPLCLEDVKDVDEVYYESLKWIKNNDVD----ES 468
Query: 640 LGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKG 699
L L F E G++ V+L P G+N+ VN++N+ Y+ L + + VTS EQ + G
Sbjct: 469 LCLNFSVEENRFGESVTVDLIPNGRNIAVNNQNKMNYLKALTEHKLVTSTEEQFNALKGG 528
Query: 700 FTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWE 759
+++ S Q F +LD +L+G D I V+DWK T+Y Y ETD + WFWE
Sbjct: 529 LNELIPDSVLQIF-----NENELDTLLNGKRD-IDVQDWKRFTDYRSYTETDDIVIWFWE 582
Query: 760 IVGRMTAEQRKVLLFFWTSVKYLPVEGFCGL----ASRLYIYKSPEPEDRLPSSHTCFYR 815
++ + E++ LL F T LP+ GF + R + + +LP +HTCF R
Sbjct: 583 LLSEWSPEKKAKLLQFATGTSRLPLSGFKDMHGSDGPRKFTIEKVGHISQLPKAHTCFNR 642
Query: 816 MCFPAYSSMTVMQERLEVITQEHIGCSFGT 845
+ P Y+S ++++L + QE G FGT
Sbjct: 643 LDIPPYNSKEELEQKLTIAIQETAG--FGT 670
>URB1_MOUSE (Q7TMY8) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2.-)
(Fragment)
Length = 2749
Score = 195 bits (495), Expect = 5e-49
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 2396 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 2455
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 2456 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 2515
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 2516 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 2572
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 2573 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 2626
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 2627 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 2686
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 2687 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 2743
Query: 843 FG 844
G
Sbjct: 2744 EG 2745
>URB1_HUMAN (Q7Z6Z7) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2.-)
(HSPC272)
Length = 3360
Score = 195 bits (495), Expect = 5e-49
Identities = 115/362 (31%), Positives = 183/362 (49%), Gaps = 18/362 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNAL 547
+ R + +S+ + R + L++ F+ EE G +LREW++++ + +FNP AL
Sbjct: 3007 VRRDHVFEDSYRELHRKSPEEMKNRLYIVFEGEEGQDAGGLLREWYMIISREMFNPMYAL 3066
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F P DR + N +S +P HL YF F GR++A A+ + R F+ + GK
Sbjct: 3067 FRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVYDNRLLECYFTRSFYKHILGKS 3126
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ D++ D Y +LE D + D LTF EV+E G +V +L P G N++
Sbjct: 3127 VRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFSTEVQEFGVCEVRDLKPNGANIL 3183
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+++Y+ L+ + R +I +Q++ F +GF +I+ F Q LEL L
Sbjct: 3184 VTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPKRLISIFTEQELEL------LI 3237
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
TI ++D K++TEY+ Y+ IQI WFW + R L F T +P++GF
Sbjct: 3238 SGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQADRAKFLQFVTGTSKVPLQGF 3297
Query: 788 CGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIGCS 842
L + I++ DRLPS+HTCF ++ PAY S ++ L + QE CS
Sbjct: 3298 AALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAYESFEKLRHMLLLAIQE---CS 3354
Query: 843 FG 844
G
Sbjct: 3355 EG 3356
>TOM1_NEUCR (Q9P4Z1) E3 ubiquitin protein ligase TOM1-like protein (EC
6.3.2.-)
Length = 4065
Score = 193 bits (490), Expect = 2e-48
Identities = 128/436 (29%), Positives = 218/436 (49%), Gaps = 26/436 (5%)
Query: 410 KELYLISKLYDGAEEKLWKVLTRQRSVVCLLIVRYAKR--TDEHQWILEHKCVTNFESRR 467
KE+ L S + L+ T + + +VR+ + + ++++ V F+++R
Sbjct: 3628 KEMLLTSPPPEDRIAGLFFTFTEEHRRILNELVRHNPKLMSGTFSLLVKNPKVLEFDNKR 3687
Query: 468 HL----AMMMFPEVKEDYEELHEMLIDRSQLLSESFE--YIARADTASLHAGLFMEFKNE 521
+ + + + + L ++ + R + +SF Y +AD L + F+ E
Sbjct: 3688 NYFNRSVHSKYQQTRHSFPPL-QLQVRREHVFHDSFRSLYYKKADELKF-GKLNIRFQGE 3745
Query: 522 EATGPG-VLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRV 580
E G V REWF ++ + +F+P LFV +DR F+PN S ++ HL +F F GR+
Sbjct: 3746 EGVDAGGVTREWFQVLSRQMFDPNYVLFVPVSSDRTTFHPNKLSPINDEHLPFFKFIGRI 3805
Query: 581 IALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDAL 640
I AL + R + ++ GK ++++D++ DP Y S +LE D +D +
Sbjct: 3806 IGKALYEGRLLECYFSRAVYKRILGKPVSVKDMESFDPDYYKSLVWMLENDI----TDII 3861
Query: 641 GLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGF 700
TF E + G+ KVV+L G+N+ V +N+ +Y+ L+++ + +TS+ +Q+ F GF
Sbjct: 3862 TETFSVEDDVFGEVKVVDLIENGRNIPVTEENKHEYVRLIVEHKLITSVKDQMKAFLTGF 3921
Query: 701 TDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEI 760
+I+ F Q LEL ++ G D I ++DWKA+TEY+ Y QI WFW
Sbjct: 3922 HEIIPEELIAIFNEQELEL-----LISGLPD-IDIDDWKANTEYHNYSAGAPQIQWFWRA 3975
Query: 761 VGRMTAEQRKVLLFFWTSVKYLPVEGFCGL-----ASRLYIYKSPEPEDRLPSSHTCFYR 815
V E+ LL F T +P+ GF L SR I++ +DRLPSSHTCF +
Sbjct: 3976 VRSFDKEELAKLLQFVTGTSKVPLNGFKELEGMNGVSRFNIHRDYGSKDRLPSSHTCFNQ 4035
Query: 816 MCFPAYSSMTVMQERL 831
+ P Y + ++ +L
Sbjct: 4036 LDLPEYENYETLRSQL 4051
>PUB1_SCHPO (Q92462) E3 ubiquitin--protein ligase pub1 (EC 6.3.2.-)
Length = 767
Score = 191 bits (485), Expect = 7e-48
Identities = 119/358 (33%), Positives = 178/358 (49%), Gaps = 17/358 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPGVL-REWFLLVCQALFNPQNAL 547
+ R+ + +S+ I R L L ++F E+ G L RE+F L+ +FNP L
Sbjct: 417 VRRNHIFEDSYAEIMRQSATDLKKRLMIKFDGEDGLDYGGLSREYFFLLSHEMFNPFYCL 476
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F D N S ++P HL YF F GRVI LA+ H+ V F+ + K
Sbjct: 477 FEYSSVDNYTLQINPHSGINPEHLNYFKFIGRVIGLAIFHRRFVDAFFVVSFYKMILQKK 536
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+TL+D++ D Y S IL+ D + L LTF E G+ ++L P G+N+
Sbjct: 537 VTLQDMESMDAEYYRSLVWILDNDI----TGVLDLTFSVEDNCFGEVVTIDLKPNGRNIE 592
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+ +Y+DL+ R I EQ + F +GF++++ F + LEL L
Sbjct: 593 VTEENKREYVDLVTVWRIQKRIEEQFNAFHEGFSELIPQELINVFDERELEL------LI 646
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
G I +EDWK HT+Y Y E D I WFWE++ + E++ LL F T +PV GF
Sbjct: 647 GGISEIDMEDWKKHTDYRSYSENDQIIKWFWELMDEWSNEKKSRLLQFTTGTSRIPVNGF 706
Query: 788 CGLAS-----RLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIG 840
L + I K+ EP ++LP +HTCF R+ P Y+S + +L + +E IG
Sbjct: 707 KDLQGSDGPRKFTIEKAGEP-NKLPKAHTCFNRLDLPPYTSKKDLDHKLSIAVEETIG 763
>TOM1_YEAST (Q03280) E3 ubiquitin protein ligase TOM1 (EC 6.3.2.-)
(Temperature dependent-organization in mitotic nucleus
protein 1) (Suppressor of snRNA protein 2)
Length = 3268
Score = 190 bits (483), Expect = 1e-47
Identities = 122/397 (30%), Positives = 209/397 (51%), Gaps = 23/397 (5%)
Query: 454 ILEHKCVTNFESRRHLAMMMFPEVKEDYEELHEMLID--RSQLLSESFEYIA-RADTASL 510
++++ V +F+++R+ ++K D +E ++ I R Q+ +S+ + + +
Sbjct: 2881 LVKNPKVLDFDNKRYF---FNAKLKSDNQERPKLPITVRREQVFLDSYRALFFKTNDEIK 2937
Query: 511 HAGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPL 569
++ L + FK E G V REW+ ++ + +FNP ALF+ P+D+ F+PN S ++P
Sbjct: 2938 NSKLEITFKGESGVDAGGVTREWYQVLSRQMFNPDYALFLPVPSDKTTFHPNRTSGINPE 2997
Query: 570 HLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILE 629
HL +F F G +I A+ + + R + + G+ ++L+D++ DP Y S ILE
Sbjct: 2998 HLSFFKFIGMIIGKAIRDQCFLDCHFSREVYKNILGRPVSLKDMESLDPDYYKSLVWILE 3057
Query: 630 MDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSI 689
D +D + TF E ++ G+ KV+ L GGK++IV N++ Y+ +++ + TS+
Sbjct: 3058 NDI----TDIIEETFSVETDDYGEHKVINLIEGGKDIIVTEANKQDYVKKVVEYKLQTSV 3113
Query: 690 SEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKE 749
EQ+ +F GF ++S F Q LEL ++ G D I V+DWK +T Y Y
Sbjct: 3114 KEQMDNFLVGFYALISKDLITIFDEQELEL-----LISGLPD-IDVDDWKNNTTYVNYTA 3167
Query: 750 TDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGLAS-----RLYIYKSPEPED 804
T ++++FW V AE+R LL F T +P+ GF L+ + I++ +
Sbjct: 3168 TCKEVSYFWRAVRSFDAEERAKLLQFVTGTSKVPLNGFKELSGVNGVCKFSIHRDFGSSE 3227
Query: 805 RLPSSHTCFYRMCFPAYSSM-TVMQERLEVITQEHIG 840
RLPSSHTCF ++ P Y S T+ L I + H G
Sbjct: 3228 RLPSSHTCFNQLNLPPYESYETLRGSLLLAINEGHEG 3264
>PUB3_SCHPO (O14326) E3 ubiquitin--protein ligase pub3 (EC 6.3.2.-)
Length = 786
Score = 190 bits (482), Expect = 2e-47
Identities = 119/357 (33%), Positives = 173/357 (48%), Gaps = 15/357 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPGVL-REWFLLVCQALFNPQNAL 547
+ R + +S+ I R L L + F E+ G L RE+F L+ +F+P L
Sbjct: 436 VRRDHIFEDSYAEIMRYSAHDLKKRLMIRFDGEDGLDYGGLSREFFFLLSHKMFDPIYCL 495
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F D N S ++P HL YF F GRVI LA+ H+ + + +L K
Sbjct: 496 FEYSAVDNYTLQINPHSSINPEHLNYFRFIGRVIGLAIFHRRFLDAFFVVSLYKKLLRKK 555
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
++L D++ D Y S K +LE D I L LTF E + G+ + VEL G+N+
Sbjct: 556 VSLADMESIDAEFYRSLKWVLENDITGI----LDLTFSVEEDHFGEVRTVELITNGENIE 611
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N++KY+DL+ + R + +Q + F GF +++S F + LEL L
Sbjct: 612 VTEENKKKYVDLVTEWRVSKRVEQQFNAFYSGFVELVSPDLVNVFDERELEL------LI 665
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
G + VEDWK+HTEY Y TD I WFWEI+ E R LL F T +PV GF
Sbjct: 666 GGISDVDVEDWKSHTEYRTYIATDPVIKWFWEIIAGWKNEDRSKLLQFATGTSRIPVNGF 725
Query: 788 CGL----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIG 840
L R + + D+LP +HTCF R+ P Y S + E+L + + +G
Sbjct: 726 RDLQGSDGPRKFTIEKAGTPDQLPVAHTCFNRLDLPDYPSKDTLHEKLSLAVENTVG 782
>RSP5_YEAST (P39940) E3 ubiquitin--protein ligase RSP5 (EC 6.3.2.-)
Length = 809
Score = 189 bits (480), Expect = 3e-47
Identities = 118/358 (32%), Positives = 176/358 (48%), Gaps = 17/358 (4%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATG-PGVLREWFLLVCQALFNPQNAL 547
+ R + ++++ I R L L ++F EE GV RE+F L+ +FNP L
Sbjct: 459 VRRKNIFEDAYQEIMRQTPEDLKKRLMIKFDGEEGLDYGGVSREFFFLLSHEMFNPFYCL 518
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F D N S ++P HL YF F GRV+ L + H+ + + + K
Sbjct: 519 FEYSAYDNYTIQINPNSGINPEHLNYFKFIGRVVGLGVFHRRFLDAFFVGALYKMMLRKK 578
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
+ L+D++ D +Y+S +LE D + L LTF + E G+ V+L P G+N+
Sbjct: 579 VVLQDMEGVDAEVYNSLNWMLENSIDGV----LDLTFSADDERFGEVVTVDLKPDGRNIE 634
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V N+++Y++L + R V + EQ F GF +++ F + LEL L
Sbjct: 635 VTDGNKKEYVELYTQWRIVDRVQEQFKAFMDGFNELIPEDLVTVFDERELEL------LI 688
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
G I +EDWK HT+Y GY+E+D I WFW+ V EQR LL F T +PV GF
Sbjct: 689 GGIAEIDIEDWKKHTDYRGYQESDEVIQWFWKCVSEWDNEQRARLLQFTTGTSRIPVNGF 748
Query: 788 CGLAS-----RLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIG 840
L R I K+ E + +LP SHTCF R+ P Y M+++L + +E IG
Sbjct: 749 KDLQGSDGPRRFTIEKAGEVQ-QLPKSHTCFNRVDLPQYVDYDSMKQKLTLAVEETIG 805
>URB1_RAT (P51593) E3 ubiquitin protein ligase URE-B1 (EC 6.3.2.-)
(Upstream regulatory element binding protein 1)
(Fragment)
Length = 322
Score = 187 bits (474), Expect = 1e-46
Identities = 108/323 (33%), Positives = 167/323 (51%), Gaps = 17/323 (5%)
Query: 527 GVLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALM 586
G+LREW++++ + +FNP ALF P DR + N +S +P HL YF F GR++A A+
Sbjct: 8 GLLREWYMIISREMFNPMYALFRTSPGDRVTYTINPSSHCNPNHLSYFKFVGRIVAKAVY 67
Query: 587 HKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVR 646
+ R F+ + GK + D++ D Y +LE D + D LTF
Sbjct: 68 DNRLLECYFTRSFYKHILGKSVRYTDMESEDYHFYQGLVYLLENDVSTLGYD---LTFST 124
Query: 647 EVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSS 706
EV+E G +V +L P G N++V +N+++Y+ L+ + R +I +Q++ F +GF +I+
Sbjct: 125 EVQEFGVCEVRDLKPNGANILVTEENKKEYVHLVCQMRMTGAIRKQLAAFLEGFYEIIPK 184
Query: 707 SKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTA 766
F Q LEL L TI ++D K++TEY+ Y+ IQI WFW +
Sbjct: 185 RLISIFTEQELEL------LISGLPTIDIDDLKSNTEYHKYQSNSIQIQWFWRALRSFDQ 238
Query: 767 EQRKVLLFFWTSVKYLPVEGFCGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAY 821
R L F T +P++GF L + I++ DRLPS+HTCF ++ PAY
Sbjct: 239 ADRAKFLQFVTGTSKVPLQGFAALEGMNGIQKFQIHRDDRSTDRLPSAHTCFNQLDLPAY 298
Query: 822 SSMTVMQERLEVITQEHIGCSFG 844
S ++ L + QE CS G
Sbjct: 299 ESFEKLRHMLLLAIQE---CSEG 318
>PTR1_SCHPO (O13834) E3 ubiquitin protein ligase ptr1 (EC 6.3.2.-)
(Poly(A)+ RNA transport protein 1)
Length = 3227
Score = 185 bits (469), Expect = 5e-46
Identities = 117/387 (30%), Positives = 195/387 (50%), Gaps = 20/387 (5%)
Query: 454 ILEHKCVTNFESRRHLAMMMFPE--VKEDYEELHEMLIDRSQLLSESFEYIARADTASLH 511
++++ V FE++R+ E KE Y L+ + + R + +S+ + D +
Sbjct: 2838 LVKNPKVLEFENKRNYFNRQLHEEAAKEQYPPLN-ITVRRDHVFLDSYRALHFKDADEVK 2896
Query: 512 -AGLFMEFKNEEATGPG-VLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPL 569
+ L + F++EE G V REW ++ + +FNP ALF+ D F+PN S V+P
Sbjct: 2897 FSKLNIHFRDEEGVDAGGVTREWLQVLARQMFNPDYALFLPVTGDATTFHPNRDSSVNPD 2956
Query: 570 HLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILE 629
HL +F F+GR+I AL + R + + + ++++D++ DP Y S +L
Sbjct: 2957 HLSFFKFTGRIIGKALYDGRLLDCHFSRAVYKHMLHRSVSVKDIESLDPDYYKSLVWMLN 3016
Query: 630 MDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSI 689
D +D + F E + G+ VV+L P G+N+ V N++ Y++ ++ + S+
Sbjct: 3017 NDI----TDIITEEFAVEKDVFGEKTVVDLIPNGRNIPVTELNKQNYVNRMVDYKLRESV 3072
Query: 690 SEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKE 749
+Q+ GF+DI+ S Q F Q LEL ++ G + I ++DWK +TEY+GY
Sbjct: 3073 KDQLKSLLDGFSDIIPSHLIQIFNEQELEL-----LISGLPE-IDIDDWKNNTEYHGYNV 3126
Query: 750 TDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGLA-----SRLYIYKSPEPED 804
+ Q+ WFW V E+R LL F T +P+ GF L R I+KS +
Sbjct: 3127 SSPQVQWFWRAVRSFDEEERAKLLQFATGTSKVPLNGFKELEGMSGFQRFNIHKSYGSLN 3186
Query: 805 RLPSSHTCFYRMCFPAYSSMTVMQERL 831
RLP SHTCF ++ P Y + ++ L
Sbjct: 3187 RLPQSHTCFNQLDLPEYDTYEQLRSML 3213
>ITCH_MOUSE (Q8C863) Itchy E3 ubiquitin protein ligase (EC 6.3.2.-)
Length = 864
Score = 182 bits (462), Expect = 3e-45
Identities = 116/360 (32%), Positives = 180/360 (49%), Gaps = 14/360 (3%)
Query: 486 EMLIDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATG-PGVLREWFLLVCQALFNPQ 544
++ + R L +SF+ I L L++ F EE GV REWF L+ + NP
Sbjct: 510 KITVTRKTLFEDSFQQIMSFSPQDLRRRLWVIFPGEEGLDYGGVAREWFFLLSHEVLNPM 569
Query: 545 NALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLA 604
LF D N AS ++P HL+YF F GR IA+AL H + F+ ++
Sbjct: 570 YCLFEYAGKDNYCLQINPASYINPDHLKYFRFIGRFIAMALFHGKFIDTGFSLPFYKRIL 629
Query: 605 GKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGK 664
K + L+D++ DP Y+S ++ + + I+ L + F + E LG+ K +L P G
Sbjct: 630 NKPVGLKDLESIDPEFYNS---LIWVKENNIEECGLEMYFSVDKEILGEIKSHDLKPNGG 686
Query: 665 NLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDW 724
N++V +N+E+YI ++ + R + EQ F +GF +IL Q + Q + ++L+
Sbjct: 687 NILVTEENKEEYIRMVAEWRLSRGVEEQTQAFFEGFNEIL-----PQQYLQYFDAKELEV 741
Query: 725 MLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPV 784
+L G ++ I + DW+ H Y Y T QI WFW+ V + E+R LL F T LPV
Sbjct: 742 LLCGMQE-IDLNDWQRHAIYRHYTRTSKQIMWFWQFVKEIDNEKRMRLLQFVTGTCRLPV 800
Query: 785 EGFCGL----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIG 840
GF L + + + E+ LP SHTCF R+ P Y S ++E+L +E G
Sbjct: 801 GGFADLMGSNGPQKFCIEKVGKENWLPRSHTCFNRLDLPPYKSYEQLKEKLLFAIEETEG 860
>ITCH_HUMAN (Q96J02) Itchy homolog E3 ubiquitin protein ligase (EC
6.3.2.-) (Itch) (Atrophin-1-interacting protein 4)
(AIP4) (NFE2-associated polypeptide 1) (NAPP1)
Length = 903
Score = 182 bits (461), Expect = 4e-45
Identities = 117/360 (32%), Positives = 179/360 (49%), Gaps = 14/360 (3%)
Query: 486 EMLIDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATG-PGVLREWFLLVCQALFNPQ 544
++ + R L +SF+ I L L++ F EE GV REWF L+ + NP
Sbjct: 549 KITVTRKTLFEDSFQQIMSFSPQDLRRRLWVIFPGEEGLDYGGVAREWFFLLSHEVLNPM 608
Query: 545 NALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLA 604
LF D N AS ++P HL+YF F GR IA+AL H + F+ ++
Sbjct: 609 YCLFEYAGKDNYCLQINPASYINPDHLKYFRFIGRFIAMALFHGKFIDTGFSLPFYKRIL 668
Query: 605 GKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGK 664
K + L+D++ DP Y+S + E + + D L + F + E LG+ K +L P G
Sbjct: 669 NKPVGLKDLESIDPEFYNSLIWVKENNIEECD---LEMYFSVDKEILGEIKSHDLKPNGG 725
Query: 665 NLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDW 724
N++V +N+E+YI ++ + R + EQ F +GF +IL Q + Q + ++L+
Sbjct: 726 NILVTEENKEEYIRMVAEWRLSRGVEEQTQAFFEGFNEIL-----PQQYLQYFDAKELEV 780
Query: 725 MLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPV 784
+L G ++ I + DW+ H Y Y T QI WFW+ V + E+R LL F T LPV
Sbjct: 781 LLCGMQE-IDLNDWQRHAIYRHYARTSKQIMWFWQFVKEIDNEKRMRLLQFVTGTCRLPV 839
Query: 785 EGFCGL----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIG 840
GF L + + + E+ LP SHTCF R+ P Y S ++E+L +E G
Sbjct: 840 GGFADLMGSNGPQKFCIEKVGKENWLPRSHTCFNRLDLPPYKSYEQLKEKLLFAIEETEG 899
>NED4_MOUSE (P46935) E3 ubiquitin-protein ligase Nedd-4 (EC 6.3.2.-)
Length = 887
Score = 179 bits (454), Expect = 3e-44
Identities = 123/369 (33%), Positives = 185/369 (49%), Gaps = 19/369 (5%)
Query: 480 DYEELHEMLIDRSQLLSESFEYIARADTASL-HAGLFMEFKNEEATG-PGVLREWFLLVC 537
D EM + R+ +L +S+ I A L A L++EF E+ GV REWF L+
Sbjct: 525 DIPNKFEMKLRRANILEDSYRRIMGVKRADLLKARLWIEFDGEKGLDYGGVAREWFFLIS 584
Query: 538 QALFNPQNALFVACPNDRRRFYPNHASKV-HPLHLEYFGFSGRVIALALMHKVQVGIVLD 596
+ +FNP LF D N S + + HL YF F GRV +A+ H +
Sbjct: 585 KEMFNPYYGLFEYSATDNYTLQINPNSGLCNEDHLSYFKFIGRVAGMAVYHGKLLDGFFI 644
Query: 597 RVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKV 656
R F+ + K ITL D++ D YSS + ILE D +D L F+ + E G T
Sbjct: 645 RPFYKMMLQKLITLHDMESVDSEYYSSLRWILENDPTELD-----LRFIIDEELFGQTHQ 699
Query: 657 VELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQS 716
EL GG ++V +KN+++YI L+I+ RFV I +Q++ F +GF +++ + F
Sbjct: 700 HELKTGGSEIVVTNKNKKEYIYLVIQWRFVNRIQKQMAAFKEGFFELIPQDLIKIFDENE 759
Query: 717 LELEDLDWMLHGSEDTISVEDWKAHTEY-NGYKETDIQITWFWEIVGRMTAEQRKVLLFF 775
LEL ++ G D + V DW+ HT+Y NGY I WFW+ V M +E+R LL F
Sbjct: 760 LEL-----LMCGLGD-VDVNDWREHTKYKNGYSMNHQVIHWFWKAVWMMDSEKRIRLLQF 813
Query: 776 WTSVKYLPVEGFCGL----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERL 831
T +P+ GF L + + + D+LP +HTCF R+ P Y S + ++L
Sbjct: 814 VTGTSRVPMNGFAELYGSNGPQSFTVEQWGTPDKLPRAHTCFNRLDLPPYESFDELWDKL 873
Query: 832 EVITQEHIG 840
++ + G
Sbjct: 874 QMAIENTQG 882
>WWP2_MOUSE (Q9DBH0) Nedd-4-like E3 ubiquitin-protein ligase WWP2
(EC 6.3.2.-) (WW domain-containing protein 2)
Length = 870
Score = 178 bits (451), Expect = 6e-44
Identities = 118/357 (33%), Positives = 173/357 (48%), Gaps = 14/357 (3%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATG-PGVLREWFLLVCQALFNPQNAL 547
+ R L +SF+ I L L++ + EE G+ REWF L+ + NP L
Sbjct: 519 VSRQTLFEDSFQQIMNMKPYDLRRRLYIIMRGEEGLDYGGIAREWFFLLSHEVLNPMYCL 578
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F + N AS ++P HL YF F GR IA+AL H + F+ ++ K
Sbjct: 579 FEYAGKNNYCLQINPASSINPDHLTYFRFIGRFIAMALYHGKFIDTGFTLPFYKRMLNKR 638
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
TL+D++ DP Y+S I E + ++ L L F++++E LG EL GG+N+
Sbjct: 639 PTLKDLESIDPEFYNSIVWIKENN---LEECGLELFFIQDMEILGKVTTHELKEGGENIR 695
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+E+YI LL RF + EQ F GF ++ + F + LEL ML
Sbjct: 696 VTEENKEEYIMLLTDWRFTRGVEEQTKAFLDGFNEVAPLEWLRYFDEKELEL-----MLC 750
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
G ++ I + DW+ + Y Y ++ QI WFW++V M E+R LL F T LPV GF
Sbjct: 751 GMQE-IDMSDWQKNAIYRHYTKSSKQIQWFWQVVKEMDNEKRIRLLQFVTGTCRLPVGGF 809
Query: 788 CGL----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIG 840
L + + E LP SHTCF R+ P Y S ++E+L +E G
Sbjct: 810 AELIGSNGPQKFCIDRVGKETWLPRSHTCFNRLDLPPYKSYEQLKEKLLYAIEETEG 866
>TOM1_ASHGO (Q756G2) Probable E3 ubiquitin protein ligase TOM1 (EC
6.3.2.-)
Length = 3258
Score = 178 bits (451), Expect = 6e-44
Identities = 117/395 (29%), Positives = 196/395 (49%), Gaps = 19/395 (4%)
Query: 454 ILEHKCVTNFESRRHLAMMMFPEVKEDYEELHEMLIDRSQLLSESFEYIARADTASLHAG 513
++++ + +F+++R+ + D +L + + R + +S+ + +
Sbjct: 2871 LVKNPKILDFDNKRYYFTAQLRAITHDRPKL-SISVRREHVFLDSYRSLFFKSNEDIKIS 2929
Query: 514 -LFMEFKNEEATGPG-VLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHL 571
L + FK E G + REW+ ++ + +FNP ALF+ +D+ F PN S ++P HL
Sbjct: 2930 KLEISFKGEAGVDAGGITREWYQVLSRQMFNPDYALFIPVASDKTTFRPNRTSGINPEHL 2989
Query: 572 EYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMD 631
+F F G +I A+ + + R + + GK + L+D++ D Y S ILE D
Sbjct: 2990 SFFKFIGMIIGKAISDQCFLDCHFSREVYKNILGKPVALKDMESLDLDYYKSLIWILEND 3049
Query: 632 SDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISE 691
+D + TF E ++ G+ KV+EL G ++ V +N+ Y+ +++ + TS+ +
Sbjct: 3050 I----TDIIEETFSVETDDYGEHKVIELIENGAHVAVTEQNKHDYVKKIVEYKLQTSVKD 3105
Query: 692 QVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETD 751
Q+ +F +GF I+ F Q LEL ++ G D I V+DWK +T Y Y T
Sbjct: 3106 QMENFLQGFYAIIPKDLISIFDEQELEL-----LVSGLPD-IDVDDWKNNTIYVNYTPTC 3159
Query: 752 IQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGLA-----SRLYIYKSPEPEDRL 806
QI +FW V E+R LL F T +P+ GF L+ S+ I++ DRL
Sbjct: 3160 KQINYFWRAVRSFDKEERAKLLQFVTGTSKVPLNGFKELSGVNGISKFSIHRDYGSIDRL 3219
Query: 807 PSSHTCFYRMCFPAYSSM-TVMQERLEVITQEHIG 840
PSSHTCF ++ PAY S T+ L I + H G
Sbjct: 3220 PSSHTCFNQLDLPAYDSYETLRGSLLLAINEGHEG 3254
>WWP2_HUMAN (O00308) Nedd-4-like E3 ubiquitin-protein ligase WWP2
(EC 6.3.2.-) (WW domain-containing protein 2) (Atropin-1
interacting protein 2) (AIP2)
Length = 870
Score = 177 bits (449), Expect = 1e-43
Identities = 118/357 (33%), Positives = 172/357 (48%), Gaps = 14/357 (3%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATG-PGVLREWFLLVCQALFNPQNAL 547
+ R L +SF+ I L L++ + EE G+ REWF L+ + NP L
Sbjct: 519 VSRQTLFEDSFQQIMNMKPYDLRRRLYIIMRGEEGLDYGGIAREWFFLLSHEVLNPMYCL 578
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F + N AS ++P HL YF F GR IA+AL H + F+ ++ K
Sbjct: 579 FEYAGKNNYCLQINPASSINPDHLTYFRFIGRFIAMALYHGKFIDTGFTLPFYKRMLNKR 638
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCPGGKNLI 667
TL+D++ DP Y+S I E + ++ L L F++++E LG EL GG+++
Sbjct: 639 PTLKDLESIDPEFYNSIVWIKENN---LEECGLELYFIQDMEILGKVTTHELKEGGESIR 695
Query: 668 VNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLH 727
V +N+E+YI LL RF + EQ F GF ++ + F + LEL ML
Sbjct: 696 VTEENKEEYIMLLTDWRFTRGVEEQTKAFLDGFNEVAPLEWLRYFDEKELEL-----MLC 750
Query: 728 GSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGF 787
G ++ I + DW+ T Y Y + QI WFW++V M E+R LL F T LPV GF
Sbjct: 751 GMQE-IDMSDWQKSTIYRHYTKNSKQIQWFWQVVKEMDNEKRIRLLQFVTGTCRLPVGGF 809
Query: 788 CGL----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIG 840
L + + E LP SHTCF R+ P Y S ++E+L +E G
Sbjct: 810 AELIGSNGPQKFCIDKVGKETWLPRSHTCFNRLDLPPYKSYEQLREKLLYAIEETEG 866
>NED4_HUMAN (P46934) E3 ubiquitin-protein ligase Nedd-4 (EC 6.3.2.-)
Length = 1000
Score = 176 bits (446), Expect = 2e-43
Identities = 121/371 (32%), Positives = 186/371 (49%), Gaps = 19/371 (5%)
Query: 478 KEDYEELHEMLIDRSQLLSESFEYIARADTAS-LHAGLFMEFKNEEATG-PGVLREWFLL 535
+ D EM + R+ +L +S+ I A L A L++EF E+ GV REWF L
Sbjct: 636 QNDIPNKFEMKLRRATVLEDSYRRIMGVKRADFLKARLWIEFDGEKGLDYGGVAREWFFL 695
Query: 536 VCQALFNPQNALFVACPNDRRRFYPNHASKV-HPLHLEYFGFSGRVIALALMHKVQVGIV 594
+ + +FNP LF D N S + + HL YF F GRV +A+ H +
Sbjct: 696 ISKEMFNPYYGLFEYSATDNYTLQINPNSGLCNEDHLSYFKFIGRVAGMAVYHGKLLDGF 755
Query: 595 LDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDT 654
R F+ + K ITL D++ D Y+S + ILE D +D L F+ + E G T
Sbjct: 756 FIRPFYKMMLHKPITLHDMESVDSEYYNSLRWILENDPTELD-----LRFIIDEELFGQT 810
Query: 655 KVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFF 714
EL GG ++V +KN+++YI L+I+ RFV I +Q++ F +GF +++ + F
Sbjct: 811 HQHELKNGGSEIVVTNKNKKEYIYLVIQWRFVNRIQKQMAAFKEGFFELIPQDLIKIFDE 870
Query: 715 QSLELEDLDWMLHGSEDTISVEDWKAHTEY-NGYKETDIQITWFWEIVGRMTAEQRKVLL 773
LEL ++ G D + V DW+ HT+Y NGY I WFW+ V M +E+R LL
Sbjct: 871 NELEL-----LMCGLGD-VDVNDWREHTKYKNGYSANHQVIQWFWKAVLMMDSEKRIRLL 924
Query: 774 FFWTSVKYLPVEGFCGL----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQE 829
F T +P+ GF L + + + ++LP +HTCF R+ P Y S + +
Sbjct: 925 QFVTGTSRVPMNGFAELYGSNGPQSFTVEQWGTPEKLPRAHTCFNRLDLPPYESFEELWD 984
Query: 830 RLEVITQEHIG 840
+L++ + G
Sbjct: 985 KLQMAIENTQG 995
>NED4_RAT (Q62940) E3 ubiquitin-protein ligase Nedd-4 (EC 6.3.2.-)
Length = 887
Score = 175 bits (444), Expect = 4e-43
Identities = 122/375 (32%), Positives = 187/375 (49%), Gaps = 30/375 (8%)
Query: 480 DYEELHEMLIDRSQLLSESFEYIARADTAS-LHAGLFMEFKNEEATG-PGVLREWFLLVC 537
D EM + R+ +L +S+ I A L A L++EF E+ GV REWF L+
Sbjct: 524 DIPNKFEMKLRRANILEDSYRRIMGVKRADFLKARLWIEFDGEKGLDYGGVAREWFFLIS 583
Query: 538 QALFNPQNALFVACPNDRRRFYPNHASKVHPL-------HLEYFGFSGRVIALALMHKVQ 590
+ +FNP LF + N+ +++P HL YF F GRV +A+ H
Sbjct: 584 KEMFNPYYGLFEYSATE-----DNYTLQINPNSGLCNEDHLSYFKFIGRVAGMAVYHGKL 638
Query: 591 VGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEE 650
+ R F+ + K ITL D++ D YSS + ILE D +D L F+ + E
Sbjct: 639 LDGFFIRPFYKMMLQKLITLHDMESVDSEYYSSLRWILENDPTELD-----LRFIIDEEL 693
Query: 651 LGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQ 710
G T EL GG ++V +KN+++YI L+I+ RFV I +Q++ F +GF +++ +
Sbjct: 694 FGQTHQHELKTGGSEVVVTNKNKKEYIYLVIQWRFVNRIQKQMAAFKEGFFELIPQDLIK 753
Query: 711 QFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEY-NGYKETDIQITWFWEIVGRMTAEQR 769
F LEL ++ G D + V DW+ HT+Y NGY I WFW+ V M +E+R
Sbjct: 754 IFDENELEL-----LMCGLGD-VDVNDWREHTKYKNGYSLNHQVIHWFWKAVLMMDSEKR 807
Query: 770 KVLLFFWTSVKYLPVEGFCGL----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMT 825
LL F T +P+ GF L + + + D+LP +HTCF R+ P Y S
Sbjct: 808 IRLLQFVTGTSRVPMNGFAELYGSNGPQSFTVEQWGTPDKLPRAHTCFNRLDLPPYESFD 867
Query: 826 VMQERLEVITQEHIG 840
+ ++L++ + G
Sbjct: 868 ELWDKLQMAIENTQG 882
>SUF1_XENLA (Q9PUN2) Smad ubiquitination regulatory factor 1 (EC
6.3.2.-) (Ubiquitin--protein ligase SMURF1)
(Smad-specific E3 ubiquitin ligase) (xSMURF1)
Length = 731
Score = 173 bits (439), Expect = 2e-42
Identities = 122/394 (30%), Positives = 177/394 (43%), Gaps = 24/394 (6%)
Query: 456 EHKCVTNFESRRHLAMMMFPEVKEDYEELHEMLIDRSQLLSESFEYIARADTASLHAGLF 515
E V + RH ++ P+ E + R ++ ES+ I + L L
Sbjct: 349 ERDLVQKLKVLRHELSLLQPQAGHCRVE-----VSREEIFEESYRQIMKMRPKDLKKRLM 403
Query: 516 MEFKNEEATG-PGVLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYF 574
++F+ EE GV REW L+C + NP LF ++ N S ++P HL YF
Sbjct: 404 VKFRGEEGLDYGGVAREWLYLLCHEMLNPYYGLFQYSTDNIYTLQINPDSSINPDHLSYF 463
Query: 575 GFSGRVIALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDF 634
F GR++ LA+ H + F+ QL GK I L D++ DP L+ S ILE D
Sbjct: 464 HFVGRIMGLAVFHGHYINGGFTVPFYKQLLGKPIQLSDLESVDPELHKSLVWILENDI-- 521
Query: 635 IDSDALGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVS 694
+ L TF E G EL P GKNL V +N+++Y+ L + RF+ I Q
Sbjct: 522 --TSVLDHTFCVEHNAFGRLLQHELKPNGKNLQVTEENKKEYVRLYVNWRFMRGIEAQFL 579
Query: 695 HFSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQI 754
KGF +++ + F + LEL + G D I + DWKA+T +
Sbjct: 580 ALQKGFNELIPQHLLKPFEQKELEL------IIGGLDKIDISDWKANTRLKHCLANSNIV 633
Query: 755 TWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGL-------ASRLY-IYKSPEPEDRL 806
WFW+ V E+R LL F T +P++GF L RL+ I+ D L
Sbjct: 634 QWFWQAVESFDEERRARLLQFVTGSTRVPLQGFKALQGSTGAAGPRLFTIHLIDANTDNL 693
Query: 807 PSSHTCFYRMCFPAYSSMTVMQERLEVITQEHIG 840
P +HTCF R+ P Y S + E+L +E G
Sbjct: 694 PKAHTCFNRIDIPPYESYEKLYEKLLTAVEETSG 727
>UPL1_ARATH (Q8GY23) E3 ubiquitin protein ligase UPL1 (EC 6.3.2.-)
(Ubiquitin-protein ligase 1)
Length = 3684
Score = 173 bits (438), Expect = 2e-42
Identities = 117/362 (32%), Positives = 177/362 (48%), Gaps = 24/362 (6%)
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPGVL-REWFLLVCQALFNPQNAL 547
+ R+ +L +S+ + L L ++F+ EE G L REW+ L+ + +F+ L
Sbjct: 3323 VRRAYVLEDSYNQLRMRSPQDLKGRLNVQFQGEEGIDAGGLTREWYQLLSRVIFDKGALL 3382
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F ND F PN S HL YF F GR++A AL + + R F+ + G
Sbjct: 3383 FTTVGNDAT-FQPNPNSVYQTEHLSYFKFVGRMVAKALFDGQLLDVYFTRSFYKHILGVK 3441
Query: 608 ITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEE-----LGDTKVV--ELC 660
+T D++ DP Y + K +LE D SD L LTF + +E T+V EL
Sbjct: 3442 VTYHDIEAVDPDYYKNLKWLLENDV----SDILDLTFSMDADEEKHILYEKTEVTDYELK 3497
Query: 661 PGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELE 720
PGG+N+ V + + +Y+DL+ +I Q++ F +GF +++ F + LEL
Sbjct: 3498 PGGRNIRVTEETKHEYVDLVAGHILTNAIRPQINAFLEGFNELIPRELVSIFNDKELEL- 3556
Query: 721 DLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVK 780
++ G + I +D KA+TEY Y I WFWE+V + E L F T
Sbjct: 3557 ----LISGLPE-IDFDDLKANTEYTSYTAGSPVIHWFWEVVKAFSKEDMARFLQFVTGTS 3611
Query: 781 YLPVEGFCGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVIT 835
+P+EGF L RL I+K+ +RLPS+HTCF ++ P Y S +QERL +
Sbjct: 3612 KVPLEGFKALQGISGPQRLQIHKAYGAPERLPSAHTCFNQLDLPEYQSKEQLQERLLLAI 3671
Query: 836 QE 837
E
Sbjct: 3672 HE 3673
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.137 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 98,906,437
Number of Sequences: 164201
Number of extensions: 4185928
Number of successful extensions: 9661
Number of sequences better than 10.0: 133
Number of HSP's better than 10.0 without gapping: 116
Number of HSP's successfully gapped in prelim test: 17
Number of HSP's that attempted gapping in prelim test: 9396
Number of HSP's gapped (non-prelim): 151
length of query: 846
length of database: 59,974,054
effective HSP length: 119
effective length of query: 727
effective length of database: 40,434,135
effective search space: 29395616145
effective search space used: 29395616145
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0288.8


