

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0284.4
(247 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
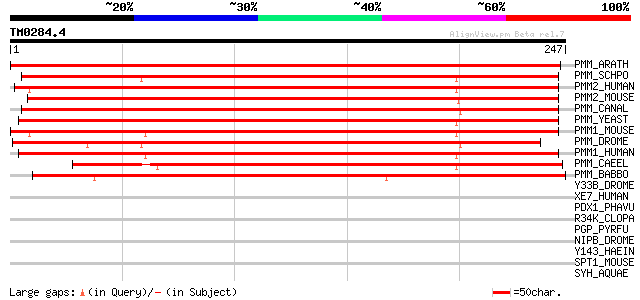
PMM_ARATH (O80840) Probable phosphomannomutase (EC 5.4.2.8) (PMM) 419 e-117
PMM_SCHPO (Q9UTJ2) Phosphomannomutase (EC 5.4.2.8) (PMM) 307 2e-83
PMM2_HUMAN (O15305) Phosphomannomutase 2 (EC 5.4.2.8) (PMM 2) 299 5e-81
PMM2_MOUSE (Q9Z2M7) Phosphomannomutase 2 (EC 5.4.2.8) (PMM 2) 297 2e-80
PMM_CANAL (P31353) Phosphomannomutase (EC 5.4.2.8) (PMM) 286 3e-77
PMM_YEAST (P07283) Phosphomannomutase (EC 5.4.2.8) (PMM) 281 8e-76
PMM1_MOUSE (O35621) Phosphomannomutase 1 (EC 5.4.2.8) (PMM 1) 271 1e-72
PMM_DROME (Q9VTZ6) Probable phosphomannomutase (EC 5.4.2.8) (PMM) 270 3e-72
PMM1_HUMAN (Q92871) Phosphomannomutase 1 (EC 5.4.2.8) (PMM 1) (P... 270 3e-72
PMM_CAEEL (Q9XUE6) Probable phosphomannomutase (EC 5.4.2.8) (PMM) 235 8e-62
PMM_BABBO (O43976) Phosphomannomutase (EC 5.4.2.8) 201 1e-51
Y33B_DROME (Q9U1M2) Hypothetical protein CG32795 in chromosome 1 36 0.075
XE7_HUMAN (Q02040) B-lymphocyte antigen precursor (B-lymphocyte ... 33 0.64
PDX1_PHAVU (Q9FT25) Probable pyridoxin biosynthesis protein PDX1... 32 1.4
R34K_CLOPA (P23160) 34.2 kDa protein in rubredoxin operon (EC 1.... 31 2.4
PGP_PYRFU (Q8U111) Phosphoglycolate phosphatase (EC 3.1.3.18) (PGP) 31 3.2
NIPB_DROME (Q7PLI2) Nipped-B protein (SCC2 homolog) 31 3.2
Y143_HAEIN (P44540) Putative HTH-type transcriptional regulator ... 30 4.1
SPT1_MOUSE (Q9R097) Kunitz-type protease inhibitor 1 precursor (... 30 4.1
SYH_AQUAE (O66522) Histidyl-tRNA synthetase (EC 6.1.1.21) (Histi... 30 5.4
>PMM_ARATH (O80840) Probable phosphomannomutase (EC 5.4.2.8) (PMM)
Length = 246
Score = 419 bits (1078), Expect = e-117
Identities = 201/245 (82%), Positives = 225/245 (91%)
Query: 1 MAVAKPGVIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGST 60
MA PGVIALFDVDGTLTAPRK A E+L F++ELRKVVT+GVVGGSDL KISEQLG T
Sbjct: 1 MAAKIPGVIALFDVDGTLTAPRKEATPELLDFIRELRKVVTIGVVGGSDLSKISEQLGKT 60
Query: 61 VTHDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTF 120
VT+DYDY FSENGLVAHK GK IG +SLK LGD+KLKE INFTLHYIADLDIPIKRGTF
Sbjct: 61 VTNDYDYCFSENGLVAHKDGKSIGIQSLKLHLGDDKLKELINFTLHYIADLDIPIKRGTF 120
Query: 121 IEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQIS 180
IEFR+GMLNVSPIGRNCSQEERDEFE+YDKVQNIRPKMV+ LRE+FAHLNLTFSIGGQIS
Sbjct: 121 IEFRNGMLNVSPIGRNCSQEERDEFERYDKVQNIRPKMVAELRERFAHLNLTFSIGGQIS 180
Query: 181 FDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQC 240
FDVFP+GWDKTYCL+YL+ F+EIHFFGDKTY+GGND+EIYES +TIGH+VTSP+DT+ +C
Sbjct: 181 FDVFPKGWDKTYCLQYLEDFSEIHFFGDKTYEGGNDYEIYESPKTIGHSVTSPDDTVAKC 240
Query: 241 KSLFL 245
K+LF+
Sbjct: 241 KALFM 245
>PMM_SCHPO (Q9UTJ2) Phosphomannomutase (EC 5.4.2.8) (PMM)
Length = 257
Score = 307 bits (786), Expect = 2e-83
Identities = 148/244 (60%), Positives = 187/244 (75%), Gaps = 5/244 (2%)
Query: 6 PGVIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQL---GSTVT 62
P + LFDVDGTLT R + EML +Q LRKVV +G VGGSDL K EQL G V
Sbjct: 12 PKTLVLFDVDGTLTPARLSVSPEMLETLQNLRKVVAIGFVGGSDLSKQQEQLSVNGENVI 71
Query: 63 HDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFIE 122
+DY F+ENGL A++ G+ + ++S +LG+EK ++ +NF LHYIADLDIP+KRGTFIE
Sbjct: 72 DSFDYAFAENGLTAYRYGQQLASQSFIAWLGEEKYQKLVNFCLHYIADLDIPVKRGTFIE 131
Query: 123 FRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFD 182
FR+GM+N+SP+GRN + EER+EFE++DK + IR MV VLREKF LTFSIGGQISFD
Sbjct: 132 FRNGMINISPVGRNANTEERNEFERFDKGRKIRATMVDVLREKFKDYGLTFSIGGQISFD 191
Query: 183 VFPQGWDKTYCLRYL--DGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQC 240
VFP GWDKTYCL+++ +GF+ IHFFGDKTYKGGND+EI+ RTIGH+VT+P+DTI +
Sbjct: 192 VFPAGWDKTYCLQHVEKEGFDTIHFFGDKTYKGGNDYEIFVDPRTIGHSVTNPDDTIAEL 251
Query: 241 KSLF 244
K +F
Sbjct: 252 KKIF 255
>PMM2_HUMAN (O15305) Phosphomannomutase 2 (EC 5.4.2.8) (PMM 2)
Length = 246
Score = 299 bits (765), Expect = 5e-81
Identities = 147/245 (60%), Positives = 189/245 (77%), Gaps = 3/245 (1%)
Query: 3 VAKPG-VIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGSTV 61
+A PG + LFDVDGTLTAPR+ EM F+Q+LR+ + +GVVGGSD K+ EQLG+ V
Sbjct: 1 MAAPGPALCLFDVDGTLTAPRQKITKEMDDFLQKLRQKIKIGVVGGSDFEKVQEQLGNDV 60
Query: 62 THDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFI 121
YDYVF ENGLVA+K GKL+ +++++ LG+ +++ IN+ L YIA + +P KRGTFI
Sbjct: 61 VEKYDYVFPENGLVAYKDGKLLCRQNIQSHLGEALIQDLINYCLSYIAKIKLPKKRGTFI 120
Query: 122 EFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISF 181
EFR+GMLNVSPIGR+CSQEER EF + DK +NIR K V+ LR++FA LTFSIGGQISF
Sbjct: 121 EFRNGMLNVSPIGRSCSQEERIEFYELDKKENIRQKFVADLRKEFAGKGLTFSIGGQISF 180
Query: 182 DVFPQGWDKTYCLRYL--DGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQ 239
DVFP GWDK YCLR++ DG+ I+FFGDKT GGNDHEI+ RT+G++VT+PEDT +
Sbjct: 181 DVFPDGWDKRYCLRHVENDGYKTIYFFGDKTMPGGNDHEIFTDPRTMGYSVTAPEDTRRI 240
Query: 240 CKSLF 244
C+ LF
Sbjct: 241 CELLF 245
>PMM2_MOUSE (Q9Z2M7) Phosphomannomutase 2 (EC 5.4.2.8) (PMM 2)
Length = 242
Score = 297 bits (760), Expect = 2e-80
Identities = 143/238 (60%), Positives = 186/238 (78%), Gaps = 2/238 (0%)
Query: 9 IALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGSTVTHDYDYV 68
+ LFD+DGTLTAPR+ EM GF+Q+LR+ +GVVGGSD K+ EQLG+ V YDYV
Sbjct: 4 LCLFDMDGTLTAPRQKITEEMDGFLQKLRQKTKIGVVGGSDFEKLQEQLGNDVVEKYDYV 63
Query: 69 FSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFIEFRSGML 128
F ENGLVA+K GKL+ ++++ LG++ +++ IN+ L YIA++ +P KRGTFIEFR+GML
Sbjct: 64 FPENGLVAYKDGKLLCKQNIQGHLGEDVIQDLINYCLSYIANIKLPKKRGTFIEFRNGML 123
Query: 129 NVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFDVFPQGW 188
NVSPIGR+CSQEER EF + DK ++IR K V+ LR++FA LTFSIGGQIS DVFP+GW
Sbjct: 124 NVSPIGRSCSQEERIEFYELDKKEHIRQKFVADLRKEFAGKGLTFSIGGQISIDVFPEGW 183
Query: 189 DKTYCLRYLD--GFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQCKSLF 244
DK YCLR+L+ G+ I+FFGDKT GGNDHEI+ RT+G+TVT+PEDT + C+ LF
Sbjct: 184 DKRYCLRHLEHAGYKTIYFFGDKTMPGGNDHEIFTDPRTVGYTVTAPEDTRRICEGLF 241
>PMM_CANAL (P31353) Phosphomannomutase (EC 5.4.2.8) (PMM)
Length = 252
Score = 286 bits (732), Expect = 3e-77
Identities = 135/241 (56%), Positives = 178/241 (73%), Gaps = 2/241 (0%)
Query: 6 PGVIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGSTVTHDY 65
P + LFDVDGTLT R + EM +++LR+ V +G VGGSDL K EQLG V +D+
Sbjct: 9 PKTLVLFDVDGTLTPARLTISEEMKKTLEKLREKVVIGFVGGSDLSKQVEQLGPNVLNDF 68
Query: 66 DYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFIEFRS 125
DY FSENGL A+K GK + ++S ++G+EK + + F L Y++D+D+PI+RGTFIEFR+
Sbjct: 69 DYCFSENGLTAYKLGKELASQSFINWIGNEKYNKLVKFILRYLSDIDLPIRRGTFIEFRN 128
Query: 126 GMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFDVFP 185
GM+NVSPIGRN S +ER+++EK+DK +IR MV L+++F LT+SIGGQISFDVFP
Sbjct: 129 GMINVSPIGRNASTQERNDYEKFDKQHHIRETMVEALKKEFPDFGLTYSIGGQISFDVFP 188
Query: 186 QGWDKTYCLRYLDG--FNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQCKSL 243
GWDKTYCL++++ F IHFFGDK+YKGGND+EIY RTIGH V SP+DTI+
Sbjct: 189 TGWDKTYCLQHVEDEHFENIHFFGDKSYKGGNDYEIYNDPRTIGHAVNSPDDTIRILNET 248
Query: 244 F 244
F
Sbjct: 249 F 249
>PMM_YEAST (P07283) Phosphomannomutase (EC 5.4.2.8) (PMM)
Length = 254
Score = 281 bits (720), Expect = 8e-76
Identities = 137/242 (56%), Positives = 175/242 (71%), Gaps = 2/242 (0%)
Query: 5 KPGVIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGSTVTHD 64
KP + LFDVDGTLT R + E+ + +LR +G VGGSDL K EQLG V +
Sbjct: 11 KPETLVLFDVDGTLTPARLTVSEEVRKTLAKLRNKCCIGFVGGSDLSKQLEQLGPNVLDE 70
Query: 65 YDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFIEFR 124
+DY FSENGL A++ GK + ++S +LG+EK + F L Y++++D+P +RGTF+EFR
Sbjct: 71 FDYSFSENGLTAYRLGKELASQSFINWLGEEKYNKLAVFILRYLSEIDLPKRRGTFLEFR 130
Query: 125 SGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFDVF 184
+GM+NVSPIGRN S EER+EFE+YDK IR K V L+++F LTFSIGGQISFDVF
Sbjct: 131 NGMINVSPIGRNASTEERNEFERYDKEHQIRAKFVEALKKEFPDYGLTFSIGGQISFDVF 190
Query: 185 PQGWDKTYCLRYL--DGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQCKS 242
P GWDKTYCL+++ DGF EIHFFGDKT GGND+EI+ ERTIGH+V SP+DT+K
Sbjct: 191 PAGWDKTYCLQHVEKDGFKEIHFFGDKTMVGGNDYEIFVDERTIGHSVQSPDDTVKILTE 250
Query: 243 LF 244
LF
Sbjct: 251 LF 252
>PMM1_MOUSE (O35621) Phosphomannomutase 1 (EC 5.4.2.8) (PMM 1)
Length = 262
Score = 271 bits (692), Expect = 1e-72
Identities = 139/254 (54%), Positives = 180/254 (70%), Gaps = 10/254 (3%)
Query: 1 MAVAKPG------VIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKIS 54
MAVA G ++ LFDVDGTLT R+ + E+ F+Q+LR V +GVVGGSD KI+
Sbjct: 1 MAVAVEGARRKERILCLFDVDGTLTPARQKIDPEVSAFLQKLRSRVQIGVVGGSDYSKIA 60
Query: 55 EQLGS--TVTHDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLD 112
EQLG V +DYVF+ENG V +K G+L+ ++++ LG+E L++ INF L Y+A L
Sbjct: 61 EQLGEGDEVIEKFDYVFAENGTVQYKHGRLLSKQTIQNHLGEELLQDLINFCLSYMALLR 120
Query: 113 IPIKRGTFIEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLT 172
+P KRGTFIEFR+GMLNVSPIGR+C+ EER EF + DK + IR K V L+ +FA L
Sbjct: 121 LPKKRGTFIEFRNGMLNVSPIGRSCTLEERIEFSELDKKEKIREKFVEALKTEFAGKGLR 180
Query: 173 FSIGGQISFDVFPQGWDKTYCLRYL--DGFNEIHFFGDKTYKGGNDHEIYESERTIGHTV 230
FS GG ISFDVFP+GWDK YCL L D F+ IHFFG++T GGND EIY RT+GH+V
Sbjct: 181 FSRGGMISFDVFPEGWDKRYCLDSLDEDSFDIIHFFGNETSPGGNDFEIYADPRTVGHSV 240
Query: 231 TSPEDTIKQCKSLF 244
SP+DT+++C+ LF
Sbjct: 241 VSPQDTVQRCRELF 254
>PMM_DROME (Q9VTZ6) Probable phosphomannomutase (EC 5.4.2.8) (PMM)
Length = 254
Score = 270 bits (689), Expect = 3e-72
Identities = 130/240 (54%), Positives = 176/240 (73%), Gaps = 5/240 (2%)
Query: 2 AVAKPGVIALFDVDGTLTAPRKVANSEMLGFM-QELRKVVTVGVVGGSDLIKISEQL-GS 59
A+ + ++ LFDVDGTLT PR V E F ++ T+G+VGGSDL K+ EQL G
Sbjct: 5 ALKRDEILLLFDVDGTLTMPRSVVTPEFEEFFYSRVKPRATIGIVGGSDLEKMFEQLNGR 64
Query: 60 TVTHDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGT 119
+ +++D++F ENGLV + GK +G +++ LG+E +K FINF L Y+++LD+PIKRGT
Sbjct: 65 KILNEFDFIFPENGLVQIEGGKEVGKQNIIMHLGEETVKRFINFVLRYLSELDVPIKRGT 124
Query: 120 FIEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQI 179
FIEFR+GM+NV PIGR C++EER+ F +YD +R KM+ L+++FA ++LT+SIGGQI
Sbjct: 125 FIEFRNGMMNVCPIGRQCTREERNMFAEYDIEHKVREKMIKDLKQEFADVDLTYSIGGQI 184
Query: 180 SFDVFPQGWDKTYCLRYLDG---FNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDT 236
SFDVFP GWDKTYCLR+++ F EIHFFGDKT GGND+EIY RTI H V +P+DT
Sbjct: 185 SFDVFPHGWDKTYCLRHIEAHYKFKEIHFFGDKTEPGGNDYEIYSDPRTISHRVYTPKDT 244
>PMM1_HUMAN (Q92871) Phosphomannomutase 1 (EC 5.4.2.8) (PMM 1)
(PMMH-22)
Length = 262
Score = 270 bits (689), Expect = 3e-72
Identities = 133/244 (54%), Positives = 176/244 (71%), Gaps = 4/244 (1%)
Query: 5 KPGVIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGS--TVT 62
K V+ LFDVDGTLT R+ + E+ F+Q+LR V +GVVGGSD KI+EQLG V
Sbjct: 11 KERVLCLFDVDGTLTPARQKIDPEVAAFLQKLRSRVQIGVVGGSDYCKIAEQLGDGDEVI 70
Query: 63 HDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFIE 122
+DYVF+ENG V +K G+L+ ++++ LG+E L++ INF L Y+A L +P KRGTFIE
Sbjct: 71 EKFDYVFAENGTVQYKHGRLLSKQTIQNHLGEELLQDLINFCLSYMALLRLPKKRGTFIE 130
Query: 123 FRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFD 182
FR+GMLN+SPIGR+C+ EER EF + DK + IR K V L+ +FA L FS GG ISFD
Sbjct: 131 FRNGMLNISPIGRSCTLEERIEFSELDKKEKIREKFVEALKTEFAGKGLRFSRGGMISFD 190
Query: 183 VFPQGWDKTYCLRYL--DGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQC 240
VFP+GWDK YCL L D F+ IHFFG++T GGND EI+ RT+GH+V SP+DT+++C
Sbjct: 191 VFPEGWDKRYCLDSLDQDSFDTIHFFGNETSPGGNDFEIFADPRTVGHSVVSPQDTVQRC 250
Query: 241 KSLF 244
+ +F
Sbjct: 251 REIF 254
>PMM_CAEEL (Q9XUE6) Probable phosphomannomutase (EC 5.4.2.8) (PMM)
Length = 245
Score = 235 bits (599), Expect = 8e-62
Identities = 122/246 (49%), Positives = 155/246 (62%), Gaps = 31/246 (12%)
Query: 29 MLGFMQELRKVVTVGVVGGSDLIKISEQLGSTVTHD------------------------ 64
M F+ E RK V + +VGGSD KI+EQL HD
Sbjct: 1 MREFLIEARKRVPLAIVGGSDFKKITEQLAD---HDKDLCMSLLIFFFELQILISSFFPV 57
Query: 65 ---YDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFI 121
+DY FSENGL K + +S++ +GD KL+E INF L Y++D+ +P+KRG F+
Sbjct: 58 LSLFDYTFSENGLYGFKGTEPYPVQSIQKAIGDAKLQELINFALRYMSDIQLPVKRGNFV 117
Query: 122 EFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISF 181
EFR+GM+N+SPIGR+CSQEER +F ++DK IR K LREKF L F+IGGQIS
Sbjct: 118 EFRNGMINLSPIGRSCSQEERMQFVEFDKKHGIRQKFTEQLREKFGQYGLQFAIGGQISV 177
Query: 182 DVFPQGWDKTYCLRYL-DGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQC 240
DVFP GWDKT+CL+YL F+ IHFFGDKT GGNDHEI+ ERT+GHTV PEDT K
Sbjct: 178 DVFPTGWDKTFCLQYLVPDFDTIHFFGDKTAPGGNDHEIFADERTVGHTVEGPEDTRKHV 237
Query: 241 KSLFLE 246
+++ E
Sbjct: 238 ENVLKE 243
>PMM_BABBO (O43976) Phosphomannomutase (EC 5.4.2.8)
Length = 246
Score = 201 bits (511), Expect = 1e-51
Identities = 102/240 (42%), Positives = 147/240 (60%), Gaps = 3/240 (1%)
Query: 11 LFDVDGTLTAPRKVANSEMLGFMQEL-RKVVTVGVVGGSDLIKISEQLGSTVTHDYDYVF 69
+FD+DGTLT P +V N+++ ++ RK + VV GS KI QL ++DYVF
Sbjct: 7 IFDMDGTLTDPVQVINNDVKDILRRCKRKNFEIAVVSGSKYEKIKGQLNDGFIDEFDYVF 66
Query: 70 SENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFIEFRSGMLN 129
SENG + + L+ + + + + KL++ + F L YIADLDIP KRGTFIE R ++N
Sbjct: 67 SENGTQVYVKNVLVKSLDITEAIPETKLRKMVEFCLRYIADLDIPTKRGTFIEHRKSLIN 126
Query: 130 VSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKF--AHLNLTFSIGGQISFDVFPQG 187
+ P GRNCS +R F +YD + ++R K++ VL+ +F L+F GGQIS DV+P+
Sbjct: 127 ICPPGRNCSMVDRRRFVEYDSIHHVRQKLIQVLKSQFDSDDCPLSFVAGGQISIDVYPKA 186
Query: 188 WDKTYCLRYLDGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQCKSLFLES 247
W K+ L ++ + IHFFGD T +GGND EIY IGHTVT +D + Q + L +S
Sbjct: 187 WSKSIALSHIGKCDVIHFFGDNTREGGNDFEIYNHPDVIGHTVTGYKDLVNQLEELLAKS 246
>Y33B_DROME (Q9U1M2) Hypothetical protein CG32795 in chromosome 1
Length = 387
Score = 36.2 bits (82), Expect = 0.075
Identities = 24/81 (29%), Positives = 39/81 (47%), Gaps = 11/81 (13%)
Query: 85 TESLKTFLGD------EKLKEFINFTLHYIADL-----DIPIKRGTFIEFRSGMLNVSPI 133
T SL+ F G EK+ + TL A L +P K G +++ G +NVS +
Sbjct: 56 TTSLRQFKGPVPAEDKEKVDDLHKMTLKRKAQLHEIEQSLPAKSGRYLQIILGDVNVSIL 115
Query: 134 GRNCSQEERDEFEKYDKVQNI 154
RN +D++EK+ + N+
Sbjct: 116 NRNDKVRYKDDYEKFKLILNV 136
>XE7_HUMAN (Q02040) B-lymphocyte antigen precursor (B-lymphocyte
surface antigen) (721P) (Protein XE7)
Length = 695
Score = 33.1 bits (74), Expect = 0.64
Identities = 15/43 (34%), Positives = 26/43 (59%), Gaps = 1/43 (2%)
Query: 145 FEKYDKVQNIRPKMVSVLREKFAHLNL-TFSIGGQISFDVFPQ 186
FEK+ +++N+ M+ RE+ N TFS GG ++F+ + Q
Sbjct: 178 FEKFGEIRNVDIPMLDPYREEMTGRNFHTFSFGGHLNFEAYVQ 220
>PDX1_PHAVU (Q9FT25) Probable pyridoxin biosynthesis protein PDX1
(pvPDX1)
Length = 312
Score = 32.0 bits (71), Expect = 1.4
Identities = 19/52 (36%), Positives = 31/52 (59%), Gaps = 1/52 (1%)
Query: 8 VIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGS 59
V+AL+D +G +T +K S +G Q LR V + VV +D +I+E+ G+
Sbjct: 8 VVALYDGNGAITETKKSPFSVKVGLAQMLRGGVIMDVV-NADQARIAEEAGA 58
>R34K_CLOPA (P23160) 34.2 kDa protein in rubredoxin operon (EC
1.6.4.-) (ORF A)
Length = 308
Score = 31.2 bits (69), Expect = 2.4
Identities = 18/80 (22%), Positives = 42/80 (52%), Gaps = 11/80 (13%)
Query: 31 GFMQELRKVVTVGVVGGSDLIKISEQLGS-----TVTHDYDYV----FSENGLVAHKQGK 81
G + + + +V VG GG+ ++ + L T+ H +DY+ +S++ L HK K
Sbjct: 141 GALYQGKDLVVVG--GGNSAVEAAIFLTKYARNITIVHQFDYLQAQKYSQDELFKHKNVK 198
Query: 82 LIGTESLKTFLGDEKLKEFI 101
+I ++ +G+ ++++ +
Sbjct: 199 IIWDSEIRNIVGENEIEKIV 218
>PGP_PYRFU (Q8U111) Phosphoglycolate phosphatase (EC 3.1.3.18) (PGP)
Length = 231
Score = 30.8 bits (68), Expect = 3.2
Identities = 26/103 (25%), Positives = 49/103 (47%), Gaps = 8/103 (7%)
Query: 13 DVDGTLTAPRKVANSEMLGFMQELRKVVTVGV---VGGSDLIKISEQLGSTVTHDYDYVF 69
D+DGT+T P ++ + E L Q +RK ++GV + + ++ +E + V
Sbjct: 9 DIDGTITYPNRMIHEEAL---QAIRKAESLGVPIMLVTGNTVQFAEAASILIGTSGPVVA 65
Query: 70 SENGLVAHKQGKLIGTE-SLKTFLGDEKLKEFINF-TLHYIAD 110
+ G +++K+ ++ T + L E K F N T H + D
Sbjct: 66 EDGGAISYKKKRIFLTSMDEEWILWGELKKRFPNVKTSHTMPD 108
>NIPB_DROME (Q7PLI2) Nipped-B protein (SCC2 homolog)
Length = 2077
Score = 30.8 bits (68), Expect = 3.2
Identities = 20/58 (34%), Positives = 32/58 (54%), Gaps = 4/58 (6%)
Query: 88 LKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFIEFRSGMLNVSP-IGRNCSQEERDE 144
+K + EKL INFTL LD+ +K +++F+ ML + P G + + E RD+
Sbjct: 1940 IKKQMSQEKLSTDINFTLTKEEKLDLVVK---YLDFKQLMLKLDPDDGDSDADESRDK 1994
>Y143_HAEIN (P44540) Putative HTH-type transcriptional regulator
HI0143
Length = 288
Score = 30.4 bits (67), Expect = 4.1
Identities = 17/64 (26%), Positives = 27/64 (41%)
Query: 31 GFMQELRKVVTVGVVGGSDLIKISEQLGSTVTHDYDYVFSENGLVAHKQGKLIGTESLKT 90
G+ E + + G+ + ++ L S VT DYV QG IGT+ +
Sbjct: 194 GYSAETAHTIKIAKQNGATTVALTHSLRSPVTEYADYVLVNGNKQGKLQGDSIGTKIAQL 253
Query: 91 FLGD 94
F+ D
Sbjct: 254 FVLD 257
>SPT1_MOUSE (Q9R097) Kunitz-type protease inhibitor 1 precursor
(Hepatocyte growth factor activator inhibitor type 1)
(HAI-1)
Length = 507
Score = 30.4 bits (67), Expect = 4.1
Identities = 14/30 (46%), Positives = 17/30 (56%), Gaps = 3/30 (10%)
Query: 185 PQGWDKTYCLRYLDGFNE---IHFFGDKTY 211
P G D+ C +Y GF+E IHF DK Y
Sbjct: 339 PDGSDEATCEKYTSGFDELQNIHFLSDKGY 368
>SYH_AQUAE (O66522) Histidyl-tRNA synthetase (EC 6.1.1.21)
(Histidine--tRNA ligase) (HisRS)
Length = 403
Score = 30.0 bits (66), Expect = 5.4
Identities = 24/81 (29%), Positives = 38/81 (46%), Gaps = 10/81 (12%)
Query: 53 ISEQLGSTVTHD--YDYVFSE-NGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHY-- 107
+S++LG T+ YDY+ E G G +G E L L DE+ KE + F + +
Sbjct: 270 VSDELGLTLIAGGRYDYLVEELGGPPTPALGFALGVERLMLLLPDEEEKEEVYFVIPFGD 329
Query: 108 -----IADLDIPIKRGTFIEF 123
+ DI K+G +E+
Sbjct: 330 VHEYALRVADILRKKGKVVEY 350
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.140 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,900,131
Number of Sequences: 164201
Number of extensions: 1310242
Number of successful extensions: 3009
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 2978
Number of HSP's gapped (non-prelim): 26
length of query: 247
length of database: 59,974,054
effective HSP length: 107
effective length of query: 140
effective length of database: 42,404,547
effective search space: 5936636580
effective search space used: 5936636580
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0284.4


