

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.6
(561 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
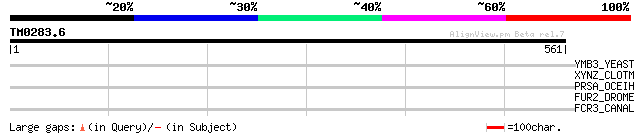
Score E
Sequences producing significant alignments: (bits) Value
YMB3_YEAST (Q04228) Hypothetical protein YML013W 33 2.0
XYNZ_CLOTM (P10478) Endo-1,4-beta-xylanase Z precursor (EC 3.2.1... 31 9.8
PRSA_OCEIH (Q8CXK4) Foldase protein prsA precursor (EC 5.2.1.8) 31 9.8
FUR2_DROME (P30432) Furin-like protease 2 precursor (EC 3.4.21.7... 31 9.8
FCR3_CANAL (Q8X229) Fluconazole resistance protein 3 31 9.8
>YMB3_YEAST (Q04228) Hypothetical protein YML013W
Length = 584
Score = 33.1 bits (74), Expect = 2.0
Identities = 20/68 (29%), Positives = 36/68 (52%), Gaps = 4/68 (5%)
Query: 478 KIPKSPWMPLHMLIAAI----NDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRLIVGD 533
K+PK+P H+ +A I DK ++ K + +A +I++++F + ++VGD
Sbjct: 182 KVPKAPTRETHIPLAEILGDTKDKDAFCELKSFKPDISFNEALRIAKEEFKFMLLILVGD 241
Query: 534 TLLRDTIT 541
T DT T
Sbjct: 242 TYDTDTDT 249
>XYNZ_CLOTM (P10478) Endo-1,4-beta-xylanase Z precursor (EC 3.2.1.8)
(Xylanase Z) (1,4-beta-D-xylan xylanohydrolase Z)
Length = 837
Score = 30.8 bits (68), Expect = 9.8
Identities = 26/101 (25%), Positives = 43/101 (41%), Gaps = 13/101 (12%)
Query: 61 GQSYLRFYLNYKKSARPERMMFYNHGEWVDYPKDIVDLVKKDFEIKN-----AAIGIRLN 115
GQ YL + Y + A P+ ++FYN DY +I DL K + N G+ ++
Sbjct: 664 GQDYLDYAFRYAREADPDALLFYN-----DY--NIEDLGPKSNAVFNMIKSMKERGVPID 716
Query: 116 GRDLMLDFLHACQMDLKTGFIQPIAWIDKVGGYF-FPEVDV 155
G F++ + Q I ++G F E+D+
Sbjct: 717 GVGFQCHFINGMSPEYLASIDQNIKRYAEIGVIVSFTEIDI 757
>PRSA_OCEIH (Q8CXK4) Foldase protein prsA precursor (EC 5.2.1.8)
Length = 299
Score = 30.8 bits (68), Expect = 9.8
Identities = 23/76 (30%), Positives = 35/76 (45%), Gaps = 5/76 (6%)
Query: 154 DVASGEESYNFGEQEGMKFHVEIKMNMVDEFQLRECIGESNALVKDA--QVKVRNMMSIM 211
DV EES E + EI +N VD+ Q++E I N L++DA Q+ V M +
Sbjct: 227 DVREKEESIGEFEDVKKELEREILLNRVDQTQIQEKI---NKLIQDAGVQINVEGMEDLF 283
Query: 212 DCGYIGEEIAKNEELG 227
+ E + + G
Sbjct: 284 EAEETEENTTEEDAQG 299
>FUR2_DROME (P30432) Furin-like protease 2 precursor (EC 3.4.21.75)
(Furin 2)
Length = 1679
Score = 30.8 bits (68), Expect = 9.8
Identities = 24/88 (27%), Positives = 38/88 (42%), Gaps = 9/88 (10%)
Query: 435 DKNNVSRDNSSGANSDSHCSGDPVQSASSAYNEIAGNGVACTQKIPKSPWMPLHMLIAAI 494
D + +DN + + C+G+ A+ A+N G GVA I + ML +
Sbjct: 443 DSDPTPQDNGDNKHG-TRCAGEV---AAVAFNNFCGVGVAYNASIGG-----VRMLDGKV 493
Query: 495 NDKVPPNDMSLIKENFELYKAKQISRDD 522
ND V +SL + ++Y A DD
Sbjct: 494 NDVVEAQALSLNPSHIDIYSASWGPEDD 521
>FCR3_CANAL (Q8X229) Fluconazole resistance protein 3
Length = 399
Score = 30.8 bits (68), Expect = 9.8
Identities = 11/35 (31%), Positives = 27/35 (76%)
Query: 437 NNVSRDNSSGANSDSHCSGDPVQSASSAYNEIAGN 471
NN+S D++ GANS++ SG+ ++++ +++ ++G+
Sbjct: 127 NNMSPDSTGGANSNNFTSGNKRKASNESFSPLSGH 161
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 66,521,770
Number of Sequences: 164201
Number of extensions: 2872975
Number of successful extensions: 5862
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 5861
Number of HSP's gapped (non-prelim): 5
length of query: 561
length of database: 59,974,054
effective HSP length: 115
effective length of query: 446
effective length of database: 41,090,939
effective search space: 18326558794
effective search space used: 18326558794
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0283.6


