

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.14.1
(63 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
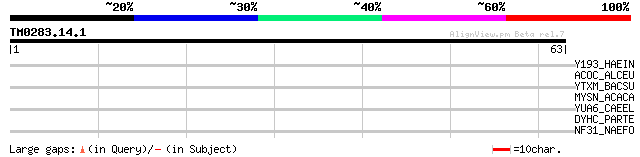
Score E
Sequences producing significant alignments: (bits) Value
Y193_HAEIN (Q57427) Putative esterase/lipase HI0193 (EC 3.1.-.-) 33 0.18
ACOC_ALCEU (P27747) Dihydrolipoyllysine-residue acetyltransferas... 31 0.53
YTXM_BACSU (P23974) Putative esterase ytxM (EC 3.1.-.-) 30 1.5
MYSN_ACACA (P05659) Myosin II heavy chain, non muscle 28 4.5
YUA6_CAEEL (P52715) Putative serine carboxypeptidase F13S12.6 pr... 28 5.8
DYHC_PARTE (Q27171) Dynein heavy chain, cytosolic (DYHC) 27 7.6
NF31_NAEFO (P42661) Virulence-related protein Nf314 (EC 3.4.16.-) 27 9.9
>Y193_HAEIN (Q57427) Putative esterase/lipase HI0193 (EC 3.1.-.-)
Length = 287
Score = 32.7 bits (73), Expect = 0.18
Identities = 18/51 (35%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Query: 7 GENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
G N K E ++ + EQ + F I +GH VH E+P R +KRF+
Sbjct: 235 GGNSSYIKIENSEKILEQFPNATAFT-INGSGHWVHAEKPDFVIRAIKRFL 284
>ACOC_ALCEU (P27747) Dihydrolipoyllysine-residue acetyltransferase
component of acetoin cleaving system (EC 2.3.1.12)
(Acetoin dehydrogenase E2 component) (Dihydrolipoamide
acetyltransferase component of acetoin cleaving system)
(Fast-migrating protein) (
Length = 373
Score = 31.2 bits (69), Expect = 0.53
Identities = 18/57 (31%), Positives = 28/57 (48%), Gaps = 4/57 (7%)
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
R+ ++WG DQI A E GAT + AGH+ +E+ +N LK+ +
Sbjct: 319 RVLVVWGGQDQIIPAAHA----EAAPPGATVKVFADAGHMSQMEKANDFNALLKKHL 371
>YTXM_BACSU (P23974) Putative esterase ytxM (EC 3.1.-.-)
Length = 274
Score = 29.6 bits (65), Expect = 1.5
Identities = 10/26 (38%), Positives = 18/26 (68%)
Query: 34 IKKAGHLVHLERPCVYNRCLKRFIAS 59
+ KAGH VH+E+P ++ + + F+ S
Sbjct: 248 VPKAGHTVHVEQPRLFGKIVSEFLTS 273
>MYSN_ACACA (P05659) Myosin II heavy chain, non muscle
Length = 1509
Score = 28.1 bits (61), Expect = 4.5
Identities = 16/44 (36%), Positives = 22/44 (49%), Gaps = 1/44 (2%)
Query: 19 KNMKEQLG-DGATFQGIKKAGHLVHLERPCVYNRCLKRFIASFF 61
KN K LG + F G++ G L +L P V + KR+ A F
Sbjct: 75 KNEKNFLGVNPPKFDGVEDMGELGYLNEPAVLHNLKKRYDADLF 118
>YUA6_CAEEL (P52715) Putative serine carboxypeptidase F13S12.6
precursor (EC 3.4.16.-)
Length = 454
Score = 27.7 bits (60), Expect = 5.8
Identities = 11/44 (25%), Positives = 23/44 (52%)
Query: 14 KRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
K ++ + + G TF ++ AGH+V ++P V ++ F+
Sbjct: 406 KGQIGGYVTQYKGSQVTFATVRGAGHMVPTDKPAVAEHIIQSFL 449
>DYHC_PARTE (Q27171) Dynein heavy chain, cytosolic (DYHC)
Length = 4540
Score = 27.3 bits (59), Expect = 7.6
Identities = 15/52 (28%), Positives = 23/52 (43%)
Query: 6 WGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
W E +IFK+ + K Q F+ IK + H++R + LK I
Sbjct: 414 WDEEYKIFKQSIVKKSVHQKDQYGQFEHIKLQKQIQHIQRLREMHENLKEVI 465
>NF31_NAEFO (P42661) Virulence-related protein Nf314 (EC 3.4.16.-)
Length = 482
Score = 26.9 bits (58), Expect = 9.9
Identities = 14/35 (40%), Positives = 19/35 (54%)
Query: 23 EQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
E+ G G TF ++ AGH+V L +P K FI
Sbjct: 443 EKSGKGLTFITVRGAGHMVPLVKPDSAFYMFKNFI 477
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.328 0.142 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,383,775
Number of Sequences: 164201
Number of extensions: 225610
Number of successful extensions: 728
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 723
Number of HSP's gapped (non-prelim): 7
length of query: 63
length of database: 59,974,054
effective HSP length: 39
effective length of query: 24
effective length of database: 53,570,215
effective search space: 1285685160
effective search space used: 1285685160
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0283.14.1


