

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0279a.6
(1582 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
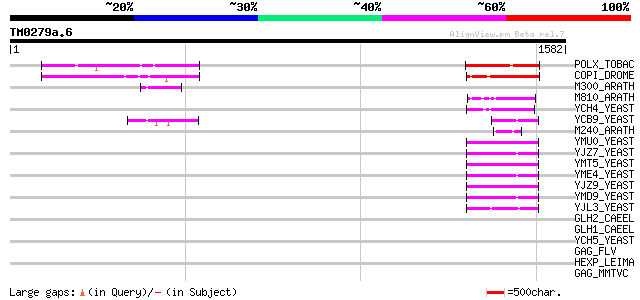
Score E
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 305 5e-82
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 163 3e-39
M300_ARATH (P93293) Hypothetical mitochondrial protein AtMg00300... 104 2e-21
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 98 2e-19
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 80 4e-14
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 54 3e-06
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 51 2e-05
YMU0_YEAST (Q04670) Transposon Ty1 protein B 50 6e-05
YJZ7_YEAST (P47098) Transposon Ty1 protein B 49 8e-05
YMT5_YEAST (Q04214) Transposon Ty1 protein B 49 1e-04
YME4_YEAST (Q04711) Transposon Ty1 protein B 49 1e-04
YJZ9_YEAST (P47100) Transposon Ty1 protein B 49 1e-04
YMD9_YEAST (Q03434) Transposon Ty1 protein B 48 2e-04
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 46 7e-04
GLH2_CAEEL (Q966L9) ATP-dependent RNA helicase glh-2 (EC 3.6.1.-... 45 0.002
GLH1_CAEEL (P34689) ATP-dependent RNA helicase glh-1 (EC 3.6.1.-... 42 0.013
YCH5_YEAST (P25601) Transposon Ty5-1 16.0 kDa hypothetical protein 41 0.023
GAG_FLV (P10262) Gag polyprotein [Contains: Core protein p15; Co... 39 0.11
HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding p... 38 0.19
GAG_MMTVC (P11284) Gag polyprotein [Contains: Protein p10; Phosp... 37 0.33
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 305 bits (782), Expect = 5e-82
Identities = 174/461 (37%), Positives = 266/461 (56%), Gaps = 19/461 (4%)
Query: 90 MSKTLMNKLFAKQRLYSLKMQEGGDLQAHVYAFNNILADLTRLGVTVDDEDKTIILLCSL 149
MSKTL NKL+ K++LY+L M EG + +H+ FN ++ L LGV +++EDK I+LL SL
Sbjct: 93 MSKTLTNKLYLKKQLYALHMSEGTNFLSHLNVFNGLITQLANLGVKIEEEDKAILLLNSL 152
Query: 150 PGSYDHLVTTLTYGKDSITLDSISSTLLQHAQRRRSVEEGGGSSGEGLFVKG-GQDRGRG 208
P SYD+L TT+ +GK +I L ++S LL + + R+ E + G+ L +G G+ R
Sbjct: 153 PSSYDNLATTILHGKTTIELKDVTSALLLNEKMRKKPE----NQGQALITEGRGRSYQRS 208
Query: 209 KGKAVDSGKKKRSKSKDR-KTAECYSCKQIGHWKRDCPN-------RSGKSGNSSSAANV 260
SG + +SK++ + + CY+C Q GH+KRDCPN SG+ + ++AA V
Sbjct: 209 SNNYGRSGARGKSKNRSKSRVRNCYNCNQPGHFKRDCPNPRKGKGETSGQKNDDNTAAMV 268
Query: 261 VQSDGSCS--EEDLLCVSYVKCTDAWVLDSGCSCHMTPHREWFNSFKSCDFGYVYLGDDK 318
+D E+ C+ WV+D+ S H TP R+ F + + DFG V +G+
Sbjct: 269 QNNDNVVLFINEEEECMHLSGPESEWVVDTAASHHATPVRDLFCRYVAGDFGTVKMGNTS 328
Query: 319 PCIIKGM*QVKIALDDGGVRTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVS 378
I G+ + I + G L VR+VP++ NLIS L +GY + R++
Sbjct: 329 YSKIAGIGDICIKTNVGCTLVLKDVRHVPDLRMNLISGIALDRDGYESYFANQK--WRLT 386
Query: 379 KGAMTVMRAKRTAGNIYKLLGSTVMGDVASVETDDDATKLWHMRLGHLSERGMMELYKRN 438
KG++ + + G +Y+ G++ + + D+ + LWH R+GH+SE+G+ L K++
Sbjct: 387 KGSLVIAKGV-ARGTLYRTNAEICQGELNAAQ-DEISVDLWHKRMGHMSEKGLQILAKKS 444
Query: 439 LLKGVRSCTIGLCKYCVLGKQCRVRFKTGQHKTKGILDYVHSDVWGPTKEPSVGGFRYFV 498
L+ + T+ C YC+ GKQ RV F+T + ILD V+SDV GP + S+GG +YFV
Sbjct: 445 LISYAKGTTVKPCDYCLFGKQHRVSFQTSSERKLNILDLVYSDVCGPMEIESMGGNKYFV 504
Query: 499 TFTDDFSRKVWVYFLKYKSEVFAKFKLWKAEVENQTGRKIK 539
TF DD SRK+WVY LK K +VF F+ + A VE +TGRK+K
Sbjct: 505 TFIDDASRKLWVYILKTKDQVFQVFQKFHALVERETGRKLK 545
Score = 244 bits (624), Expect = 1e-63
Identities = 116/211 (54%), Positives = 153/211 (71%), Gaps = 3/211 (1%)
Query: 1300 AKKILGMEIHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKTA 1359
A++ILGM+I RE+ ++KLWLSQ Y+E VL RF+M A PVSTPLA H KLS + CP T
Sbjct: 1041 AQQILGMKIVRERTSRKLWLSQEKYIERVLERFNMKNAKPVSTPLAGHLKLSKKMCPTTV 1100
Query: 1360 SEIEGMSKISHASAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKG 1419
E M+K+ ++SAVG LMY MVCTRPD+A A V +F+ PGK+HWEAVKWILRYL+G
Sbjct: 1101 EEKGNMAKVPYSSAVGSLMYAMVCTRPDIAHAVGVVSRFLENPGKEHWEAVKWILRYLRG 1160
Query: 1420 TADRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAM 1479
T + F G P++ GY D+D AGD+D+R+S+TGY+FT +GG I W+S +Q VA+
Sbjct: 1161 TTGDCLCFG---GSDPILKGYTDADMAGDIDNRKSSTGYLFTFSGGAISWQSKLQKCVAL 1217
Query: 1480 STTEAEYMAVAEAVKEALWLTGLVKKLGVEQ 1510
STTEAEY+A E KE +WL +++LG+ Q
Sbjct: 1218 STTEAEYIAATETGKEMIWLKRFLQELGLHQ 1248
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 163 bits (413), Expect = 3e-39
Identities = 86/210 (40%), Positives = 130/210 (60%), Gaps = 10/210 (4%)
Query: 1301 KKILGMEIHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKTAS 1360
K +G+ I ++ K++LSQ++YV+ +LS+F+M N VSTPL + L + +
Sbjct: 1121 KHFIGIRIEMQED--KIYLSQSAYVKKILSKFNMENCNAVSTPLPSKINYELLNSDEDCN 1178
Query: 1361 EIEGMSKISHASAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGT 1420
S +GCLMY+M+CTRPDL A + + ++ SK + W+ +K +LRYLKGT
Sbjct: 1179 T-------PCRSLIGCLMYIMLCTRPDLTTAVNILSRYSSKNNSELWQNLKRVLRYLKGT 1231
Query: 1421 ADRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAG-GPICWKSSVQSIVAM 1479
D ++F + ++GYVDSD+AG DR+STTGY+F + ICW + Q+ VA
Sbjct: 1232 IDMKLIFKKNLAFENKIIGYVDSDWAGSEIDRKSTTGYLFKMFDFNLICWNTKRQNSVAA 1291
Query: 1480 STTEAEYMAVAEAVKEALWLTGLVKKLGVE 1509
S+TEAEYMA+ EAV+EALWL L+ + ++
Sbjct: 1292 SSTEAEYMALFEAVREALWLKFLLTSINIK 1321
Score = 140 bits (352), Expect = 4e-32
Identities = 122/470 (25%), Positives = 209/470 (43%), Gaps = 35/470 (7%)
Query: 92 KTLMNKLFAKQRLYSLKMQEGGDLQAHVYAFNNILADLTRLGVTVDDEDKTIILLCSLPG 151
K+L ++L ++RL SLK+ L +H + F+ ++++L G +++ DK LL +LP
Sbjct: 91 KSLASQLALRKRLLSLKLSSEMSLLSHFHIFDELISELLAAGAKIEEMDKISHLLITLPS 150
Query: 152 SYDHLVTTL-TYGKDSITLDSISSTLLQHAQRRRSVEEGGGSSGEGLFVKGGQDRGRGK- 209
YD ++T + T ++++TL + + LL + ++ V + +
Sbjct: 151 CYDGIITAIETLSEENLTLAFVKNRLLDQEIKIKNDHNDTSKKVMNAIVHNNNNTYKNNL 210
Query: 210 GKAVDSGKKKRSKSKDRKTAECYSCKQIGHWKRDCPNRSGKSGNSSSAANVVQSDGSCSE 269
K + KK K + +C+ C + GH K+DC + + N+ + N Q + S
Sbjct: 211 FKNRVTKPKKIFKGNSKYKVKCHHCGREGHIKKDCFHYK-RILNNKNKENEKQVQTATSH 269
Query: 270 EDLLCVSYVKCTD-----AWVLDSGCSCHMTPHREWFNS----FKSCDFGYVYLGDDKPC 320
V V T +VLDSG S H+ + G+
Sbjct: 270 GIAFMVKEVNNTSVMDNCGFVLDSGASDHLINDESLYTDSVEVVPPLKIAVAKQGEFIYA 329
Query: 321 IIKGM*QVKIALDDGGVRTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVSKG 380
+G+ + L + TL V + E NL+S+ L E G S E ++ + +SK
Sbjct: 330 TKRGI----VRLRNDHEITLEDVLFCKEAAGNLMSVKRLQEAGMSI--EFDKSGVTISKN 383
Query: 381 AMTVMRAKRTAGNIYKLLGSTVMGDVASVET-DDDATKLWHMRLGHLSERGMMELYKRNL 439
+ V++ N+ + S+ + +LWH R GH+S+ ++E+ ++N+
Sbjct: 384 GLMVVKNSGMLNNV-----PVINFQAYSINAKHKNNFRLWHERFGHISDGKLLEIKRKNM 438
Query: 440 LKGVR-------SCTIGLCKYCVLGKQCRVRFKTGQHKT--KGILDYVHSDVWGPTKEPS 490
SC I C+ C+ GKQ R+ FK + KT K L VHSDV GP +
Sbjct: 439 FSDQSLLNNLELSCEI--CEPCLNGKQARLPFKQLKDKTHIKRPLFVVHSDVCGPITPVT 496
Query: 491 VGGFRYFVTFTDDFSRKVWVYFLKYKSEVFAKFKLWKAEVENQTGRKIKY 540
+ YFV F D F+ Y +KYKS+VF+ F+ + A+ E K+ Y
Sbjct: 497 LDDKNYFVIFVDQFTHYCVTYLIKYKSDVFSMFQDFVAKSEAHFNLKVVY 546
>M300_ARATH (P93293) Hypothetical mitochondrial protein AtMg00300
(ORF145a) (ORF1451)
Length = 145
Score = 104 bits (259), Expect = 2e-21
Identities = 51/116 (43%), Positives = 69/116 (58%), Gaps = 1/116 (0%)
Query: 374 ILRVSKGAMTVMRAKRTAGNIYKLLGSTVMGDVASVETDDDATKLWHMRLGHLSERGMME 433
+L+V KG T+++ R ++Y L GS G+ ET D T+LWH RL H+S+RGM
Sbjct: 28 VLKVLKGCRTILKGNRH-DSLYILQGSVETGESNLAETAKDETRLWHSRLAHMSQRGMEL 86
Query: 434 LYKRNLLKGVRSCTIGLCKYCVLGKQCRVRFKTGQHKTKGILDYVHSDVWGPTKEP 489
L K+ L + ++ C+ C+ GK RV F TGQH TK LDYVHSD+WG P
Sbjct: 87 LVKKGFLDSSKVSSLKFCEDCIYGKTHRVNFSTGQHTTKNPLDYVHSDLWGAPSVP 142
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 97.8 bits (242), Expect = 2e-19
Identities = 68/195 (34%), Positives = 97/195 (48%), Gaps = 12/195 (6%)
Query: 1304 LGMEIHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKTASEIE 1363
LG++I L+LSQ Y E +L+ M P+STPL L L TA +
Sbjct: 43 LGIQIKTHPSG--LFLSQTKYAEQILNNAGMLDCKPMSTPLP----LKLNSSVSTAKYPD 96
Query: 1364 GMSKISHASAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADR 1423
S VG L Y+ + TRPD++ A + VC+ M +P ++ +K +LRY+KGT
Sbjct: 97 PSD---FRSIVGALQYLTL-TRPDISYAVNIVCQRMHEPTLADFDLLKRVLRYVKGTIFH 152
Query: 1424 GIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTE 1483
G+ + + V + DSD+AG RRSTTG+ L I W + Q V+ S+TE
Sbjct: 153 GLYIHKNSKLN--VQAFCDSDWAGCTSTRRSTTGFCTFLGCNIISWSAKRQPTVSRSSTE 210
Query: 1484 AEYMAVAEAVKEALW 1498
EY A+A E W
Sbjct: 211 TEYRALALTAAELTW 225
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 80.1 bits (196), Expect = 4e-14
Identities = 56/195 (28%), Positives = 93/195 (46%), Gaps = 10/195 (5%)
Query: 1302 KILGMEIHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKTASE 1361
K LG+ IH+ + LS Y+ S +++ TPL N L T+
Sbjct: 122 KFLGLNIHQSTNGD-ITLSLQDYIAKAASESEINTFKLTQTPLCNSKPLF----ETTSPH 176
Query: 1362 IEGMSKISHASAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTA 1421
++ ++ + S VG L++ RPD++ S + +F+ +P H E+ + +LRYL T
Sbjct: 177 LKDITP--YQSIVGQLLFCANTGRPDISYPVSLLSRFLREPRAIHLESARRVLRYLYTT- 233
Query: 1422 DRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKS-SVQSIVAMS 1480
R + G + Y D+ + D ST GYV LAG P+ W S ++ ++ +
Sbjct: 234 -RSMCLKYRSGSQVALTVYCDASHGAIHDLPHSTGGYVTLLAGAPVTWSSKKLKGVIPVP 292
Query: 1481 TTEAEYMAVAEAVKE 1495
+TEAEY+ +E V E
Sbjct: 293 STEAEYITASETVME 307
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 53.9 bits (128), Expect = 3e-06
Identities = 53/226 (23%), Positives = 94/226 (41%), Gaps = 27/226 (11%)
Query: 336 GVRTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVSKGAMTVMRAKRTAGNIY 395
G +T + + P + +L+SL L + + R+ L S G + K G+ Y
Sbjct: 505 GTKTSIKALHTPNIAYDLLSLSELANQNIT--ACFTRNTLERSDGTVLAPIVKH--GDFY 560
Query: 396 KLLGSTVMGDVASVETDDDATK----------LWHMRLGHLSERGMMELYKRNLLKGVRS 445
L ++ S T ++ K L H LGH + R + + K+N + ++
Sbjct: 561 WLSKKYLIPSHISKLTINNVNKSKSVNKYPYPLIHRMLGHANFRSIQKSLKKNAVTYLKE 620
Query: 446 CTIGL-------CKYCVLGKQCRVRFKTGQH----KTKGILDYVHSDVWGPTKEPSVGGF 494
I C C++GK + R G ++ Y+H+D++GP
Sbjct: 621 SDIEWSNASTYQCPDCLIGKSTKHRHVKGSRLKYQESYEPFQYLHTDIFGPVHHLPKSAP 680
Query: 495 RYFVTFTDDFSRKVWVYFLKYKSE--VFAKFKLWKAEVENQTGRKI 538
YF++FTD+ +R WVY L + E + F A ++NQ ++
Sbjct: 681 SYFISFTDEKTRFQWVYPLHDRREESILNVFTSILAFIKNQFNARV 726
Score = 47.4 bits (111), Expect = 3e-04
Identities = 38/135 (28%), Positives = 68/135 (50%), Gaps = 3/135 (2%)
Query: 1374 VGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADRGIMFSREQGV 1433
+G YV R DL + + + + P +Q + +++++ T D+ +++ + +
Sbjct: 1555 IGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPT 1614
Query: 1434 VP--LVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAE 1491
P +V D+ Y G+ +S G +F L G I KS+ S+ STTEAE AV+E
Sbjct: 1615 KPDNKLVAISDASY-GNQPYYKSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAEIHAVSE 1673
Query: 1492 AVKEALWLTGLVKKL 1506
A+ L+ LV++L
Sbjct: 1674 AIPLLNNLSHLVQEL 1688
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 51.2 bits (121), Expect = 2e-05
Identities = 29/81 (35%), Positives = 46/81 (55%), Gaps = 3/81 (3%)
Query: 1378 MYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADRGIMFSREQGVVPLV 1437
MY+ + TRPDL A +++ +F S +AV +L Y+KGT +G+ +S + +
Sbjct: 1 MYLTI-TRPDLTFAVNRLSQFSSASRTAQMQAVYKVLHYVKGTVGQGLFYSATSDL--QL 57
Query: 1438 VGYVDSDYAGDLDDRRSTTGY 1458
+ DSD+A D RRS TG+
Sbjct: 58 KAFADSDWASCPDTRRSVTGF 78
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 49.7 bits (117), Expect = 6e-05
Identities = 55/208 (26%), Positives = 96/208 (45%), Gaps = 6/208 (2%)
Query: 1303 ILGMEIHREKGAKKLWLSQNSYVEGV--LSRFDMSKANPVSTPLANHFKLSLEQCPKTAS 1360
ILG+EI ++G +NS E + L+ K +S P L ++Q
Sbjct: 1041 ILGLEIKYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAP--GQPGLYIDQQELELE 1098
Query: 1361 EIEGMSKISHASA-VGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKG 1419
E + K+ +G YV R DL + + + + P KQ + +++++
Sbjct: 1099 EDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWN 1158
Query: 1420 TADRGIMFSREQGVVPLVVGYVDSDYA-GDLDDRRSTTGYVFTLAGGPICWKSSVQSIVA 1478
T D+ +++ + + V P V SD + G+ +S G ++ L G I KS+ S+
Sbjct: 1159 TRDKQLIWHKSKPVKPTNKLVVISDASYGNQPYYKSQIGNIYLLNGKVIGGKSTKASLTC 1218
Query: 1479 MSTTEAEYMAVAEAVKEALWLTGLVKKL 1506
STTEAE A++E+V L+ LV++L
Sbjct: 1219 TSTTEAEIHAISESVPLLNNLSHLVQEL 1246
Score = 43.5 bits (101), Expect = 0.005
Identities = 50/219 (22%), Positives = 89/219 (39%), Gaps = 27/219 (12%)
Query: 338 RTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVSKGAMTVMRAKRTAGNIY-- 395
+T +V + P + +L+SL L + +++L S G TV+ G+ Y
Sbjct: 84 KTSIKVLHTPNIAYDLLSLNELA--AVDITACFTKNVLERSDG--TVLAPIVKYGDFYWV 139
Query: 396 ---KLLGSTV-MGDVASVETDDDATK----LWHMRLGHLSERGMMELYKRNLLKGVRSCT 447
LL S + + + +V T + K H L H + + + K N +
Sbjct: 140 SKKYLLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESD 199
Query: 448 IGL-------CKYCVLGKQCRVRFKTGQH----KTKGILDYVHSDVWGPTKEPSVGGFRY 496
+ C C++GK + R G + Y+H+D++GP Y
Sbjct: 200 VDWSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSY 259
Query: 497 FVTFTDDFSRKVWVYFLKYKSE--VFAKFKLWKAEVENQ 533
F++FTD+ ++ WVY L + E + F A ++NQ
Sbjct: 260 FISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 298
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 49.3 bits (116), Expect = 8e-05
Identities = 54/209 (25%), Positives = 96/209 (45%), Gaps = 8/209 (3%)
Query: 1303 ILGMEIHREKGAKKLWLSQNSYVEGV--LSRFDMSKANPVSTPLANHFKLSLEQCPKTAS 1360
ILG+EI ++G +NS E + L+ K +S P L ++Q
Sbjct: 1468 ILGLEIKYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAP--GQPGLYIDQDELEID 1525
Query: 1361 EIEGMSKISHASA-VGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKG 1419
E E K+ +G YV R DL + + + + P +Q + +++++
Sbjct: 1526 EDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWD 1585
Query: 1420 TADRGIMFSREQGVVP--LVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIV 1477
T D+ +++ + + P +V D+ Y G+ +S G +F L G I KS+ S+
Sbjct: 1586 TRDKQLIWHKNKPTEPDNKLVAISDASY-GNQPYYKSQIGNIFLLNGKVIGGKSTKASLT 1644
Query: 1478 AMSTTEAEYMAVAEAVKEALWLTGLVKKL 1506
STTEAE A++E+V L+ L+++L
Sbjct: 1645 CTSTTEAEIHAISESVPLLNNLSYLIQEL 1673
Score = 43.5 bits (101), Expect = 0.005
Identities = 50/219 (22%), Positives = 89/219 (39%), Gaps = 27/219 (12%)
Query: 338 RTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVSKGAMTVMRAKRTAGNIY-- 395
+T +V + P + +L+SL L + +++L S G TV+ G+ Y
Sbjct: 511 KTSIKVLHTPNIAYDLLSLNELA--AVDITACFTKNVLERSDG--TVLAPIVQYGDFYWV 566
Query: 396 ---KLLGSTV-MGDVASVETDDDATK----LWHMRLGHLSERGMMELYKRNLLKGVRSCT 447
LL S + + + +V T + K H L H + + + K N +
Sbjct: 567 SKRYLLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESD 626
Query: 448 IGL-------CKYCVLGKQCRVRFKTGQH----KTKGILDYVHSDVWGPTKEPSVGGFRY 496
+ C C++GK + R G + Y+H+D++GP Y
Sbjct: 627 VDWSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSY 686
Query: 497 FVTFTDDFSRKVWVYFLKYKSE--VFAKFKLWKAEVENQ 533
F++FTD+ ++ WVY L + E + F A ++NQ
Sbjct: 687 FISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 725
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 48.9 bits (115), Expect = 1e-04
Identities = 54/208 (25%), Positives = 96/208 (45%), Gaps = 6/208 (2%)
Query: 1303 ILGMEIHREKGAKKLWLSQNSYVEGV--LSRFDMSKANPVSTPLANHFKLSLEQCPKTAS 1360
ILG+EI ++G +NS E + L+ K +S P L ++Q
Sbjct: 1041 ILGLEIKYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAP--GQPGLYIDQQELELE 1098
Query: 1361 EIEGMSKISHASA-VGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKG 1419
E + K+ +G YV R DL + + + + P KQ + +++++
Sbjct: 1099 EDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWN 1158
Query: 1420 TADRGIMFSREQGVVPLVVGYVDSDYA-GDLDDRRSTTGYVFTLAGGPICWKSSVQSIVA 1478
T D+ +++ + + V P V SD + G+ +S G ++ L G I KS+ S+
Sbjct: 1159 TRDKQLIWHKSKPVKPTNKLVVISDASYGNQPYYKSQIGNIYLLNGKVIGGKSTKASLTC 1218
Query: 1479 MSTTEAEYMAVAEAVKEALWLTGLVKKL 1506
STTEAE A++E+V L+ L+++L
Sbjct: 1219 TSTTEAEIHAISESVPLLNNLSYLIQEL 1246
Score = 43.5 bits (101), Expect = 0.005
Identities = 50/219 (22%), Positives = 89/219 (39%), Gaps = 27/219 (12%)
Query: 338 RTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVSKGAMTVMRAKRTAGNIY-- 395
+T +V + P + +L+SL L + +++L S G TV+ G+ Y
Sbjct: 84 KTSIKVLHTPNIAYDLLSLNELA--AVDITACFTKNVLERSDG--TVLAPIVKYGDFYWV 139
Query: 396 ---KLLGSTV-MGDVASVETDDDATK----LWHMRLGHLSERGMMELYKRNLLKGVRSCT 447
LL S + + + +V T + K H L H + + + K N +
Sbjct: 140 SKKYLLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESD 199
Query: 448 IGL-------CKYCVLGKQCRVRFKTGQH----KTKGILDYVHSDVWGPTKEPSVGGFRY 496
+ C C++GK + R G + Y+H+D++GP Y
Sbjct: 200 VDWSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSY 259
Query: 497 FVTFTDDFSRKVWVYFLKYKSE--VFAKFKLWKAEVENQ 533
F++FTD+ ++ WVY L + E + F A ++NQ
Sbjct: 260 FISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 298
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 48.9 bits (115), Expect = 1e-04
Identities = 54/209 (25%), Positives = 96/209 (45%), Gaps = 8/209 (3%)
Query: 1303 ILGMEIHREKGAKKLWLSQNSYVEGV--LSRFDMSKANPVSTPLANHFKLSLEQCPKTAS 1360
ILG+EI ++G +NS E + L+ K +S P L ++Q
Sbjct: 1041 ILGLEIKYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAP--GQPGLYIDQDELEID 1098
Query: 1361 EIEGMSKISHASA-VGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKG 1419
E E K+ +G YV R DL + + + + P +Q + +++++
Sbjct: 1099 EDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWD 1158
Query: 1420 TADRGIMFSREQGVVP--LVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIV 1477
T D+ +++ + + P +V D+ Y G+ +S G ++ L G I KS+ S+
Sbjct: 1159 TRDKQLIWHKNKPTEPDNKLVAISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLT 1217
Query: 1478 AMSTTEAEYMAVAEAVKEALWLTGLVKKL 1506
STTEAE A++E+V L+ LV++L
Sbjct: 1218 CTSTTEAEIHAISESVPLLNNLSHLVQEL 1246
Score = 43.5 bits (101), Expect = 0.005
Identities = 50/219 (22%), Positives = 89/219 (39%), Gaps = 27/219 (12%)
Query: 338 RTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVSKGAMTVMRAKRTAGNIY-- 395
+T +V + P + +L+SL L + +++L S G TV+ G+ Y
Sbjct: 84 KTSIKVLHTPNIAYDLLSLNELA--AVDITACFTKNVLERSDG--TVLAPIVKYGDFYWV 139
Query: 396 ---KLLGSTV-MGDVASVETDDDATK----LWHMRLGHLSERGMMELYKRNLLKGVRSCT 447
LL S + + + +V T + K H L H + + + K N +
Sbjct: 140 SKKYLLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESD 199
Query: 448 IGL-------CKYCVLGKQCRVRFKTGQH----KTKGILDYVHSDVWGPTKEPSVGGFRY 496
+ C C++GK + R G + Y+H+D++GP Y
Sbjct: 200 VDWSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSY 259
Query: 497 FVTFTDDFSRKVWVYFLKYKSE--VFAKFKLWKAEVENQ 533
F++FTD+ ++ WVY L + E + F A ++NQ
Sbjct: 260 FISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 298
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 48.9 bits (115), Expect = 1e-04
Identities = 54/208 (25%), Positives = 96/208 (45%), Gaps = 6/208 (2%)
Query: 1303 ILGMEIHREKGAKKLWLSQNSYVEGV--LSRFDMSKANPVSTPLANHFKLSLEQCPKTAS 1360
ILG+EI ++G +NS E + L+ K +S P L ++Q
Sbjct: 1468 ILGLEIKYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAP--GQPGLYIDQQELELE 1525
Query: 1361 EIEGMSKISHASA-VGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKG 1419
E + K+ +G YV R DL + + + + P KQ + +++++
Sbjct: 1526 EDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELIQFIWN 1585
Query: 1420 TADRGIMFSREQGVVPLVVGYVDSDYA-GDLDDRRSTTGYVFTLAGGPICWKSSVQSIVA 1478
T D+ +++ + + V P V SD + G+ +S G ++ L G I KS+ S+
Sbjct: 1586 TRDKQLIWHKSKPVKPTNKLVVISDASYGNQPYYKSQIGNIYLLNGKVIGGKSTKASLTC 1645
Query: 1479 MSTTEAEYMAVAEAVKEALWLTGLVKKL 1506
STTEAE A++E+V L+ L+++L
Sbjct: 1646 TSTTEAEIHAISESVPLLNNLSYLIQEL 1673
Score = 43.5 bits (101), Expect = 0.005
Identities = 50/219 (22%), Positives = 89/219 (39%), Gaps = 27/219 (12%)
Query: 338 RTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVSKGAMTVMRAKRTAGNIY-- 395
+T +V + P + +L+SL L + +++L S G TV+ G+ Y
Sbjct: 511 KTSIKVLHTPNIAYDLLSLNELA--AVDITACFTKNVLERSDG--TVLAPIVKYGDFYWV 566
Query: 396 ---KLLGSTV-MGDVASVETDDDATK----LWHMRLGHLSERGMMELYKRNLLKGVRSCT 447
LL S + + + +V T + K H L H + + + K N +
Sbjct: 567 SKKYLLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESD 626
Query: 448 IGL-------CKYCVLGKQCRVRFKTGQH----KTKGILDYVHSDVWGPTKEPSVGGFRY 496
+ C C++GK + R G + Y+H+D++GP Y
Sbjct: 627 VDWSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSY 686
Query: 497 FVTFTDDFSRKVWVYFLKYKSE--VFAKFKLWKAEVENQ 533
F++FTD+ ++ WVY L + E + F A ++NQ
Sbjct: 687 FISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 725
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 48.1 bits (113), Expect = 2e-04
Identities = 53/209 (25%), Positives = 96/209 (45%), Gaps = 8/209 (3%)
Query: 1303 ILGMEIHREKGAKKLWLSQNSYVEGV--LSRFDMSKANPVSTPLANHFKLSLEQCPKTAS 1360
ILG+EI ++G +NS E + L+ K +S P L ++Q
Sbjct: 1041 ILGLEIKYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAP--GQPGLYIDQDELEID 1098
Query: 1361 EIEGMSKISHASA-VGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKG 1419
E E K+ +G YV R DL + + + + P +Q + +++++
Sbjct: 1099 EDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWD 1158
Query: 1420 TADRGIMFSREQGVVP--LVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIV 1477
T D+ +++ + + P +V D+ Y G+ +S G ++ L G I KS+ S+
Sbjct: 1159 TRDKQLIWHKNKPTEPDNKLVAISDASY-GNQPYYKSQIGNIYLLNGKVIGGKSTKASLT 1217
Query: 1478 AMSTTEAEYMAVAEAVKEALWLTGLVKKL 1506
STTEAE A++E+V L+ L+++L
Sbjct: 1218 CTSTTEAEIHAISESVPLLNNLSYLIQEL 1246
Score = 43.9 bits (102), Expect = 0.004
Identities = 53/219 (24%), Positives = 91/219 (41%), Gaps = 27/219 (12%)
Query: 338 RTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVSKGAMTVMRAKRTAGNIY-- 395
+T +V + P + +L+SL L + +++L S G TV+ G+ Y
Sbjct: 84 KTSIKVLHTPNIAYDLLSLNELA--AVDITACFTKNVLERSDG--TVLAPIVKYGDFYWV 139
Query: 396 ---KLLGSTV-MGDVASVETDDDATK----LWHMRLGHLSERGMMELYKRNLLKGV---- 443
LL S + + + +V T + K H L H + + + K N +
Sbjct: 140 SKKYLLPSNISVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESD 199
Query: 444 --RSCTIGL-CKYCVLGKQCRVRFKTGQH----KTKGILDYVHSDVWGPTKEPSVGGFRY 496
RS I C C++GK + R G + Y+H+D++GP Y
Sbjct: 200 VDRSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSY 259
Query: 497 FVTFTDDFSRKVWVYFLKYKSE--VFAKFKLWKAEVENQ 533
F++FTD+ ++ WVY L + E + F A ++NQ
Sbjct: 260 FISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQ 298
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 46.2 bits (108), Expect = 7e-04
Identities = 49/210 (23%), Positives = 95/210 (44%), Gaps = 8/210 (3%)
Query: 1303 ILGMEIHREKGAKKLWLSQNSYVEGVLSRF--DMSKANPVSTPLANHFKLSLEQCPKTAS 1360
ILGM++ K + L+ S++ + ++ ++ K S P + +K+ ++ S
Sbjct: 1503 ILGMDLVYNKRLGTIDLTLKSFINRMDKKYNEELKKIRKSSIPHMSTYKIDPKKDVLQMS 1562
Query: 1361 EIE---GMSKISHASAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYL 1417
E E G+ K+ +G L YV R D+ A +V + ++ P ++ + + I++YL
Sbjct: 1563 EEEFRQGVLKLQQL--LGELNYVRHKCRYDIEFAVKKVARLVNYPHERVFYMIYKIIQYL 1620
Query: 1418 KGTADRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIV 1477
D GI + R+ V+ D+ + D +S G + S+ +
Sbjct: 1621 VRYKDIGIHYDRDCNKDKKVIAITDASVGSEYD-AQSRIGVILWYGMNIFNVYSNKSTNR 1679
Query: 1478 AMSTTEAEYMAVAEAVKEALWLTGLVKKLG 1507
+S+TEAE A+ E ++ L +K+LG
Sbjct: 1680 CVSSTEAELHAIYEGYADSETLKVTLKELG 1709
>GLH2_CAEEL (Q966L9) ATP-dependent RNA helicase glh-2 (EC 3.6.1.-)
(Germline helicase-2)
Length = 974
Score = 45.1 bits (105), Expect = 0.002
Identities = 30/91 (32%), Positives = 40/91 (42%), Gaps = 24/91 (26%)
Query: 189 GGGSSGEGLFVKGGQDRGRGKGKA-VDSGKKKRS----------------KSKDRKTAEC 231
GGG+SG F GG G G G D G++ + K+R+ C
Sbjct: 225 GGGNSGGSGFGSGGNSNGFGSGGGGQDRGERNNNCFNCQQPGHRSNDCPEPKKEREPRVC 284
Query: 232 YSCKQIGHWKRDCP-------NRSGKSGNSS 255
Y+C+Q GH RDCP R+G +G SS
Sbjct: 285 YNCQQPGHNSRDCPEERKPREGRNGFTGGSS 315
Score = 42.7 bits (99), Expect = 0.008
Identities = 27/77 (35%), Positives = 38/77 (49%), Gaps = 11/77 (14%)
Query: 189 GGGSSGEGLFVKGGQDRGRGKGKAVDSGKKKRSKS------KDRKTAECYSCKQIGHWKR 242
GG S+G G GGQDRG + + + K+R+ CY+C+Q GH R
Sbjct: 351 GGNSNGFGSG-GGGQDRGERNNNCFNCQQPGHRSNDCPEPKKEREPRVCYNCQQPGHNSR 409
Query: 243 DCPN----RSGKSGNSS 255
DCP R G++G +S
Sbjct: 410 DCPEERKPREGRNGFTS 426
Score = 35.4 bits (80), Expect = 1.3
Identities = 29/103 (28%), Positives = 40/103 (38%), Gaps = 23/103 (22%)
Query: 181 QRRRSVEEGGGSSGEGLFVKGGQDRGRGKGKAVDSGKKKRS--------KSKDRKTAEC- 231
+R+ G +SG G GG D G G G A G + K + ++AEC
Sbjct: 414 ERKPREGRNGFTSGFG----GGNDGGFGGGNAEGFGNNEERGPMKCFNCKGEGHRSAECP 469
Query: 232 ------YSCKQIGHWKRDCPN----RSGKSGNSSSAANVVQSD 264
++C + GH +CPN R G G A V D
Sbjct: 470 EPPRGCFNCGEQGHRSNECPNPAKPREGAEGEGPKATYVPVED 512
Score = 35.0 bits (79), Expect = 1.6
Identities = 19/57 (33%), Positives = 27/57 (47%), Gaps = 5/57 (8%)
Query: 189 GGGSSGEGLFVKGGQDRGRGKGKAVDSGKKKRSKSKDRKTAECYSCKQIGHWKRDCP 245
G G++GE F GG G G G + R + + C++C+Q GH DCP
Sbjct: 336 GSGNNGESGFGSGGFG-GNSNGFGSGGGGQDRGERNNN----CFNCQQPGHRSNDCP 387
>GLH1_CAEEL (P34689) ATP-dependent RNA helicase glh-1 (EC 3.6.1.-)
(Germline helicase-1)
Length = 763
Score = 42.0 bits (97), Expect = 0.013
Identities = 26/85 (30%), Positives = 37/85 (42%), Gaps = 7/85 (8%)
Query: 189 GGGSSGEGLFVKGGQDRGRGKGKAVDSGKKKRSKSKDRKTAE---CYSCKQIGHWKRDCP 245
GGG+SG G GG +R G + + RK E CY+C+Q GH R+C
Sbjct: 140 GGGNSGFGEGGHGGGERNNNCFNCQQPGHRSSDCPEPRKEREPRVCYNCQQPGHTSRECT 199
Query: 246 N----RSGKSGNSSSAANVVQSDGS 266
R G++G A + G+
Sbjct: 200 EERKPREGRTGGFGGGAGFGNNGGN 224
Score = 35.0 bits (79), Expect = 1.6
Identities = 28/100 (28%), Positives = 38/100 (38%), Gaps = 16/100 (16%)
Query: 181 QRRRSVEEGGGSSGEGLFVKGGQDR-----GRGKGKAVDSGKKKRSKSKDRKTAEC---- 231
++ R GG G G GG D G G G+ K K + ++AEC
Sbjct: 202 RKPREGRTGGFGGGAGFGNNGGNDGFGGDGGFGGGEERGPMKCFNCKGEGHRSAECPEPP 261
Query: 232 ---YSCKQIGHWKRDCPN----RSGKSGNSSSAANVVQSD 264
++C + GH +CPN R G G A V D
Sbjct: 262 RGCFNCGEQGHRSNECPNPAKPREGVEGEGPKATYVPVED 301
>YCH5_YEAST (P25601) Transposon Ty5-1 16.0 kDa hypothetical protein
Length = 146
Score = 41.2 bits (95), Expect = 0.023
Identities = 37/118 (31%), Positives = 52/118 (43%), Gaps = 16/118 (13%)
Query: 250 KSGNSSSAANVV-------QSDGSCSEEDLLCV--SYVKCTDA--WVLDSGCSCHMTPHR 298
K+ SSSA+ + Q+ G + + LC+ S V T + W+ D+GC+ HM R
Sbjct: 34 KTSRSSSASVAIPDYETQGQTAGQITPKSWLCMLSSTVPATKSSEWIFDTGCTSHMCHDR 93
Query: 299 EWFNSFKSCDFGYVYLGDDKPCIIKGM*QVKIALDDGGVRTLSQVRYVPEVTKNLISL 356
F+SF G I G V I G L V YVP++ NLIS+
Sbjct: 94 SIFSSFTRSSRKDFVRGVGGSIPIMGSGTVNI-----GTVQLHDVSYVPDLPVNLISV 146
>GAG_FLV (P10262) Gag polyprotein [Contains: Core protein p15; Core
protein p12; Core protein p30; Core protein p10]
Length = 580
Score = 38.9 bits (89), Expect = 0.11
Identities = 21/62 (33%), Positives = 29/62 (45%), Gaps = 6/62 (9%)
Query: 199 VKGGQDRGRGKGKAVDSGKKKRSKSKDRKTAECYSCKQIGHWKRDCPNRSGKSGNSSSAA 258
V +D+ R + K D K K +C CK+ GHW RDCP R K +S+
Sbjct: 523 VAQNRDKDREENKLGDQRKIPLGKD------QCAYCKEKGHWVRDCPKRPRKKPANSTLL 576
Query: 259 NV 260
N+
Sbjct: 577 NL 578
>HEXP_LEIMA (Q04832) DNA-binding protein HEXBP (Hexamer-binding
protein)
Length = 271
Score = 38.1 bits (87), Expect = 0.19
Identities = 33/107 (30%), Positives = 41/107 (37%), Gaps = 16/107 (14%)
Query: 179 HAQRRRSVEEGGGSSGEGLFVKGGQDRGRGKGKAVDSGKKKRSKSKDRKTAECYSCKQIG 238
+ Q+R GG SG+ K G D G + +G+ S + DR CY C G
Sbjct: 123 YGQKRGRSGAQGGYSGDRTCYKCG-DAGH-ISRDCPNGQGGYSGAGDRT---CYKCGDAG 177
Query: 239 HWKRDCPNRSG-----------KSGNSSSAANVVQSDGSCSEEDLLC 274
H RDCPN G K G S + S GS D C
Sbjct: 178 HISRDCPNGQGGYSGAGDRKCYKCGESGHMSRECPSAGSTGSGDRAC 224
Score = 35.8 bits (81), Expect = 0.96
Identities = 23/71 (32%), Positives = 26/71 (36%), Gaps = 17/71 (23%)
Query: 179 HAQRRRSVEEGGGSSGEGLFVKGGQDRGRGKGKAVDSGKKKRSKSKDRKTAECYSCKQIG 238
H R +GG G G Q RGR + SG + CY C G
Sbjct: 107 HLSRDCPSSQGGSRGGYG------QKRGRSGAQGGYSGDRT-----------CYKCGDAG 149
Query: 239 HWKRDCPNRSG 249
H RDCPN G
Sbjct: 150 HISRDCPNGQG 160
Score = 35.0 bits (79), Expect = 1.6
Identities = 23/71 (32%), Positives = 26/71 (36%), Gaps = 17/71 (23%)
Query: 227 KTAECYSCKQIGHWKRDCPNRSGKSGNS-------SSAANVVQSDGSCSEEDLLCVSYVK 279
K ECY C Q GH RDCP+ G S S A D +C K
Sbjct: 95 KGFECYKCGQEGHLSRDCPSSQGGSRGGYGQKRGRSGAQGGYSGDRTC----------YK 144
Query: 280 CTDAWVLDSGC 290
C DA + C
Sbjct: 145 CGDAGHISRDC 155
Score = 33.5 bits (75), Expect = 4.8
Identities = 13/30 (43%), Positives = 19/30 (63%), Gaps = 2/30 (6%)
Query: 225 DRKTAECYSCKQIGHWKRDCPN--RSGKSG 252
D ++ C+ C + GH R+CPN RSG +G
Sbjct: 39 DERSTTCFRCGEEGHMSRECPNEARSGAAG 68
>GAG_MMTVC (P11284) Gag polyprotein [Contains: Protein p10;
Phosphorylated protein pp21; Protein p3; Protein p8;
Major core protein p27; Nucleic acid binding protein
p14]
Length = 591
Score = 37.4 bits (85), Expect = 0.33
Identities = 22/65 (33%), Positives = 29/65 (43%), Gaps = 2/65 (3%)
Query: 189 GGGSSGEGLFVKGGQDRGRGKGKAVDSGKKKRSKSKDRKTAECYSCKQIGHWKRDCPNRS 248
GGG G+G KG GK + K+ SK C CK+ HWK +C ++
Sbjct: 514 GGGKGGQGS--KGPVCFSCGKTGHIKRDCKEEKGSKRAPPGLCPRCKKGYHWKSECKSKF 571
Query: 249 GKSGN 253
K GN
Sbjct: 572 DKDGN 576
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.341 0.149 0.500
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 167,135,227
Number of Sequences: 164201
Number of extensions: 6645323
Number of successful extensions: 21710
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 32
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 21557
Number of HSP's gapped (non-prelim): 132
length of query: 1582
length of database: 59,974,054
effective HSP length: 124
effective length of query: 1458
effective length of database: 39,613,130
effective search space: 57755943540
effective search space used: 57755943540
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0279a.6


