

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0278.5.1
(30 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
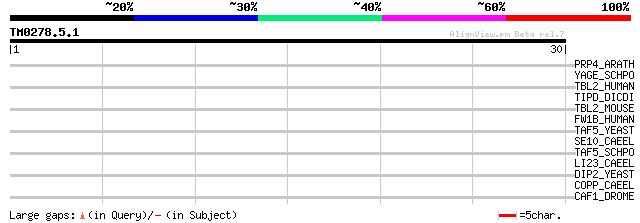
Score E
Sequences producing significant alignments: (bits) Value
PRP4_ARATH (O22212) Hypothetical Trp-Asp repeats containing prot... 36 0.018
YAGE_SCHPO (Q9UTN4) Hypothetical WD-repeat protein C12G12.14c in... 30 1.7
TBL2_HUMAN (Q9Y4P3) Transducin beta-like 2 protein (WS beta-tran... 28 3.8
TIPD_DICDI (O15736) Protein tipD 28 4.9
TBL2_MOUSE (Q9R099) Transducin beta-like 2 protein 28 4.9
FW1B_HUMAN (Q9UKB1) F-box/WD-repeat protein 11 (F-box/WD-repeat ... 28 4.9
TAF5_YEAST (P38129) Transcription initiation factor TFIID subuni... 28 6.4
SE10_CAEEL (Q93794) F-box/WD-repeat protein sel-10 (Suppressor/e... 28 6.4
TAF5_SCHPO (O13282) Transcription initiation factor TFIID subuni... 27 8.4
LI23_CAEEL (Q09990) F-box/WD-repeat protein lin-23 (Abnormal cel... 27 8.4
DIP2_YEAST (Q12220) DOM34 interacting protein 2 27 8.4
COPP_CAEEL (Q20168) Probable coatomer beta' subunit (Beta'-coat ... 27 8.4
CAF1_DROME (Q24572) Chromatin assembly factor 1 P55 subunit (CAF... 27 8.4
>PRP4_ARATH (O22212) Hypothetical Trp-Asp repeats containing protein
At2g41500
Length = 554
Score = 36.2 bits (82), Expect = 0.018
Identities = 17/30 (56%), Positives = 22/30 (72%), Gaps = 3/30 (10%)
Query: 4 IVTVSHDRTIKLWSSNMTNDNE---HTMDV 30
I TVSHDRTIKLW+S+ +D + TMD+
Sbjct: 523 IATVSHDRTIKLWTSSGNDDEDEEKETMDI 552
>YAGE_SCHPO (Q9UTN4) Hypothetical WD-repeat protein C12G12.14c in
chromosome I
Length = 509
Score = 29.6 bits (65), Expect = 1.7
Identities = 12/29 (41%), Positives = 18/29 (61%)
Query: 1 GGYIVTVSHDRTIKLWSSNMTNDNEHTMD 29
G + T S+D+T + WS + +D E TMD
Sbjct: 372 GHLLCTGSNDKTTRFWSRSRPDDKESTMD 400
>TBL2_HUMAN (Q9Y4P3) Transducin beta-like 2 protein (WS
beta-transducin repeats protein) (WS-betaTRP)
(Williams-Beuren syndrome chromosome region 13 protein)
(UNQ563/PRO1125)
Length = 447
Score = 28.5 bits (62), Expect = 3.8
Identities = 11/26 (42%), Positives = 16/26 (61%)
Query: 1 GGYIVTVSHDRTIKLWSSNMTNDNEH 26
G Y+ T + DRTI++WS+ EH
Sbjct: 102 GKYLATCADDRTIRIWSTKDFLQREH 127
>TIPD_DICDI (O15736) Protein tipD
Length = 612
Score = 28.1 bits (61), Expect = 4.9
Identities = 11/13 (84%), Positives = 12/13 (91%)
Query: 4 IVTVSHDRTIKLW 16
+VT SHDRTIKLW
Sbjct: 422 VVTGSHDRTIKLW 434
>TBL2_MOUSE (Q9R099) Transducin beta-like 2 protein
Length = 442
Score = 28.1 bits (61), Expect = 4.9
Identities = 10/26 (38%), Positives = 16/26 (61%)
Query: 1 GGYIVTVSHDRTIKLWSSNMTNDNEH 26
G Y+ T + DRT+++WS+ EH
Sbjct: 98 GKYLATCADDRTVRIWSTKDFLQREH 123
>FW1B_HUMAN (Q9UKB1) F-box/WD-repeat protein 11 (F-box/WD-repeat
protein 1B) (F-box and WD-repeats protein beta-TrCP2)
Length = 542
Score = 28.1 bits (61), Expect = 4.9
Identities = 11/17 (64%), Positives = 15/17 (87%)
Query: 3 YIVTVSHDRTIKLWSSN 19
YIV+ S DRTIK+WS++
Sbjct: 375 YIVSASGDRTIKVWSTS 391
>TAF5_YEAST (P38129) Transcription initiation factor TFIID subunit 5
(TBP-associated factor 5) (TBP-associated factor 90 kDa)
(TAFII-90)
Length = 798
Score = 27.7 bits (60), Expect = 6.4
Identities = 10/17 (58%), Positives = 12/17 (69%)
Query: 1 GGYIVTVSHDRTIKLWS 17
G Y T SHD+T +LWS
Sbjct: 579 GHYFATASHDQTARLWS 595
>SE10_CAEEL (Q93794) F-box/WD-repeat protein sel-10
(Suppressor/enhancer of lin-12) (Sel-10 protein)
Length = 587
Score = 27.7 bits (60), Expect = 6.4
Identities = 11/18 (61%), Positives = 15/18 (83%)
Query: 1 GGYIVTVSHDRTIKLWSS 18
G YIV+ S DRT+K+WS+
Sbjct: 308 GRYIVSGSTDRTVKVWST 325
>TAF5_SCHPO (O13282) Transcription initiation factor TFIID subunit 5
(Transcription initiation factor TFIID 72 kDa subunit)
(TAFII-72)
Length = 643
Score = 27.3 bits (59), Expect = 8.4
Identities = 10/17 (58%), Positives = 12/17 (69%)
Query: 1 GGYIVTVSHDRTIKLWS 17
G Y T SHD+T +LWS
Sbjct: 433 GHYFATASHDQTAQLWS 449
>LI23_CAEEL (Q09990) F-box/WD-repeat protein lin-23 (Abnormal cell
lineage protein 23)
Length = 665
Score = 27.3 bits (59), Expect = 8.4
Identities = 11/15 (73%), Positives = 13/15 (86%)
Query: 3 YIVTVSHDRTIKLWS 17
YIV+ S DRTIK+WS
Sbjct: 357 YIVSASGDRTIKVWS 371
>DIP2_YEAST (Q12220) DOM34 interacting protein 2
Length = 943
Score = 27.3 bits (59), Expect = 8.4
Identities = 8/16 (50%), Positives = 14/16 (87%)
Query: 1 GGYIVTVSHDRTIKLW 16
GG++V+ SHD +I++W
Sbjct: 669 GGFVVSSSHDHSIRIW 684
>COPP_CAEEL (Q20168) Probable coatomer beta' subunit (Beta'-coat
protein) (Beta'-COP)
Length = 1000
Score = 27.3 bits (59), Expect = 8.4
Identities = 12/26 (46%), Positives = 18/26 (69%), Gaps = 1/26 (3%)
Query: 4 IVTVSHDRTIKLWSSNMTNDNEHTMD 29
I+T S D T++LW +N T EHT++
Sbjct: 244 IITGSEDSTVRLWHAN-TYRQEHTLN 268
>CAF1_DROME (Q24572) Chromatin assembly factor 1 P55 subunit (CAF-1
P55 subunit) (dCAF-1) (Nucleosome remodeling factor 55
kDa subunit) (NURF-55)
Length = 430
Score = 27.3 bits (59), Expect = 8.4
Identities = 11/28 (39%), Positives = 15/28 (53%)
Query: 2 GYIVTVSHDRTIKLWSSNMTNDNEHTMD 29
GY+++ S D TI LW N T +D
Sbjct: 195 GYLLSASDDHTICLWDINATPKEHRVID 222
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.128 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,611,788
Number of Sequences: 164201
Number of extensions: 54456
Number of successful extensions: 525
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 497
Number of HSP's gapped (non-prelim): 29
length of query: 30
length of database: 59,974,054
effective HSP length: 6
effective length of query: 24
effective length of database: 58,988,848
effective search space: 1415732352
effective search space used: 1415732352
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0278.5.1


