

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0269b.2
(2064 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
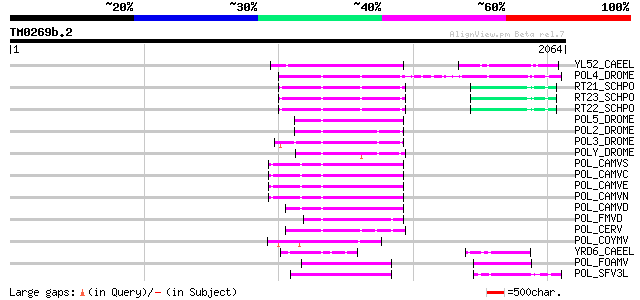
Score E
Sequences producing significant alignments: (bits) Value
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 229 6e-59
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 226 7e-58
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 196 6e-49
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 193 4e-48
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 193 4e-48
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 190 3e-47
POL2_DROME (P20825) Retrovirus-related Pol polyprotein from tran... 189 5e-47
POL3_DROME (P04323) Retrovirus-related Pol polyprotein from tran... 187 2e-46
POLY_DROME (P10401) Retrovirus-related Pol polyprotein from tran... 156 6e-37
POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic pro... 149 8e-35
POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic pro... 147 2e-34
POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic pro... 146 7e-34
POL_CAMVN (Q00962) Enzymatic polyprotein [Contains: Aspartic pro... 145 9e-34
POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic pro... 144 3e-33
POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic prot... 130 4e-29
POL_CERV (P05400) Enzymatic polyprotein [Contains: Aspartic prot... 126 7e-28
POL_COYMV (P19199) Putative polyprotein [Contains: Coat protein;... 123 6e-27
YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II 119 9e-26
POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse transcript... 116 7e-25
POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.2... 113 5e-24
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 229 bits (584), Expect = 6e-59
Identities = 153/504 (30%), Positives = 262/504 (51%), Gaps = 15/504 (2%)
Query: 969 FSKKMEAQDPLEEVDLGDGLEKRPTYISALIDPELKDRMV-KLLKEFKDCFAWDYDEMPG 1027
FS+ + E V + E+ T L DR + ++++F+D FA DE+
Sbjct: 867 FSEIETDTNSCEVVKTAETYERFTTICEHLKRENGDDRKIWDVIEQFQDVFAISDDELGR 926
Query: 1028 LSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVV 1087
S E + +KE +P++Q PR + +I++ I+++L K IR ++ W + VV
Sbjct: 927 NSG--TECVIELKEGAEPIRQKPRPIPLALKPEIRKMIQKMLNQKVIRESKS-PWSSPVV 983
Query: 1088 PVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIA 1147
V KK+G +R+CID+R +N + + +P E + S AG + ++ D +G+ QI +
Sbjct: 984 LVKKKDGSIRMCIDYRKVNKVVKNNAHPLPNIEATLQSLAGKKLYTVFDMIAGFWQIPLD 1043
Query: 1148 EEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVK 1207
E+ TAF L +EW V+PFGL + A +Q M I D + VY+DD+++
Sbjct: 1044 EKSKEITAFAIGSEL--FEWNVLPFGLVISPALFQGTMEEIIGDLLGVCAFVYVDDLLIA 1101
Query: 1208 SPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAIL 1267
S + HL ++++ R+RK G+K+ KC ++LG V G+E + K +
Sbjct: 1102 SKDMEQHLQDVKEALTRIRKSGMKLRASKCHIAKKEVEYLGHKVTLDGVETQEVKTDKMK 1161
Query: 1268 DTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDEL 1327
S PT+ K+LQS LG + + R+FI N ++ S + L+ K A+ WE E + AF EL
Sbjct: 1162 QFSRPTNVKELQSFLGLVGYYRKFILNFAQIASSLTSLISAKV--AWIWEKEQEIAFQEL 1219
Query: 1328 KVYLSSPPIMASP-----IRG-KPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLN 1381
K + P++A P ++G +P +Y A+ IG++LAQE D ++ I + S+ L+
Sbjct: 1220 KKLVCQTPVLAQPDVEAALKGDRPFMIYTDASRKGIGAVLAQEGPDGQQHPIAFASKALS 1279
Query: 1382 DAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALA 1441
AETRY + + L + F+ + K I + VF+ + + +L L R+ +W++
Sbjct: 1280 PAETRYHITDLEALAMMFALRRFKTIIYGTAITVFTDHKPLISLLKGSPLADRLWRWSIE 1339
Query: 1442 LTEYSLTYAPLKAVKGQAIADFLA 1465
+ E+ + L A K A+AD L+
Sbjct: 1340 ILEFDVKIVYL-AGKANAVADALS 1362
Score = 119 bits (297), Expect = 1e-25
Identities = 95/379 (25%), Positives = 174/379 (45%), Gaps = 20/379 (5%)
Query: 1669 YVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPT 1728
Y I+G L ++ + E + +H+G+ H GIK W + + YWP
Sbjct: 1440 YKIVGGVLKNTEIEEQSRSVVPEKIRTPLLKELHEGMLAGH-FGIKKMWRMVHRKFYWPQ 1498
Query: 1729 I---MKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQH 1785
+ +++C+ +C H+ + S L +P A DL+ S + +
Sbjct: 1499 MRVCVENCVRTCAKCLCANDHSKL----TSSLTPYRMTFPLEIVACDLMDV--GLSVQGN 1552
Query: 1786 KYIIVAIDYFTKWVEAIPLQNVTQDTVIE-FISSIVYRFG-LPESLTTDQGTVFVGQKVA 1843
+YI+ ID FTK+ A+P+ + +TV++ F+ G +P L TDQG FV A
Sbjct: 1553 RYILTIIDLFTKYGTAVPIPDKKAETVLKAFVERWAIGEGRIPLKLLTDQGKEFVNGLFA 1612
Query: 1844 SFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNS 1903
F I+ +T+ Y ++ANG VE NKT++ ++KK P W + ++AY N
Sbjct: 1613 QFTHMLKIEHITTKGYNSRANGAVERFNKTIMHIMKKKTA-VPMEWDDQVVYAVYAYNNC 1671
Query: 1904 PKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDEERLLV 1963
E+TG TP L +G++ + P+E+ + ++ ++Y +++ EL+ + + +
Sbjct: 1672 VHENTGETPMFLMHGRDVMGPLEMSGEDAVGINYADM--DEYKHLLTQELLKVQK---IA 1726
Query: 1964 LDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVL--PINKKDKRYRKWVPNWEGPFTVE 2021
+ R+++ +++K +K +V+L P K + K V W GP+ V
Sbjct: 1727 KEHAMREQESYKSLFDQKYASKKHRFPQPGSRVLLEIPSEKLGAQCPKLVNKWSGPYRVI 1786
Query: 2022 KVLVNNAYSIKELGGNRQM 2040
N+A LG + +
Sbjct: 1787 SCSENSAEITPVLGKRKHI 1805
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-); Reverse
transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 226 bits (575), Expect = 7e-58
Identities = 241/1062 (22%), Positives = 461/1062 (42%), Gaps = 110/1062 (10%)
Query: 1001 PEL-KDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLV 1059
PEL K ++ + E+ D FA + + P + + +L +K+D+ + R H V
Sbjct: 272 PELFKSQLENICSEYIDIFALESE--PITVNNLYKQQLRLKDDEPVYTKNYRSPHSQV-E 328
Query: 1060 KIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNG------KMRVCIDFRDLNAATPKDE 1113
+I+ ++++L++ K + + + + ++ V KK+ K R+ ID+R +N D+
Sbjct: 329 EIQAQVQKLIKDKIVEPS-VSQYNSPLLLVPKKSSPNSDKKKWRLVIDYRQINKKLLADK 387
Query: 1114 YHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFG 1173
+ +P + ++D +Y S LD SG++QI + E T+F G+Y + +PFG
Sbjct: 388 FPLPRIDDILDQLGRAKYFSCLDLMSGFHQIELDEGSRDITSFSTSN--GSYRFTRLPFG 445
Query: 1174 LKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMN 1233
LK A ++QR+M F + +Y+DD++V S L +L + F + R+Y LK++
Sbjct: 446 LKIAPNSFQRMMTIAFSGIEPSQAFLYMDDLIVIGCSEKHMLKNLTEVFGKCREYNLKLH 505
Query: 1234 PLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIA 1293
P KC+F + FLG KGI + K I + P + + N+ RRFI
Sbjct: 506 PEKCSFFMHEVTFLGHKCTDKGILPDDKKYDVIQNYPVPHDADSARRFVAFCNYYRRFIK 565
Query: 1294 NLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISAT 1353
N ++ ++ + L KK F W E QKAF LK L +P ++ P K + A+
Sbjct: 566 NFADYSRHITRL--CKKNVPFEWTDECQKAFIHLKSQLINPTLLQYPDFSKEFCITTDAS 623
Query: 1354 DGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDV 1413
G++L Q + + + + Y SR E+ + E+ ++++ + + YI
Sbjct: 624 KQACGAVLTQ-NHNGHQLPVAYASRAFTKGESNKSTTEQELAAIHWAIIHFRPYIYGKHF 682
Query: 1414 MVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTLPQEI 1473
V + + + ++ S S++ + L L EY+ T LK K +AD L+ T+ +
Sbjct: 683 TVKTDHRPLTYLFSMVNPSSKLTRIRLELEEYNFTVEYLKG-KDNHVADALSRITIKE-- 739
Query: 1474 ITYVGIQPWKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGL 1533
K+ TG + + T+F+ R KS + + + T
Sbjct: 740 -----------------LKDITG------NILKVTTRFQSRQKSCAGKEQLDLQKQTK-- 774
Query: 1534 EILIALGAKNVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRV 1593
EI V+ + V+ + C+ ++ ++A++D L +
Sbjct: 775 EIASEPNVYEVITNDEVRKVVTLQLNDSICLFKH-------GKKIIARYDVGDLYTNGIL 827
Query: 1594 DNQEANELAQIASGYMVDKCRLKELIEVKEKLNPSDLDILVIDNMAPNDWRKPIVDYLQN 1653
D + + ++ +G + ++ ++K MAP W+K I +++
Sbjct: 828 DLDQFLQRLELQAG-------IYDISQIK---------------MAP--WKK-IFEHV-- 860
Query: 1654 PVGTTDRKTKYRAMSYVIMGNELFKKNVDGTLLKCL----SEDDAFIAISAVHDGLCGAH 1709
+ D+ + MGN++ KN+ LL + +E + +S +HD
Sbjct: 861 ---SIDK--------FKNMGNKIL-KNLKVALLNPVTQINNEKEKEAILSTLHDDPIQGG 908
Query: 1710 QAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGW 1769
GI ++ YW + K EY ++CQ CQ+ +H + F
Sbjct: 909 HTGITKTLAKVKRHYYWKNMSKYIKEYVRKCQKCQKAKTTKHTKTPMTITETPEHAFDRV 968
Query: 1770 ALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFI-SSIVYRFGLPES 1828
+D IG + P S ++Y + I TK++ AIP+ N + TV + I S + ++G ++
Sbjct: 969 VVDTIGPL-PKSENGNEYAVTLICDLTKYLVAIPIANKSAKTVAKAIFESFILKYGPMKT 1027
Query: 1829 LTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKN 1888
TD GT + + + IK +TST ++ Q G VE +++TL I+ ++ +
Sbjct: 1028 FITDMGTEYKNSIITDLCKYLKIKNITSTAHHHQTVGVVERSHRTLNEYIRSYISTDKTD 1087
Query: 1889 WHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNM 1948
W L ++ + + P+ L +G+ + LP ++ E I + D +
Sbjct: 1088 WDVWLQYFVYCFNTTQSMVHNYCPYELVFGRTSNLPKHFN----KLHSIEPIYNID--DY 1141
Query: 1949 MLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYR 2008
+ L+ L K++ + Y+ KV+ +GD KV+L +++
Sbjct: 1142 AKESKYRLEVAYARARKLLEAHKEKNKENYDLKVKDIELEVGD---KVLL----RNEVGH 1194
Query: 2009 KWVPNWEGPFTVEKVLVNNAYSIKELGGNRQMTINGKYLKTY 2050
K + GP+ +E + NN ++ N++ ++ LK +
Sbjct: 1195 KLDFKYTGPYKIESIGDNNNITLL-TNKNKKQIVHKDRLKKF 1235
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 196 bits (498), Expect = 6e-49
Identities = 136/484 (28%), Positives = 251/484 (51%), Gaps = 29/484 (5%)
Query: 1000 DPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKED--KKPVKQLPRRFHPDV 1057
+PEL D + KEFKD A E + +E ++ + ++ + P++ P P
Sbjct: 371 EPELPD----IYKEFKDITAETNTEKLPKPIKGLEFEVELTQENYRLPIRNYP--LPPGK 424
Query: 1058 LVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMP 1117
+ + +EI + L+ IR ++ ++ V+ V KK G +R+ +D++ LN + Y +P
Sbjct: 425 MQAMNDEINQGLKSGIIRESKAIN-ACPVMFVPKKEGTLRMVVDYKPLNKYVKPNIYPLP 483
Query: 1118 IAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNA 1177
+ E ++ G + LD S Y+ I + + D K AFRCP G +E++VMP+G+ A
Sbjct: 484 LIEQLLAKIQGSTIFTKLDLKSAYHLIRVRKGDEHKLAFRCPR--GVFEYLVMPYGISTA 541
Query: 1178 GATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKC 1237
A +Q +NTI + E+ + Y+DDI++ S S +H+ H++ ++++ L +N KC
Sbjct: 542 PAHFQYFINTILGEAKESHVVCYMDDILIHSKSESEHVKHVKDVLQKLKNANLIINQAKC 601
Query: 1238 AFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSE 1297
F F+G+ + +KG + +L P ++K+L+ LG +N+LR+FI S+
Sbjct: 602 EFHQSQVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSVNYLRKFIPKTSQ 661
Query: 1298 KTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTI 1357
T + L LKK+ ++W +A + +K L SPP++ K + L A+D +
Sbjct: 662 LTHPLNNL--LKKDVRWKWTPTQTQAIENIKQCLVSPPVLRHFDFSKKILLETDASDVAV 719
Query: 1358 GSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYY----IKPIDV 1413
G++L+Q+ +D K + Y S ++ A+ Y++ +K L + S ++Y I+P +
Sbjct: 720 GAVLSQKHDDDKYYPVGYYSAKMSKAQLNYSVSDKEMLAIIKSLKHWRHYLESTIEPFKI 779
Query: 1414 MVFSHYDIIKHML--SKPILHSRIGKWALALTEYS--LTYAPLKAVKGQAIADFLA---D 1466
+ H ++I + S+P + R+ +W L L +++ + Y P A IAD L+ D
Sbjct: 780 LT-DHRNLIGRITNESEP-ENKRLARWQLFLQDFNFEINYRPGSA---NHIADALSRIVD 834
Query: 1467 QTLP 1470
+T P
Sbjct: 835 ETEP 838
Score = 84.3 bits (207), Expect = 3e-15
Identities = 78/335 (23%), Positives = 130/335 (38%), Gaps = 43/335 (12%)
Query: 1712 GIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSII---KPWPFRG 1768
GI++ + + W I K EY + C CQ + H P L I +PW
Sbjct: 929 GIELLTNIILRRFTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQPIPPSERPW--ES 986
Query: 1769 WALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQN--VTQDTVIEFISSIVYRFGLP 1826
++D I + S + + V +D F+K +P + T F ++ FG P
Sbjct: 987 LSMDFITALPESSG--YNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVIAYFGNP 1044
Query: 1827 ESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKP 1886
+ + D +F Q FA + + S PY Q +GQ E N+T+ L++ P
Sbjct: 1045 KEIIADNDHIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRCVCSTHP 1104
Query: 1887 KNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYW 1946
W + V +Y N+ +T TPF + + L E+PS
Sbjct: 1105 NTWVDHISLVQQSYNNAIHSATQMTPFEIVHRYSPAL------------SPLELPS---- 1148
Query: 1947 NMMLDELVNLDEERLLVLDT----LTRQKDRIAKAYNKKVR-AKSFMLGDYVWKVVLPIN 2001
D+ +E + V T L ++ K ++ K++ + F GD V +
Sbjct: 1149 --FSDKTDENSQETIQVFQTVKEHLNTNNIKMKKYFDMKIQEIEEFQPGDLV------MV 1200
Query: 2002 KKDK-----RYRKWVPNWEGPFTVEKVLVNNAYSI 2031
K+ K + K P++ GPF V + N Y +
Sbjct: 1201 KRTKTGFLHKSNKLAPSFAGPFYVLQKSGPNNYEL 1235
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 193 bits (491), Expect = 4e-48
Identities = 135/484 (27%), Positives = 251/484 (50%), Gaps = 29/484 (5%)
Query: 1000 DPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKED--KKPVKQLPRRFHPDV 1057
+PEL D + KEFKD A E + +E ++ + ++ + P++ P P
Sbjct: 371 EPELPD----IYKEFKDITAETNTEKLPKPIKGLEFEVELTQENYRLPIRNYP--LPPGK 424
Query: 1058 LVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMP 1117
+ + +EI + L+ IR ++ ++ V+ V KK G +R+ +D++ LN + Y +P
Sbjct: 425 MQAMNDEINQGLKSGIIRESKAIN-ACPVMFVPKKEGTLRMVVDYKPLNKYVKPNIYPLP 483
Query: 1118 IAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNA 1177
+ E ++ G + LD S Y+ I + + D K AFRCP G +E++VMP+G+ A
Sbjct: 484 LIEQLLAKIQGSTIFTKLDLKSAYHLIRVRKGDEHKLAFRCPR--GVFEYLVMPYGISIA 541
Query: 1178 GATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKC 1237
A +Q +NTI + E+ + Y+D+I++ S S +H+ H++ ++++ L +N KC
Sbjct: 542 PAHFQYFINTILGEVKESHVVCYMDNILIHSKSESEHVKHVKDVLQKLKNANLIINQAKC 601
Query: 1238 AFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSE 1297
F F+G+ + +KG + +L P ++K+L+ LG +N+LR+FI S+
Sbjct: 602 EFHQSQVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSVNYLRKFIPKTSQ 661
Query: 1298 KTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTI 1357
T + L LKK+ ++W +A + +K L SPP++ K + L A+D +
Sbjct: 662 LTHPLNNL--LKKDVRWKWTPTQTQAIENIKQCLVSPPVLRHFDFSKKILLETDASDVAV 719
Query: 1358 GSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYY----IKPIDV 1413
G++L+Q+ +D K + Y S ++ A+ Y++ +K L + S ++Y I+P +
Sbjct: 720 GAVLSQKHDDDKYYPVGYYSAKMSKAQLNYSVSDKEMLAIIKSLKHWRHYLESTIEPFKI 779
Query: 1414 MVFSHYDIIKHML--SKPILHSRIGKWALALTEYS--LTYAPLKAVKGQAIADFLA---D 1466
+ H ++I + S+P + R+ +W L L +++ + Y P A IAD L+ D
Sbjct: 780 LT-DHRNLIGRITNESEP-ENKRLARWQLFLQDFNFEINYRPGSA---NHIADALSRIVD 834
Query: 1467 QTLP 1470
+T P
Sbjct: 835 ETEP 838
Score = 84.3 bits (207), Expect = 3e-15
Identities = 78/335 (23%), Positives = 130/335 (38%), Gaps = 43/335 (12%)
Query: 1712 GIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSII---KPWPFRG 1768
GI++ + + W I K EY + C CQ + H P L I +PW
Sbjct: 929 GIELLTNIILRRFTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQPIPPSERPW--ES 986
Query: 1769 WALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQN--VTQDTVIEFISSIVYRFGLP 1826
++D I + S + + V +D F+K +P + T F ++ FG P
Sbjct: 987 LSMDFITALPESSG--YNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVIAYFGNP 1044
Query: 1827 ESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKP 1886
+ + D +F Q FA + + S PY Q +GQ E N+T+ L++ P
Sbjct: 1045 KEIIADNDHIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRCVCSTHP 1104
Query: 1887 KNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYW 1946
W + V +Y N+ +T TPF + + L E+PS
Sbjct: 1105 NTWVDHISLVQQSYNNAIHSATQMTPFEIVHRYSPAL------------SPLELPS---- 1148
Query: 1947 NMMLDELVNLDEERLLVLDT----LTRQKDRIAKAYNKKVR-AKSFMLGDYVWKVVLPIN 2001
D+ +E + V T L ++ K ++ K++ + F GD V +
Sbjct: 1149 --FSDKTDENSQETIQVFQTVKEHLNTNNIKMKKYFDMKIQEIEEFQPGDLV------MV 1200
Query: 2002 KKDK-----RYRKWVPNWEGPFTVEKVLVNNAYSI 2031
K+ K + K P++ GPF V + N Y +
Sbjct: 1201 KRTKTGFLHKSNKLAPSFAGPFYVLQKSGPNNYEL 1235
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 193 bits (491), Expect = 4e-48
Identities = 135/484 (27%), Positives = 251/484 (50%), Gaps = 29/484 (5%)
Query: 1000 DPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKED--KKPVKQLPRRFHPDV 1057
+PEL D + KEFKD A E + +E ++ + ++ + P++ P P
Sbjct: 371 EPELPD----IYKEFKDITAETNTEKLPKPIKGLEFEVELTQENYRLPIRNYP--LPPGK 424
Query: 1058 LVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYHMP 1117
+ + +EI + L+ IR ++ ++ V+ V KK G +R+ +D++ LN + Y +P
Sbjct: 425 MQAMNDEINQGLKSGIIRESKAIN-ACPVMFVPKKEGTLRMVVDYKPLNKYVKPNIYPLP 483
Query: 1118 IAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNA 1177
+ E ++ G + LD S Y+ I + + D K AFRCP G +E++VMP+G+ A
Sbjct: 484 LIEQLLAKIQGSTIFTKLDLKSAYHLIRVRKGDEHKLAFRCPR--GVFEYLVMPYGISIA 541
Query: 1178 GATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKC 1237
A +Q +NTI + E+ + Y+D+I++ S S +H+ H++ ++++ L +N KC
Sbjct: 542 PAHFQYFINTILGEVKESHVVCYMDNILIHSKSESEHVKHVKDVLQKLKNANLIINQAKC 601
Query: 1238 AFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSE 1297
F F+G+ + +KG + +L P ++K+L+ LG +N+LR+FI S+
Sbjct: 602 EFHQSQVKFIGYHISEKGFTPCQENIDKVLQWKQPKNRKELRQFLGSVNYLRKFIPKTSQ 661
Query: 1298 KTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTI 1357
T + L LKK+ ++W +A + +K L SPP++ K + L A+D +
Sbjct: 662 LTHPLNNL--LKKDVRWKWTPTQTQAIENIKQCLVSPPVLRHFDFSKKILLETDASDVAV 719
Query: 1358 GSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYY----IKPIDV 1413
G++L+Q+ +D K + Y S ++ A+ Y++ +K L + S ++Y I+P +
Sbjct: 720 GAVLSQKHDDDKYYPVGYYSAKMSKAQLNYSVSDKEMLAIIKSLKHWRHYLESTIEPFKI 779
Query: 1414 MVFSHYDIIKHML--SKPILHSRIGKWALALTEYS--LTYAPLKAVKGQAIADFLA---D 1466
+ H ++I + S+P + R+ +W L L +++ + Y P A IAD L+ D
Sbjct: 780 LT-DHRNLIGRITNESEP-ENKRLARWQLFLQDFNFEINYRPGSA---NHIADALSRIVD 834
Query: 1467 QTLP 1470
+T P
Sbjct: 835 ETEP 838
Score = 84.3 bits (207), Expect = 3e-15
Identities = 78/335 (23%), Positives = 130/335 (38%), Gaps = 43/335 (12%)
Query: 1712 GIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSII---KPWPFRG 1768
GI++ + + W I K EY + C CQ + H P L I +PW
Sbjct: 929 GIELLTNIILRRFTWKGIRKQIQEYVQNCHTCQINKSRNHKPYGPLQPIPPSERPW--ES 986
Query: 1769 WALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQN--VTQDTVIEFISSIVYRFGLP 1826
++D I + S + + V +D F+K +P + T F ++ FG P
Sbjct: 987 LSMDFITALPESSG--YNALFVVVDRFSKMAILVPCTKSITAEQTARMFDQRVIAYFGNP 1044
Query: 1827 ESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKP 1886
+ + D +F Q FA + + S PY Q +GQ E N+T+ L++ P
Sbjct: 1045 KEIIADNDHIFTSQTWKDFAHKYNFVMKFSLPYRPQTDGQTERTNQTVEKLLRCVCSTHP 1104
Query: 1887 KNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYW 1946
W + V +Y N+ +T TPF + + L E+PS
Sbjct: 1105 NTWVDHISLVQQSYNNAIHSATQMTPFEIVHRYSPAL------------SPLELPS---- 1148
Query: 1947 NMMLDELVNLDEERLLVLDT----LTRQKDRIAKAYNKKVR-AKSFMLGDYVWKVVLPIN 2001
D+ +E + V T L ++ K ++ K++ + F GD V +
Sbjct: 1149 --FSDKTDENSQETIQVFQTVKEHLNTNNIKMKKYFDMKIQEIEEFQPGDLV------MV 1200
Query: 2002 KKDK-----RYRKWVPNWEGPFTVEKVLVNNAYSI 2031
K+ K + K P++ GPF V + N Y +
Sbjct: 1201 KRTKTGFLHKSNKLAPSFAGPFYVLQKSGPNNYEL 1235
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 190 bits (483), Expect = 3e-47
Identities = 131/422 (31%), Positives = 217/422 (51%), Gaps = 22/422 (5%)
Query: 1060 KIKEEIERLLRCKFIRTARYVD----WLANVVPVIKKNGKMRVCIDFRDLNAATPKDEYH 1115
+++ +I+ LL+ IR + W+ P + R+ +DF+ LN T D Y
Sbjct: 138 EVERQIDELLQDGIIRPSNSPYNSPIWIVPKKPKPNGEKQYRMVVDFKRLNTVTIPDTYP 197
Query: 1116 MPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLK 1175
+P + S +Y + LD SG++QI + E D+ KTAF G YE++ +PFGLK
Sbjct: 198 IPDINATLASLGNAKYFTTLDLTSGFHQIHMKESDIPKTAFSTLN--GKYEFLRLPFGLK 255
Query: 1176 NAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPL 1235
NA A +QR+++ I + I VYIDDI+V S D H +LR + K L++N
Sbjct: 256 NAPAIFQRMIDDILREHIGKVCYVYIDDIIVFSEDYDTHWKNLRLVLASLSKANLQVNLE 315
Query: 1236 KCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIANL 1295
K F +FLG++V GI+ + K +AI + PPTS K+L+ LG ++ R+FI +
Sbjct: 316 KSHFLDTQVEFLGYIVTADGIKADPKKVRAISEMPPPTSVKELKRFLGMTSYYRKFIQDY 375
Query: 1296 SEKTKSFSPLLR-----LKKEDAFR----WEAEHQKAFDELKVYLSSPPIMASPIRGKPM 1346
++ K + L R +K + + + ++F++LK L S I+A P KP
Sbjct: 376 AKVAKPLTNLTRGLYANIKSSQSSKVPITLDETALQSFNDLKSILCSSEILAFPCFTKPF 435
Query: 1347 KLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKY 1406
L A++ IG++L+Q+D+ ++R I Y+SR LN E Y IEK L + +S L+
Sbjct: 436 HLTTDASNWAIGAVLSQDDQ-GRDRPIAYISRSLNKTEENYATIEKEMLAIIWSLDNLRA 494
Query: 1407 YIKPI-DVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYS--LTYAPLKAVKGQAIADF 1463
Y+ + V++ + + L ++++ +W + EY+ L Y P K +AD
Sbjct: 495 YLYGAGTIKVYTDHQPLTFALGNRNFNAKLKRWKARIEEYNCELIYKP---GKSNVVADA 551
Query: 1464 LA 1465
L+
Sbjct: 552 LS 553
Score = 38.5 bits (88), Expect = 0.20
Identities = 35/172 (20%), Positives = 74/172 (42%), Gaps = 7/172 (4%)
Query: 1708 AHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFR 1767
AH+ +++ L + Y+P + CQ C+ + +H L +P P
Sbjct: 704 AHRGPTEIRLQLLEK-YYFPRMSSTIRLQTSSCQCCKLYKYERHPNKPNL----QPTPIP 758
Query: 1768 GWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPE 1827
+ +++ I+ + + Y+ ID F+K+ + LQ+ + E + ++ F P+
Sbjct: 759 NYPCEIL-HIDIFALEKRLYLS-CIDKFSKFAKLFHLQSKASVHLRETLVEALHYFTAPK 816
Query: 1828 SLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIK 1879
L +D + V ++ S I L + ++ NGQVE + T + + +
Sbjct: 817 VLVSDNERGLLCPTVLNYLRSLDIDLYYAPTQKSEVNGQVERFHSTFLEIYR 868
>POL2_DROME (P20825) Retrovirus-related Pol polyprotein from
transposon 297 [Contains: Protease (EC 3.4.23.-); Reverse
transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1059
Score = 189 bits (481), Expect = 5e-47
Identities = 124/413 (30%), Positives = 213/413 (51%), Gaps = 17/413 (4%)
Query: 1059 VKIKEEIERLLRCKFIRTARYV----DWLANVVPVIKKNGKMRVCIDFRDLNAATPKDEY 1114
++++ +++ +L IR + W+ P K RV ID+R LN T D Y
Sbjct: 220 IEVENQVQEMLNQGLIRESNSPYNSPTWVVPKKPDASGANKYRVVIDYRKLNEITIPDRY 279
Query: 1115 HMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGL 1174
+P + ++ +Y + +D G++QI + EE +SKTAF G YE++ MPFGL
Sbjct: 280 PIPNMDEILGKLGKCQYFTTIDLAKGFHQIEMDEESISKTAFSTKS--GHYEYLRMPFGL 337
Query: 1175 KNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNP 1234
+NA AT+QR MN I + VY+DDI++ S S +HL ++ F ++ LK+
Sbjct: 338 RNAPATFQRCMNNILRPLLNKHCLVYLDDIIIFSTSLTEHLNSIQLVFTKLADANLKLQL 397
Query: 1235 LKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIAN 1294
KC F +FLG +V GI+ N K KAI+ PT K++++ LG + R+FI N
Sbjct: 398 DKCEFLKKEANFLGHIVTPDGIKPNPIKVKAIVSYPIPTKDKEIRAFLGLTGYYRKFIPN 457
Query: 1295 LSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATD 1354
++ K + L+ K+ + E+ +AF++LK + PI+ P K L A++
Sbjct: 458 YADIAKPMTSCLK-KRTKIDTQKLEYIEAFEKLKALIIRDPILQLPDFEKKFVLTTDASN 516
Query: 1355 GTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVM 1414
+G++L+Q I ++SR LND E Y+ IEK L + ++ ++Y+ +
Sbjct: 517 LALGAVLSQNG-----HPISFISRTLNDHELNYSAIEKELLAIVWATKTFRHYLLGRQFL 571
Query: 1415 VFSHYDIIK--HMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADFLA 1465
+ S + ++ H L +P +++ +W + L+EY +K K ++AD L+
Sbjct: 572 IASDHQPLRWLHNLKEP--GAKLERWRVRLSEYQFKIDYIKG-KENSVADALS 621
>POL3_DROME (P04323) Retrovirus-related Pol polyprotein from
transposon 17.6 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1058
Score = 187 bits (476), Expect = 2e-46
Identities = 139/494 (28%), Positives = 234/494 (47%), Gaps = 27/494 (5%)
Query: 986 DGLEKRPTYISALIDPEL----------KDRMVKLLKEFKDCFAWDYDEMPGLSREVVEL 1035
D + ++P IS +++ +L K R+ LL+++ D + D++ ++ +
Sbjct: 142 DTMLRQPNKISPILESDLYRLEHLNNEEKQRLCALLQKYHDIQYHEGDKLTFTNQTKHTI 201
Query: 1036 KLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVD----WLANVVPVIK 1091
P+ + +V + +I+ +L IRT+ W+
Sbjct: 202 NTKHNLPLYSKYSYPQAYEQEV----ESQIQDMLNQGIIRTSNSPYNSPIWVVPKKQDAS 257
Query: 1092 KNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDV 1151
K R+ ID+R LN T D + +P + ++ Y + +D G++QI + E V
Sbjct: 258 GKQKFRIVIDYRKLNEITVGDRHPIPNMDEILGKLGRCNYFTTIDLAKGFHQIEMDPESV 317
Query: 1152 SKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSR 1211
SKTAF G YE++ MPFGLKNA AT+QR MN I + VY+DDI+V S S
Sbjct: 318 SKTAFSTKH--GHYEYLRMPFGLKNAPATFQRCMNDILRPLLNKHCLVYLDDIIVFSTSL 375
Query: 1212 DDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSP 1271
D+HL L FE++ K LK+ KC F FLG V+ GI+ N K +AI
Sbjct: 376 DEHLQSLGLVFEKLAKANLKLQLDKCEFLKQETTFLGHVLTPDGIKPNPEKIEAIQKYPI 435
Query: 1272 PTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYL 1331
PT K++++ LG + R+FI N ++ K + L+ K E+ AF +LK +
Sbjct: 436 PTKPKEIKAFLGLTGYYRKFIPNFADIAKPMTKCLK-KNMKIDTTNPEYDSAFKKLKYLI 494
Query: 1332 SSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIE 1391
S PI+ P K L A+D +G++L+Q+ + Y+SR LN+ E Y+ IE
Sbjct: 495 SEDPILKVPDFTKKFTLTTDASDVALGAVLSQDG-----HPLSYISRTLNEHEINYSTIE 549
Query: 1392 KLCLCLYFSCVKLKYYIKPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAP 1451
K L + ++ ++Y+ + S + + + +S++ +W + L+E+
Sbjct: 550 KELLAIVWATKTFRHYLLGRHFEISSDHQPLSWLYRMKDPNSKLTRWRVKLSEFDFDIKY 609
Query: 1452 LKAVKGQAIADFLA 1465
+K K +AD L+
Sbjct: 610 IKG-KENCVADALS 622
>POLY_DROME (P10401) Retrovirus-related Pol polyprotein from
transposon gypsy [Contains: Reverse transcriptase (EC
2.7.7.49); Endonuclease]
Length = 1035
Score = 156 bits (394), Expect = 6e-37
Identities = 115/426 (26%), Positives = 211/426 (48%), Gaps = 26/426 (6%)
Query: 1061 IKEEIERLLRCKFIRTARYVDWLANVVPVIKK------NGKMRVCIDFRDLNAATPKDEY 1114
+ E+++LL+ IR +R + + V KK N R+ IDFR LN T D Y
Sbjct: 197 VNNEVKQLLKDGIIRPSRS-PYNSPTWVVDKKGTDAFGNPNKRLVIDFRKLNEKTIPDRY 255
Query: 1115 HMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGL 1174
MP M++ + ++ + LD SGY+QI++AE D KT+F G G YE+ +PFGL
Sbjct: 256 PMPSIPMILANLGKAKFFTTLDLKSGYHQIYLAEHDREKTSFSVNG--GKYEFCRLPFGL 313
Query: 1175 KNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNP 1234
+NA + +QR ++ + + I VY+DD+++ S + DH+ H+ + + ++++
Sbjct: 314 RNASSIFQRALDDVLREQIGKICYVYVDDVIIFSENESDHVRHIDTVLKCLIDANMRVSQ 373
Query: 1235 LKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFIAN 1294
K F + ++LGF+V K G + + K KAI + P +++S LG ++ R FI +
Sbjct: 374 EKTRFFKESVEYLGFIVSKDGTKSDPEKVKAIQEYPEPDCVYKVRSFLGLASYYRVFIKD 433
Query: 1295 LSEKTKSFSPLLR---------LKKEDAFRWEAEHQKAFDELKVYLSSPP-IMASPIRGK 1344
+ + + +L+ + K+ + + AF L+ L+S I+ P K
Sbjct: 434 FAAIARPITDILKGENGSVSKHMSKKIPVEFNETQRNAFQRLRNILASEDVILKYPDFKK 493
Query: 1345 PMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKL 1404
P L A+ IG++L+QE R I +SR L E Y E+ L + ++ KL
Sbjct: 494 PFDLTTDASASGIGAVLSQEG-----RPITMISRTLKQPEQNYATNERELLAIVWALGKL 548
Query: 1405 KYYI-KPIDVMVFSHYDIIKHMLSKPILHSRIGKWALALTEYSLTYAPLKAVKGQAIADF 1463
+ ++ ++ +F+ + + ++ +++I +W + +++ K K +AD
Sbjct: 549 QNFLYGSREINIFTDHQPLTFAVADRNTNAKIKRWKSYIDQHNAKVF-YKPGKENFVADA 607
Query: 1464 LADQTL 1469
L+ Q L
Sbjct: 608 LSRQNL 613
Score = 42.0 bits (97), Expect = 0.018
Identities = 45/202 (22%), Positives = 84/202 (41%), Gaps = 14/202 (6%)
Query: 1679 KNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAK 1738
KNV +L +++ ++A H+ A Q IK + Y+P + E
Sbjct: 726 KNV---VLDITDKNEQIEIVTAEHNRAHRAAQENIKQ----VLRDYYFPKMGSLAKEVVA 778
Query: 1739 RCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKW 1798
C+ C + +H EL P + G + + S K + ID F+K+
Sbjct: 779 NCRVCTQAKYDRHPKKQELGETPIP-SYTGEMVHI-----DIFSTDRKLFLTCIDKFSKY 832
Query: 1799 VEAIPLQNVTQDTVIEFISSIVYRFGLPESLTTDQGTVFVGQKVASFAE-SWGIKLLTST 1857
P+ + T + + I+ F +++ D F + V S + S+GI ++ +
Sbjct: 833 AIVQPVVSRTIVDITAPLLQIINLFPNIKTVYCDNEPAFNSETVTSMLKNSFGIDIVNAP 892
Query: 1858 PYYAQANGQVEAANKTLISLIK 1879
P ++ +NGQVE + TL + +
Sbjct: 893 PLHSSSNGQVERFHSTLAEIAR 914
>POL_CAMVS (P03554) Enzymatic polyprotein [Contains: Aspartic protease
(EC 3.4.23.-); Endonuclease; Reverse transcriptase (EC
2.7.7.49)]
Length = 679
Score = 149 bits (376), Expect = 8e-35
Identities = 135/521 (25%), Positives = 242/521 (45%), Gaps = 40/521 (7%)
Query: 964 KPVVDFSKKMEAQDPLEEVD-LGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDY 1022
+PV + K+E +PLEE+ L +G +R + I + ++ +LL+ K C
Sbjct: 172 EPVNISTNKIE--NPLEEIAILSEG--RRLSEEKLFITQQRMQKIEELLE--KVCSENPL 225
Query: 1023 DEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDW 1082
D P +++ ++ + + + K +K P ++ P + ++I+ LL K I+ ++
Sbjct: 226 D--PNKTKQWMKASIKLSDPSKAIKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKS--- 280
Query: 1083 LANVVPVI-------KKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLL 1135
++ P K+ GK R+ ++++ +N AT D Y++P + ++ G + S
Sbjct: 281 -PHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATVGDAYNLPNKDELLTLIRGKKIFSSF 339
Query: 1136 DGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIET 1195
D SG+ Q+ + +E TAF CP G YEW V+PFGLK A + +QR M+ F F
Sbjct: 340 DCKSGFWQVLLDQESRPLTAFTCP--QGHYEWNVVPFGLKQAPSIFQRHMDEAFRVF-RK 396
Query: 1196 FMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKG 1255
F VY+DDI+V S + +DHLLH+ ++ ++G+ ++ K +FLG + +G
Sbjct: 397 FCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLFKKKINFLGLEI-DEG 455
Query: 1256 IEINKNKAKAILDTSPPT--SKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDA 1313
+ ++ P T KKQLQ LG + + +I L++ K +LK+
Sbjct: 456 THKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIRKPLQ--AKLKENVP 513
Query: 1314 FRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQ---EDEDSKE 1370
+RW E ++K L P + P+ + + + A+D G ML + + E
Sbjct: 514 WRWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKAIKINEGTNTE 573
Query: 1371 RAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMV---FSHYDIIKHMLS 1427
Y S AE Y +K L + + K Y+ P+ ++ +H+ ++
Sbjct: 574 LICRYASGSFKAAEKNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTDNTHFKSFVNLNY 633
Query: 1428 KPILHSRIG---KWALALTEYSLTYAPLKAVKGQAIADFLA 1465
K S++G +W L+ YS +K ADFL+
Sbjct: 634 KG--DSKLGRNIRWQAWLSHYSFDVEHIKGTDNH-FADFLS 671
>POL_CAMVC (P03555) Enzymatic polyprotein [Contains: Aspartic protease
(EC 3.4.23.-); Endonuclease; Reverse transcriptase (EC
2.7.7.49)]
Length = 679
Score = 147 bits (372), Expect = 2e-34
Identities = 134/521 (25%), Positives = 242/521 (45%), Gaps = 40/521 (7%)
Query: 964 KPVVDFSKKMEAQDPLEEVD-LGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDY 1022
+PV + K+E +PLEE+ L +G +R + I + ++ +LL+ K C
Sbjct: 172 EPVNISTNKIE--NPLEEIAILSEG--RRLSEEKLFITQQRMQKIEELLE--KVCSENPL 225
Query: 1023 DEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDW 1082
D P +++ ++ + + + K +K P ++ P + ++I+ LL K I+ ++
Sbjct: 226 D--PNKTKQWMKASIKLSDPSKAIKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKS--- 280
Query: 1083 LANVVPVI-------KKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLL 1135
++ P K+ GK R+ ++++ +N AT D Y++P + ++ G + S
Sbjct: 281 -PHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATIGDAYNLPNKDELLTLIRGKKIFSSF 339
Query: 1136 DGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIET 1195
D SG+ Q+ + +E TAF CP G YEW V+PFGLK A + +QR M+ F F
Sbjct: 340 DCKSGFWQVLLDQESRPLTAFTCP--QGHYEWNVVPFGLKQAPSIFQRHMDEAFRVF-RK 396
Query: 1196 FMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKG 1255
F VY+DDI+V S + +DHLLH+ ++ ++G+ ++ K +FLG + +G
Sbjct: 397 FCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLFKKKINFLGLEI-DEG 455
Query: 1256 IEINKNKAKAILDTSPPT--SKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDA 1313
+ ++ P T KKQLQ LG + + +I L++ K +LK+
Sbjct: 456 THKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIRKPLQ--AKLKENVP 513
Query: 1314 FRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQ---EDEDSKE 1370
++W E ++K L P + P+ + + + A+D G ML + + E
Sbjct: 514 WKWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKAIKINEGTNTE 573
Query: 1371 RAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMV---FSHYDIIKHMLS 1427
Y S AE Y +K L + + K Y+ P+ ++ +H+ ++
Sbjct: 574 LICRYASGSFKAAERNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTDNTHFKSFVNLNY 633
Query: 1428 KPILHSRIG---KWALALTEYSLTYAPLKAVKGQAIADFLA 1465
K S++G +W L+ YS +K ADFL+
Sbjct: 634 KG--DSKLGRNIRWQAWLSHYSFDVEHIKGTDNH-FADFLS 671
>POL_CAMVE (Q02964) Enzymatic polyprotein [Contains: Aspartic protease
(EC 3.4.23.-); Endonuclease; Reverse transcriptase (EC
2.7.7.49)]
Length = 679
Score = 146 bits (368), Expect = 7e-34
Identities = 133/521 (25%), Positives = 242/521 (45%), Gaps = 40/521 (7%)
Query: 964 KPVVDFSKKMEAQDPLEEVD-LGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDY 1022
+PV + K+E +PL+E+ L +G +R + I + ++ +LL+ K C
Sbjct: 172 EPVNISTNKIE--NPLKEIAILSEG--RRLSEEKLFITQQRMQKIEELLE--KVCSENPL 225
Query: 1023 DEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDW 1082
D P +++ ++ + + + K +K P ++ P + ++I+ LL K I+ ++
Sbjct: 226 D--PNKTKQWMKASIKLSDPSKAIKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKS--- 280
Query: 1083 LANVVPVI-------KKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLL 1135
++ P K+ GK R+ ++++ +N AT D Y++P + ++ G + S
Sbjct: 281 -PHMAPAFLVNNEAEKRRGKKRMVVNYKAMNKATIGDAYNLPNKDELLTLIRGKKIFSSF 339
Query: 1136 DGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIET 1195
D SG+ Q+ + +E TAF CP G YEW V+PFGLK A + +QR M+ F F
Sbjct: 340 DCKSGFWQVLLDQESRPLTAFTCP--QGHYEWNVVPFGLKQAPSIFQRHMDEAFRVF-RK 396
Query: 1196 FMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKG 1255
F VY+DDI+V S + +DHLLH+ ++ ++G+ ++ K +FLG + +G
Sbjct: 397 FCCVYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLFKKKINFLGLEI-DEG 455
Query: 1256 IEINKNKAKAILDTSPPT--SKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDA 1313
+ ++ P T KKQLQ LG + + +I L++ K +LK+
Sbjct: 456 THKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIRKPLQ--AKLKENVP 513
Query: 1314 FRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQ---EDEDSKE 1370
++W E ++K L P + P+ + + + A+D G ML + + E
Sbjct: 514 WKWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKAIKINEGTNTE 573
Query: 1371 RAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMV---FSHYDIIKHMLS 1427
Y S AE Y +K L + + K Y+ P+ ++ +H+ ++
Sbjct: 574 LICRYASGSFKAAERNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTDNTHFKSFVNLNY 633
Query: 1428 KPILHSRIG---KWALALTEYSLTYAPLKAVKGQAIADFLA 1465
K S++G +W L+ YS +K ADFL+
Sbjct: 634 KG--DSKLGRNIRWQAWLSHYSFDVEHIKGTDNH-FADFLS 671
>POL_CAMVN (Q00962) Enzymatic polyprotein [Contains: Aspartic protease
(EC 3.4.23.-); Endonuclease; Reverse transcriptase (EC
2.7.7.49)]
Length = 680
Score = 145 bits (367), Expect = 9e-34
Identities = 134/521 (25%), Positives = 240/521 (45%), Gaps = 40/521 (7%)
Query: 964 KPVVDFSKKMEAQDPLEEVD-LGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDY 1022
+PV + K+E +PLEE+ L +G +R + I + + +LL+ K C
Sbjct: 173 EPVNISTNKIE--NPLEEIAILSEG--RRLSEEKLFITQQRMQKTEELLE--KVCSENPL 226
Query: 1023 DEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDW 1082
D P +++ ++ + + + K +K P ++ P + ++I+ LL K I+ ++
Sbjct: 227 D--PNKTKQWMKASIKLSDPSKAIKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKS--- 281
Query: 1083 LANVVPVIKKN-------GKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLL 1135
++ P N G R+ ++++ +N AT D Y++P + ++ G + S
Sbjct: 282 -PHMAPAFLVNNEAENGRGNKRMVVNYKAMNKATVGDAYNLPNKDELLTLIRGKKIFSSF 340
Query: 1136 DGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIET 1195
D SG+ Q+ + +E TAF CP G YEW V+PFGLK A + +QR M+ F F
Sbjct: 341 DCKSGFWQVLLDQESRPLTAFTCP--QGHYEWNVVPFGLKQAPSIFQRHMDEAFRVF-RK 397
Query: 1196 FMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKG 1255
F VY+DDIVV S + +DHLLH+ ++ ++G+ ++ K +FLG + +G
Sbjct: 398 FCCVYVDDIVVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLFKKKINFLGLEI-DEG 456
Query: 1256 IEINKNKAKAILDTSPPT--SKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDA 1313
+ ++ P T KKQLQ LG + + +I NL++ + +LK+
Sbjct: 457 THKPQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPNLAQMRQPLQ--AKLKENVP 514
Query: 1314 FRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQ---EDEDSKE 1370
++W E ++K L P + P+ + + + A+D G ML + + E
Sbjct: 515 WKWTKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKAIKINEGTNTE 574
Query: 1371 RAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMV---FSHYDIIKHMLS 1427
Y S AE Y +K L + + K Y+ P+ ++ +H+ ++
Sbjct: 575 LICRYRSGSFKAAERNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTDNTHFKSFVNLNY 634
Query: 1428 KPILHSRIG---KWALALTEYSLTYAPLKAVKGQAIADFLA 1465
K S++G +W L+ YS +K ADFL+
Sbjct: 635 KG--DSKLGRNIRWQAWLSHYSFDVEHIKGTDNH-FADFLS 672
>POL_CAMVD (P03556) Enzymatic polyprotein [Contains: Aspartic protease
(EC 3.4.23.-); Endonuclease; Reverse transcriptase (EC
2.7.7.49)]
Length = 674
Score = 144 bits (363), Expect = 3e-33
Identities = 117/458 (25%), Positives = 211/458 (45%), Gaps = 31/458 (6%)
Query: 1026 PGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLAN 1085
P +++ ++ + + + K +K P ++ P + ++I+ LL K I+ ++ +
Sbjct: 222 PNKTKQWMKASIKLSDPSKAIKVKPMKYSPMDREEFDKQIKELLDLKVIKPSKS----PH 277
Query: 1086 VVPVI-------KKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGY 1138
+ P K+ GK R+ ++++ +N AT D Y+ P + ++ G + S D
Sbjct: 278 MAPAFLVNNEAEKRRGKKRMVVNYKAMNKATVGDAYNPPNKDELLTLIRGKKIFSSFDCK 337
Query: 1139 SGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQ 1198
SG+ Q+ + +E TAF CP G YEW V+PFGLK A + +QR M+ F F F
Sbjct: 338 SGFWQVLLDQESRPLTAFTCP--QGHYEWNVVPFGLKQAPSIFQRHMDEAFRVF-RKFCC 394
Query: 1199 VYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEI 1258
VY+DDI+V S + +DHLLH+ ++ ++G+ ++ K +FLG + +G
Sbjct: 395 VYVDDILVFSNNEEDHLLHVAMILQKCNQHGIILSKKKAQLFKKKINFLGLEI-DEGTHK 453
Query: 1259 NKNKAKAILDTSPPT--SKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRW 1316
+ ++ P T KKQLQ LG + + +I L++ K +LK+ ++W
Sbjct: 454 PQGHILEHINKFPDTLEDKKQLQRFLGILTYASDYIPKLAQIRKPLQ--AKLKENVPWKW 511
Query: 1317 EAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQ---EDEDSKERAI 1373
E ++K L P + P+ + + + A+D G ML + + E
Sbjct: 512 TKEDTLYMQKVKKNLQGFPPLHHPLPEEKLIIETDASDDYWGGMLKAIKINEGTNTELIC 571
Query: 1374 FYLSRVLNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMV---FSHYDIIKHMLSKPI 1430
Y S AE Y +K L + + K Y+ P+ ++ +H+ ++ K
Sbjct: 572 RYASGSFKAAEKNYHSNDKETLAVINTIKKFSIYLTPVHFLIRTDNTHFKSFVNLNYKG- 630
Query: 1431 LHSRIG---KWALALTEYSLTYAPLKAVKGQAIADFLA 1465
S++G +W L+ YS +K ADFL+
Sbjct: 631 -DSKLGRNIRWQAWLSHYSFDVEHIKGTDNH-FADFLS 666
>POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic protease
(EC 3.4.23.-); Endonuclease; Reverse transcriptase (EC
2.7.7.49)]
Length = 666
Score = 130 bits (327), Expect = 4e-29
Identities = 104/382 (27%), Positives = 170/382 (44%), Gaps = 17/382 (4%)
Query: 1091 KKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEED 1150
++ GK R+ ++++ +N AT D +++P + ++ G S D SG+ Q+ + EE
Sbjct: 288 RRRGKKRMVVNYKAINQATIGDSHNLPNMQELLTLLRGKSIFSSFDCKSGFWQVVLDEES 347
Query: 1151 VSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPS 1210
TAF CP G ++W V+PFGLK A + +QR M T + + F VY+DDI+V S S
Sbjct: 348 QKLTAFTCP--QGHFQWKVVPFGLKQAPSIFQRHMQTALNG-ADKFCMVYVDDIIVFSNS 404
Query: 1211 RDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTS 1270
DH H+ + + KYG+ ++ K +FLG + KG +N +
Sbjct: 405 ELDHYNHVYAVLKIVEKYGIILSKKKANLFKEKINFLGLEI-DKGTHCPQNHILENIHKF 463
Query: 1271 PP--TSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELK 1328
P KK LQ LG + + +I L+E K ++LKK+ + W ++K
Sbjct: 464 PDRLEDKKHLQRFLGVLTYAETYIPKLAEIRKPLQ--VKLKKDVTWNWTQSDSDYVKKIK 521
Query: 1329 VYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYT 1388
L S P + P + + A+D G +L D E Y S AE Y
Sbjct: 522 KNLGSFPKLYLPKPEDHLIIETDASDSFWGGVLKARALDGVELICRYSSGSFKQAEKNYH 581
Query: 1389 MIEKLCLCLYFSCVKLKYYIKPIDVMV------FSHYDIIKHMLSKPILHSRIGKWALAL 1442
+K L + K Y+ P+ V F+++ ++ L R+ +W
Sbjct: 582 SNDKELLAVKQVITKFSAYLTPVRFTVRTDNKNFTYF--LRINLKGDSKQGRLVRWQNWF 639
Query: 1443 TEYSLTYAPLKAVKGQAIADFL 1464
++Y L+ VK +AD L
Sbjct: 640 SKYQFDVEHLEGVK-NVLADCL 660
>POL_CERV (P05400) Enzymatic polyprotein [Contains: Aspartic protease
(EC 3.4.23.-); Endonuclease; Reverse transcriptase (EC
2.7.7.49)]
Length = 659
Score = 126 bits (316), Expect = 7e-28
Identities = 109/456 (23%), Positives = 195/456 (41%), Gaps = 21/456 (4%)
Query: 1026 PGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLAN 1085
P S++ + + + + K VK P + P + +I+ LL K I+ ++
Sbjct: 209 PEKSKQWMTATIELIDPKTVVKVKPMSYSPSDREEFDRQIKELLELKVIKPSKSTHMSPA 268
Query: 1086 VV---PVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYN 1142
+ ++ GK R+ ++++ +N AT D +++P + ++ G + S D SG
Sbjct: 269 FLVENEAERRRGKKRMVVNYKAMNKATKGDAHNLPNKDELLTLVRGKKIYSSFDCKSGLW 328
Query: 1143 QIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYID 1202
Q+ + +E TAF CP G Y+W V+PFGLK A + + + + + VY+D
Sbjct: 329 QVLLDKESQLLTAFTCP--QGHYQWNVVPFGLKQAPSIFPKTYANSHSNQYSKYCCVYVD 386
Query: 1203 DIVV-KSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKN 1261
DI+V + R +H +H+ R K G+ ++ K +FLG + +G +N
Sbjct: 387 DILVFSNTGRKEHYIHVLNILRRCEKLGIILSKKKAQLFKEKINFLGLEI-DQGTHCPQN 445
Query: 1262 KAKAILDTSPP--TSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAE 1319
+ P KKQLQ LG + + +I L+ K +LK++ + W
Sbjct: 446 HILEHIHKFPDRIEDKKQLQRFLGILTYASDYIPKLASIRKPLQS--KLKEDSTWTWNDT 503
Query: 1320 HQKAFDELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRV 1379
+ ++K L S P + P + + A++ G +L + +S E Y S
Sbjct: 504 DSQYMAKIKKNLKSFPKLYHPEPNDKLVIETDASEEFWGGIL-KAIHNSHEYICRYASGS 562
Query: 1380 LNDAETRYTMIEKLCLCLYFSCVKLKYYIKPIDVMV------FSHYDIIKHMLSKPILHS 1433
AE Y EK L + K Y+ P ++ F+H+ + L
Sbjct: 563 FKAAERNYHSNEKELLAVIRVIKKFSIYLTPSRFLIRTDNKNFTHF--VNINLKGDRKQG 620
Query: 1434 RIGKWALALTEYSLTYAPLKAVKGQAIADFLADQTL 1469
R+ +W + L++Y + K ADFL + TL
Sbjct: 621 RLVRWQMWLSQYDFDVEHIAGTK-NVFADFLQENTL 655
>POL_COYMV (P19199) Putative polyprotein [Contains: Coat protein;
Protease (EC 3.4.23.-); Reverse transcriptase (EC
2.7.7.49); Ribonuclease H (EC 3.1.26.4)]
Length = 1886
Score = 123 bits (308), Expect = 6e-27
Identities = 122/458 (26%), Positives = 201/458 (43%), Gaps = 42/458 (9%)
Query: 957 DGPLGFEKPVVDFSKKMEAQDPLEEVDLGDGLEK----RPTYIS--------ALIDPELK 1004
+G L EK ++ F K + + + + + +E+ Y++ + +D E
Sbjct: 1301 EGGLRIEKDIITFYKLVTSIETSRTTQVANSIEELELSEDEYLNIAASVETPSFLDQEFA 1360
Query: 1005 DRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPI-KEDKKPVKQLPRRFHPDVLVKIKE 1063
+ LLKE K+ + M ++ KL I D K + + + P +
Sbjct: 1361 RKNKDLLKEMKEMKYIGENPMEFWKNNKIKCKLNIINPDIKIMGRPIKHVTPGDEEAMTR 1420
Query: 1064 EIERLLRCKFIR--------TARYVDWLANVVPVI--KKNGKMRVCIDFRDLNAATPKDE 1113
+I LL+ K IR TA V + P+ +K GK R+ +++ LN T D+
Sbjct: 1421 QINLLLQMKVIRPSESKHRSTAFIVRSGTEIDPITGKEKKGKERMVFNYKLLNENTESDQ 1480
Query: 1114 YHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFG 1173
Y +P ++ + S D SG+ Q+ + EE V TAF L YEW+VMPFG
Sbjct: 1481 YSLPGINTIISKVGRSKIYSKFDLKSGFWQVAMEEESVPWTAFLAGNKL--YEWLVMPFG 1538
Query: 1174 LKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMN 1233
LKNA A +QR M+ +F E F+ VYIDDI+V S + + H HL + ++ GL ++
Sbjct: 1539 LKNAPAIFQRKMDNVFKG-TEKFIAVYIDDILVFSETAEQHSQHLYTMLQLCKENGLILS 1597
Query: 1234 PLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPP--TSKKQLQSLLGKINFLRRF 1291
P K G DFLG + I++ + I D S + + ++S LG +++ R +
Sbjct: 1598 PTKMKIGTPEIDFLGASLGCTKIKLQPHIISKICDFSDEKLATPEGMRSWLGILSYARNY 1657
Query: 1292 IANLSEKTKSFSPL-LRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKPMKLYI 1350
I ++ K PL ++ R E K ++K + + P + P + I
Sbjct: 1658 IQDIG---KLVQPLRQKMAPTGDKRMNPETWKMVRQIKEKVKNLPDLQLPPKD---SFII 1711
Query: 1351 SATDGTIGS-------MLAQEDEDSKERAIFYLSRVLN 1381
TDG + +++ D S ER Y S N
Sbjct: 1712 IETDGCMTGWGAVCKWKMSKHDPRSTERICAYASGSFN 1749
>YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II
Length = 1268
Score = 119 bits (298), Expect = 9e-26
Identities = 95/289 (32%), Positives = 147/289 (49%), Gaps = 16/289 (5%)
Query: 1008 VKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKPVKQLPRRFHPDVLVKIKEEIER 1067
V L +F + F D + ++E E + +E+ PV + R L ++ E+ R
Sbjct: 408 VMLKNDFPEVFK---DGLGLCTKEKAEFRT--EENAVPVFKRARPVPYGSLEAVETELNR 462
Query: 1068 LLRCKFIRTARYVDWLANVVPVIKKN-GKMRVCIDFR--DLNAATPKDEYH-MPIAEMMV 1123
L I Y W A +V + KK GK+RVC DF+ LNAA KDE+H +P +E +
Sbjct: 463 LQEMGVIVPITYAKWAAPIVVIKKKGTGKIRVCADFKCSGLNAAL-KDEFHPLPTSEDIF 521
Query: 1124 DSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQR 1183
G Y S +D Y Q+ + EE G ++++ M FGLK A A++Q+
Sbjct: 522 SRLKGTVY-SQIDLKDAYLQVELDEEAQKLAVINTHR--GIFKYLRMTFGLKPAPASFQK 578
Query: 1184 VMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIA 1243
+M+ + T + VY DDI++ + S ++H LR+ FER ++YG +++ KCAF
Sbjct: 579 IMDKMVSGL--TGVAVYWDDIIISASSIEEHEKILRELFERFKEYGFRVSAEKCAFAQKQ 636
Query: 1244 GDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKINFLRRFI 1292
FLGF V + G + K +AI PT +KQL S LG ++L R +
Sbjct: 637 VTFLGF-VDEHGRRPDSKKTEAIRSMKAPTDQKQLASFLGAADWLSRMM 684
Score = 87.4 bits (215), Expect = 4e-16
Identities = 64/245 (26%), Positives = 112/245 (45%), Gaps = 24/245 (9%)
Query: 1696 IAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPAS 1755
I + +H+G H ++MK R ++W + D + C +CQ ++ + V
Sbjct: 785 IVLKQLHEG----HPGIVQMKQKA-RSFVFWRGLDSDIENMVRHCNNCQENSKMPRVVP- 838
Query: 1756 ELHSIIKPWP-----FRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQD 1810
+ PWP ++ +D G +N C Y++V +D TK+ E ++++
Sbjct: 839 -----LNPWPVPEAPWKRIHIDFAGPLNGC------YLLVVVDAKTKYAEVKLTRSISAV 887
Query: 1811 TVIEFISSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAA 1870
T I+ + I G PE++ +D GT A +S GI+ TS YY ++NG E
Sbjct: 888 TTIDLLEEIFSIHGYPETIISDNGTQLTSHLFAQMCQSHGIEHKTSAVYYPRSNGAAERF 947
Query: 1871 NKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKES-TGATPFRLAYGQEAVLPVEVYL 1929
TL I K G N Q L + L +YRN+P + G+TP +G++ + + +
Sbjct: 948 VDTLKRGIAKIKGEGSVN-QQILNKFLISYRNTPHSALNGSTPAECHFGRKIRTTMSLLM 1006
Query: 1930 QSCRI 1934
+ R+
Sbjct: 1007 PTDRV 1011
>POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 886
Score = 116 bits (290), Expect = 7e-25
Identities = 86/332 (25%), Positives = 153/332 (45%), Gaps = 7/332 (2%)
Query: 1086 VVPVIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIF 1145
V PV K +G+ R+ +D+R++N P + ++ + +Y + LD +G+
Sbjct: 5 VYPVPKPDGRWRMVLDYREVNKTIPLTAAQNQHSAGILATIVRQKYKTTLDLANGFWAHP 64
Query: 1146 IAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIV 1205
I E TAF G Y W +P G N+ A + + + + +QVY+DDI
Sbjct: 65 ITPESYWLTAFTWQGK--QYCWTRLPQGFLNSPALFTADVVDLLKEIPN--VQVYVDDIY 120
Query: 1206 VKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKA 1265
+ +H+ L K F+ + + G ++ K G +FLGF + K+G +
Sbjct: 121 LSHDDPKEHVQQLEKVFQILLQAGYVVSLKKSEIGQKTVEFLGFNITKEGRGLTDTFKTK 180
Query: 1266 ILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFD 1325
+L+ +PP KQLQS+LG +NF R FI N +E + L+ K W E+ K +
Sbjct: 181 LLNITPPKDLKQLQSILGLLNFARNFIPNFAELVQPLYNLIASAKGKYIEWSEENTKQLN 240
Query: 1326 ELKVYLSSPPIMASPIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAET 1385
+ L++ + R +L I + +E K + I YL+ V + AE
Sbjct: 241 MVIEALNTASNLEE--RLPEQRLVIKVNTSPSAGYVRYYNETGK-KPIMYLNYVFSKAEL 297
Query: 1386 RYTMIEKLCLCLYFSCVKLKYYIKPIDVMVFS 1417
+++M+EKL ++ + +K +++V+S
Sbjct: 298 KFSMLEKLLTTMHKALIKAMDLAMGQEILVYS 329
Score = 84.0 bits (206), Expect = 4e-15
Identities = 55/216 (25%), Positives = 99/216 (45%), Gaps = 4/216 (1%)
Query: 1725 YWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQ 1784
+WP + KD ++ RCQ C L PF + +D IG + P S+
Sbjct: 635 WWPNMRKDVVKQLGRCQQCLITNASNKASGPILRPDRPQKPFDKFFIDYIGPLPP--SQG 692
Query: 1785 HKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPESLTTDQGTVFVGQKVAS 1844
+ Y++V +D T + P + + ++ ++ ++ +P+ + +DQG F A
Sbjct: 693 YLYVLVVVDGMTGFTWLYPTKAPSTSATVKSLN-VLTSIAIPKVIHSDQGAAFTSSTFAE 751
Query: 1845 FAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSP 1904
+A+ GI L STPY+ Q+ +VE N + L+ K + +P W+ L V A N+
Sbjct: 752 WAKERGIHLEFSTPYHPQSGSKVERKNSDIKRLLTKLLVGRPTKWYDLLPVVQLALNNTY 811
Query: 1905 KESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEI 1940
TP +L +G ++ P + + R+EE+
Sbjct: 812 SPVLKYTPHQLLFGIDSNTPF-ANQDTLDLTREEEL 846
>POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1157
Score = 113 bits (283), Expect = 5e-24
Identities = 96/374 (25%), Positives = 163/374 (42%), Gaps = 10/374 (2%)
Query: 1044 KPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFR 1103
+P KQ P +P I+ I LL+ + + V PV K +GK R+ +D+R
Sbjct: 177 RPQKQYP--INPKAKASIQTVINDLLKQGVLIQQNSI-MNTPVYPVPKPDGKWRMVLDYR 233
Query: 1104 DLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALG 1163
++N P + ++ S +Y + LD +G+ I E TAF G
Sbjct: 234 EVNKTIPLIAAQNQHSAGILSSIFRGKYKTTLDLSNGFWAHSITPESYWLTAFTWLGQ-- 291
Query: 1164 TYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFE 1223
Y W +P G N+ A + + + + +QVY+DDI + +HL L K F
Sbjct: 292 QYCWTRLPQGFLNSPALFTADVVDLLKEVPN--VQVYVDDIYISHDDPREHLEQLEKVFS 349
Query: 1224 RMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLG 1283
+ G ++ K +FLGF + K+G + + + +L+ +PP KQLQS+LG
Sbjct: 350 LLLNAGYVVSLKKSEIAQHEVEFLGFNITKEGRGLTETFKQKLLNITPPRDLKQLQSILG 409
Query: 1284 KINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRG 1343
+NF R FI N SE K ++ W ++ + + L+S + R
Sbjct: 410 LLNFARNFIPNFSELVKPLYNIIATANGKYITWTTDNSQQLQNIISMLNSAENLEE--RN 467
Query: 1344 KPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVK 1403
++L + + +E +K R I YL+ V AE ++T EKL ++ +K
Sbjct: 468 PEVRLIMKVNTSPSAGYIRFYNEFAK-RPIMYLNYVYTKAEVKFTNTEKLLTTIHKGLIK 526
Query: 1404 LKYYIKPIDVMVFS 1417
+++V+S
Sbjct: 527 ALDLGMGQEILVYS 540
Score = 90.1 bits (222), Expect = 6e-17
Identities = 79/333 (23%), Positives = 147/333 (43%), Gaps = 44/333 (13%)
Query: 1725 YWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPW----PFRGWALDLIGEINPC 1780
+WP + KD ++ ++C+ C + + I++P PF + +D IG + P
Sbjct: 845 WWPNLRKDVVKVIRQCKQCL----VTNAATLAAPPILRPERPVKPFDKFFIDYIGPLPPS 900
Query: 1781 SSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLPESLTTDQGTVFVGQ 1840
+ H ++V +D T +V P + + ++ ++ + +P+ + +DQG F
Sbjct: 901 NGYLH--VLVVVDSMTGFVWLYPTKAPSTSATVKALNMLT-SIAVPKVIHSDQGAAFTSA 957
Query: 1841 KVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAY 1900
A +A++ GI+L STPY+ Q++G+VE N + L+ K + +P W+ L V A
Sbjct: 958 TFADWAKNKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTKLLVGRPAKWYDLLPVVQLAL 1017
Query: 1901 RNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDEER 1960
NS S+ TP +L +G ++ P + + R+EE+ +L E+
Sbjct: 1018 NNSYSPSSKYTPHQLLFGIDSNTPF-ANSDTLDLSREEEL-------SLLQEI------- 1062
Query: 1961 LLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVW-KVVLPINKKDKRYRKWVPNWEGPFT 2019
R + +RA S +G V +V P + + P W P
Sbjct: 1063 --------RSSLYLPSTPPASIRAWSPSVGQLVQERVARPASLR--------PRWHKPTP 1106
Query: 2020 VEKVLVNNAYSIKE-LGGNRQMTINGKYLKTYK 2051
V +V+ A I + LG R ++++ L Y+
Sbjct: 1107 VLEVINPRAVVILDHLGNRRTVSVDNLKLTAYQ 1139
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.334 0.144 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 220,767,386
Number of Sequences: 164201
Number of extensions: 9035764
Number of successful extensions: 34000
Number of sequences better than 10.0: 145
Number of HSP's better than 10.0 without gapping: 86
Number of HSP's successfully gapped in prelim test: 60
Number of HSP's that attempted gapping in prelim test: 33460
Number of HSP's gapped (non-prelim): 361
length of query: 2064
length of database: 59,974,054
effective HSP length: 126
effective length of query: 1938
effective length of database: 39,284,728
effective search space: 76133802864
effective search space used: 76133802864
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0269b.2


