

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
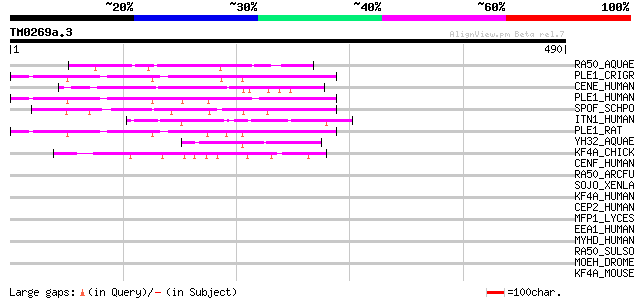
Query= TM0269a.3
(490 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
RA50_AQUAE (O67124) Probable DNA double-strand break repair rad5... 53 2e-06
PLE1_CRIGR (Q9JI55) Plectin 1 (PLTN) (PCN) (300-kDa intermediate... 50 1e-05
CENE_HUMAN (Q02224) Centromeric protein E (CENP-E protein) 49 2e-05
PLE1_HUMAN (Q15149) Plectin 1 (PLTN) (PCN) (Hemidesmosomal prote... 49 4e-05
SPOF_SCHPO (Q10411) Sporulation-specific protein 15 48 5e-05
ITN1_HUMAN (Q15811) Intersectin 1 (SH3 domain-containing protein... 46 2e-04
PLE1_RAT (P30427) Plectin 1 (PLTN) (PCN) 46 2e-04
YH32_AQUAE (O67622) Hypothetical UPF0144 protein AQ_1732 45 4e-04
KF4A_CHICK (Q90640) Chromosome-associated kinesin KIF4A (Chromok... 44 7e-04
CENF_HUMAN (P49454) CENP-F kinetochore protein (Centromere prote... 44 0.001
RA50_ARCFU (O29230) DNA double-strand break repair rad50 ATPase 44 0.001
SOJO_XENLA (Q9PW73) Cytoskeletal protein Sojo (p170) 43 0.002
KF4A_HUMAN (O95239) Chromosome-associated kinesin KIF4A (Chromok... 43 0.002
CEP2_HUMAN (Q9BV73) Centrosomal protein 2 (Centrosomal Nek2-asso... 43 0.002
MFP1_LYCES (P93203) MAR binding filament-like protein 1 43 0.002
EEA1_HUMAN (Q15075) Early endosome antigen 1 (Endosome-associate... 42 0.003
MYHD_HUMAN (Q9UKX3) Myosin heavy chain, skeletal muscle, extraoc... 42 0.005
RA50_SULSO (Q97WH0) DNA double-strand break repair rad50 ATPase 41 0.006
MOEH_DROME (P46150) Moesin/ezrin/radixin homolog 1 (Ezrin-moesin... 41 0.006
KF4A_MOUSE (P33174) Chromosome-associated kinesin KIF4A (Chromok... 41 0.006
>RA50_AQUAE (O67124) Probable DNA double-strand break repair rad50
ATPase
Length = 978
Score = 52.8 bits (125), Expect = 2e-06
Identities = 55/232 (23%), Positives = 115/232 (48%), Gaps = 28/232 (12%)
Query: 53 PSLKDVIEQEATNLSDQHKRIS-----VRDLACKFDKNLAAAAKLSNEA-KIREVASLEG 106
P+ K+V+E+ NL ++ K + +R K ++ + +LS K++E+ +LE
Sbjct: 199 PTKKEVLEKTLKNLEEELKELKETEEKLRQELKKAEEKDSLERELSQVVTKLKELENLEK 258
Query: 107 HVLLKKLRDALEYLR-----VRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVK 161
V +KLR+ LE+ R V A R E+++K ++ ++ KLT+ L E +
Sbjct: 259 EV--EKLREKLEFSRKVAPYVPIAKRI-EEIDKKLTELKVRKNKLTKELAVLKDELSFAQ 315
Query: 162 KLLNFLKQASEDAKKLVNQEKSF-----ACAEIESARSVVLRIGEALEEQEQDSQASKPQ 216
+ LN ++ E K+ +EK EI+ + ++ +L+E+E++ + +K Q
Sbjct: 316 EELNRIEAEKEKFKEEKEREKELEHRLKKLQEIKEILKELSQLSSSLKEKEREYEQAK-Q 374
Query: 217 DVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKEL 268
+ + L E V++ +++ V E +L +++ E+ S+K+++ L
Sbjct: 375 EFEDLSERVEKGKKL--------VAETEEKLEKIKELFSEEEYTSLKMKERL 418
Score = 41.6 bits (96), Expect = 0.005
Identities = 41/162 (25%), Positives = 82/162 (50%), Gaps = 16/162 (9%)
Query: 129 KEDVEKAISMVEALAVKLTQNEGELIQ-------EKFEVKKLLNFLKQASEDAKKLVNQE 181
KE++EK + VE L ++ +N E I+ EK +++ LN ++A ED +K +
Sbjct: 538 KEEMEKLRNEVEELRKEIPENLKERIKKLEELRIEKEKLEHKLNKYRKALEDRQK----Q 593
Query: 182 KSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVM 241
K A A++ A++ + + E + E+ + + K +E+ +E+ + ++ + SK+
Sbjct: 594 KEEAQAKLHKAQTELELLKEKIREKSRLVKEFKELYRVERLEDYEESLKEEINYINSKLQ 653
Query: 242 AMEYELRALRDQIREKSIFSIKLQKEL-----TMSKRDEENK 278
+E + + LR E S KL+ EL +++ +EE K
Sbjct: 654 EIEEKEKKLRKHFEELSSRKSKLEGELSALNESINSLEEERK 695
>PLE1_CRIGR (Q9JI55) Plectin 1 (PLTN) (PCN) (300-kDa intermediate
filament-associated protein) (IFAP300) (Fragment)
Length = 4473
Score = 50.1 bits (118), Expect = 1e-05
Identities = 70/320 (21%), Positives = 134/320 (41%), Gaps = 46/320 (14%)
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVED--------NQG 52
+ K+ E+E+ + +A + + S + K +L A++ E E+ +
Sbjct: 1683 LAKVRAEMEVLLASKAR---AEEESRSTSEKSKQRLEAEADRFRELAEEAARLRALAEEA 1739
Query: 53 PSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKK 112
+ + E++A + +R+ LA ++ A +L EA+I + L++
Sbjct: 1740 KRQRQLAEEDAARQRAEAERVLTEKLAA-----ISEATRLKTEAEIALKEKEAENERLRR 1794
Query: 113 LRDALEYLRVRF---AGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQ 169
L + + R R A +K D+E E LA +E EL ++K V+ L +Q
Sbjct: 1795 LAEDEAFQRRRLEEQAALHKADIE------ERLAQLRKASESELERQKGLVEDTLRQRRQ 1848
Query: 170 ASEDAKKL-VNQEKSFA--------CAEIESARSVVLRIGEALE-----------EQEQD 209
E+ L V+ EK+ A I S+ +R E E E+EQ
Sbjct: 1849 VEEEILALKVSFEKAAAGKAELELELGRIRSSAEDTMRSKEQAEQEAARQRQLAAEEEQR 1908
Query: 210 SQASKPQDVDGLVEEVQEARRIKL-LHQPSKVMAMEYELRALRDQIREKSIFSIKLQKEL 268
+ ++ + L E + AR+ K L + ++ A E R LR++ ++S ++L +E
Sbjct: 1909 RREAEERVQKSLAAEEEAARQRKAALEEVERLKAKVEEARRLRERAEQESARQLQLAQEA 1968
Query: 269 TMSKRDEENKSHPYMLHGSE 288
+ E K+H +++ E
Sbjct: 1969 AQKRLQAEEKAHAFVVQQRE 1988
Score = 34.7 bits (78), Expect = 0.57
Identities = 51/257 (19%), Positives = 107/257 (40%), Gaps = 21/257 (8%)
Query: 35 KLGADSQILEETVEDNQGPSLKDVIEQEATNLSDQ-----HKRISVRDLACKFDKNLAAA 89
+L A+ +++ E QG + ++E+E L + HKR + K +
Sbjct: 1634 RLAAEQELIRLRAETEQGEQQRQLLEEELARLQREATAATHKRQELEAELAKVRAEMEVL 1693
Query: 90 AKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQN 149
L+++A+ E E +K + LE RF E+ + ++ E +
Sbjct: 1694 --LASKARAEE----ESRSTSEKSKQRLEAEADRFR-ELAEEAARLRALAEEAKRQRQLA 1746
Query: 150 EGELIQEKFEVKKLLNFLKQASEDAKKLVNQ-EKSFACAEIESARSVVLRIGEALEEQEQ 208
E + +++ E +++L A +A +L + E + E E+ R L EA + +
Sbjct: 1747 EEDAARQRAEAERVLTEKLAAISEATRLKTEAEIALKEKEAENERLRRLAEDEAFQRRRL 1806
Query: 209 DSQAS-KPQDVDGLVEEVQEARRIKLLHQPSKV-------MAMEYELRALRDQIREKSIF 260
+ QA+ D++ + ++++A +L Q V +E E+ AL+ + +
Sbjct: 1807 EEQAALHKADIEERLAQLRKASESELERQKGLVEDTLRQRRQVEEEILALKVSFEKAAAG 1866
Query: 261 SIKLQKELTMSKRDEEN 277
+L+ EL + E+
Sbjct: 1867 KAELELELGRIRSSAED 1883
Score = 34.7 bits (78), Expect = 0.57
Identities = 55/305 (18%), Positives = 121/305 (39%), Gaps = 26/305 (8%)
Query: 127 RNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFAC 186
R +D + EA + + E QEK + L L+ +E+A++ + Q ++
Sbjct: 1427 RQVQDESQRKRQAEAELALRVKAQAEAAQEKQRALQALEELRLQAEEAERRLRQAQAERA 1486
Query: 187 AEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEE-------VQEARRIKLLHQPSK 239
+++ A R E + ++ S A K ++ ++E ++E + Q
Sbjct: 1487 RQVQVALETAQRSAEVELQSKRASFAEKTAQLERTLQEEHVTVTQLREKAERRAQQQAEA 1546
Query: 240 VMAMEYELRAL-RDQIREKSIFSIKLQKELTMSKRD----EENKSHPYMLHGSEALGSYL 294
A E R L R Q++ ++LQ E +R + K + G
Sbjct: 1547 ERAREEAERELERWQLKANEALRLRLQAEEVAQQRSLAQADAEKQKEEAEREARRRG--- 1603
Query: 295 KVQPRSGEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVG----RILQVDIV 350
K + ++ ++++ + R +EG+ ++ ++ + I + G ++L+ ++
Sbjct: 1604 KAEEQAVRQRELAEQELEKQRQLAEGTAQQRLAAEQELIRLRAETEQGEQQRQLLEEELA 1663
Query: 351 SNGKKLTLTTNPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHSFNVGRMR 410
++ T T+ Q +EA L K + V+++ + SRST + R+
Sbjct: 1664 RLQREATAATHKRQ-------ELEAELAKVRAEMEVLLASKARAEEESRSTSEKSKQRLE 1716
Query: 411 IKLCR 415
+ R
Sbjct: 1717 AEADR 1721
Score = 33.5 bits (75), Expect = 1.3
Identities = 58/260 (22%), Positives = 107/260 (40%), Gaps = 45/260 (17%)
Query: 33 KYKLGADSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKL 92
K K AD+++ + Q K +EQE T L Q + K++
Sbjct: 2084 KQKQAADAEMEKHKKFAEQTLRQKAQVEQELTTLRLQLEETD-------HQKSILDEELQ 2136
Query: 93 SNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGE 152
+A++ E A V + L +RV+ E++ K + +EA +N
Sbjct: 2137 RLKAEVTEAARQRSQV-----EEELFSVRVQM-----EELGKLKARIEA------ENRAL 2180
Query: 153 LIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQA 212
++++K ++ FL++ +E K++ + + A E+AR + + EE +A
Sbjct: 2181 ILRDKDNTQR---FLEEEAEKMKQVAEEAARLSVAAQEAAR-----LRQLAEEDLAQQRA 2232
Query: 213 SKPQDVDGLVEEVQEARRIKL---LHQPSKVMAMEYELRALRD------QIREKS----- 258
+ + ++ VQEA R+K L Q K +A E R D Q+ E++
Sbjct: 2233 LAEKMLKEKMQAVQEATRLKAEAELLQQQKELAQEQARRLQEDKEQMAQQLVEETQGFQR 2292
Query: 259 IFSIKLQKELTMSKRDEENK 278
++ Q++L MS E K
Sbjct: 2293 TLEVERQRQLEMSAEAERLK 2312
Score = 30.8 bits (68), Expect = 8.3
Identities = 38/196 (19%), Positives = 80/196 (40%), Gaps = 9/196 (4%)
Query: 69 QHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRN 128
Q +++ LA K AAA K E ++ + S + K + E R R
Sbjct: 1845 QRRQVEEEILALKVSFEKAAAGKAELELELGRIRSSAEDTMRSKEQAEQEAARQRQLAAE 1904
Query: 129 KED--------VEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQ 180
+E V+K+++ E A + E+ + K +V++ ++A +++ + +
Sbjct: 1905 EEQRRREAEERVQKSLAAEEEAARQRKAALEEVERLKAKVEEARRLRERAEQESARQLQL 1964
Query: 181 EKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKV 240
+ A +++ + + EE+ Q + + ++ L E + ARR + ++
Sbjct: 1965 AQEAAQKRLQAEEKAHAFVVQQREEELQQTLQQEQSMLERLRGEAEAARRAAEEAEEARE 2024
Query: 241 MAMEYELRALRDQIRE 256
A E E R Q+ E
Sbjct: 2025 QA-EREAAQSRKQVEE 2039
>CENE_HUMAN (Q02224) Centromeric protein E (CENP-E protein)
Length = 2663
Score = 49.3 bits (116), Expect = 2e-05
Identities = 57/250 (22%), Positives = 121/250 (47%), Gaps = 27/250 (10%)
Query: 44 EETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVAS 103
+ET++ L+ ++ ++ +S+ K DL D A K+ E +I +
Sbjct: 1734 QETID-----KLRGIVSEKTNEISNMQK-----DLEHSNDALKAQDLKIQEELRIAHMHL 1783
Query: 104 LEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKL 163
E + KLR + + + K D+E + + ++ +L NE +LI K +V +
Sbjct: 1784 KEQQETIDKLRGIVSEKTDKLSNMQK-DLENSNAKLQEKIQELKANEHQLITLKKDVNET 1842
Query: 164 LNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEE-----QEQDS----QASK 214
+ + + K++ +Q + + EIE+ ++ + E LEE +E+D+ + +
Sbjct: 1843 QKKVSEMEQLKKQIKDQSLTLSKLEIENL-NLAQELHENLEEMKSVMKERDNLRRVEETL 1901
Query: 215 PQDVDGLVEEVQE--ARRIKLLHQ--PSKVMAMEYE--LRALRDQIREKSIFSIKLQKEL 268
+ D L E +QE AR +++ + +++++ E++ + LR++I EK+I +QK+L
Sbjct: 1902 KLERDQLKESLQETKARDLEIQQELKTARMLSKEHKETVDKLREKISEKTIQISDIQKDL 1961
Query: 269 TMSKRDEENK 278
SK + + K
Sbjct: 1962 DKSKDELQKK 1971
>PLE1_HUMAN (Q15149) Plectin 1 (PLTN) (PCN) (Hemidesmosomal protein 1)
(HD1)
Length = 4684
Score = 48.5 bits (114), Expect = 4e-05
Identities = 64/319 (20%), Positives = 138/319 (43%), Gaps = 44/319 (13%)
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVED--------NQG 52
+ K+ E+E+ + +A + + S + K +L A++ E E+ +
Sbjct: 1894 LAKVRAEMEVLLASKAK---AEEESRSTSEKSKQRLEAEAGRFRELAEEAARLRALAEEA 1950
Query: 53 PSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKK 112
+ + E++A + +R+ LA + A +L EA+I + L++
Sbjct: 1951 KRQRQLAEEDAARQRAEAERVLAEKLAA-----IGEATRLKTEAEIALKEKEAENERLRR 2005
Query: 113 LRDALEYLRVRF---AGRNKEDVEKAISMV-EALAVKLTQNEG---ELIQEKFEVKKLLN 165
L + + R R A ++K D+E+ ++ + +A +L + +G + ++++ +V++ +
Sbjct: 2006 LAEDEAFQRRRLEEQAAQHKADIEERLAQLRKASDSELERQKGLVEDTLRQRRQVEEEIL 2065
Query: 166 FLKQASEDA---------------KKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDS 210
LK + E A + +S AE+E+AR L E +E +
Sbjct: 2066 ALKASFEKAAAGKAELELELGRIRSNAEDTLRSKEQAELEAARQRQLAAEEERRRREAEE 2125
Query: 211 QASKPQDVDGLVEEVQEARRIKL-LHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELT 269
+ K L E + AR+ K L + ++ A E R+LR++ ++S ++L +E
Sbjct: 2126 RVQK-----SLAAEEEAARQRKAALEEVERLKAKVEEARSLRERAEQESARQLQLAQEAA 2180
Query: 270 MSKRDEENKSHPYMLHGSE 288
+ E K+H + + E
Sbjct: 2181 QKRLQAEEKAHAFAVQQKE 2199
Score = 36.6 bits (83), Expect = 0.15
Identities = 55/259 (21%), Positives = 112/259 (43%), Gaps = 25/259 (9%)
Query: 35 KLGADSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSN 94
+L A+ +++ E QG + ++E+E L R+ A K A+L+
Sbjct: 1845 RLAAEQELIRLRAETEQGEQQRQLLEEELARLQ--------REAAAATQKRQELEAELAK 1896
Query: 95 EAKIREVASLEGHVLLKKLRDALEYLRVRF---AGRNKEDVEKAISMVEALAVKLTQN-- 149
EV ++ R E + R AGR +E E+A + + ALA + +
Sbjct: 1897 VRAEMEVLLASKAKAEEESRSTSEKSKQRLEAEAGRFRELAEEA-ARLRALAEEAKRQRQ 1955
Query: 150 --EGELIQEKFEVKKLLNFLKQASEDAKKLVNQ-EKSFACAEIESARSVVLRIGEALEEQ 206
E + +++ E +++L A +A +L + E + E E+ R L EA + +
Sbjct: 1956 LAEEDAARQRAEAERVLAEKLAAIGEATRLKTEAEIALKEKEAENERLRRLAEDEAFQRR 2015
Query: 207 EQDSQASKPQ-DVDGLVEEVQEARRIKLLHQPSKV-------MAMEYELRALRDQIREKS 258
+ QA++ + D++ + ++++A +L Q V +E E+ AL+ + +
Sbjct: 2016 RLEEQAAQHKADIEERLAQLRKASDSELERQKGLVEDTLRQRRQVEEEILALKASFEKAA 2075
Query: 259 IFSIKLQKELTMSKRDEEN 277
+L+ EL + + E+
Sbjct: 2076 AGKAELELELGRIRSNAED 2094
Score = 32.3 bits (72), Expect = 2.8
Identities = 61/348 (17%), Positives = 133/348 (37%), Gaps = 33/348 (9%)
Query: 91 KLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAIS-------MVEALA 143
+L EA R+ EG + + R + R A E + + + E
Sbjct: 1595 RLQLEATERQRGGAEGELQALRARAEEAEAQKRQAQEEAERLRRQVQDESQRKRQAEVEL 1654
Query: 144 VKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEAL 203
+ E E +EK + L L+ +E+A++ + Q + +++ A R EA
Sbjct: 1655 ASRVKAEAEAAREKQRALQALEELRLQAEEAERWLCQAEVERARQVQVALETAQRSAEAE 1714
Query: 204 EEQEQDSQASKPQDVDGLVEEV--------QEARRIKLLHQPSKVMAMEYELRALRDQIR 255
+ ++ S A K ++ ++E +EA R ++ E E + R Q++
Sbjct: 1715 LQSKRASFAEKTAQLERSLQEEHVAVAQLREEAERRAQQQAEAERAREEAERQLERWQLK 1774
Query: 256 EKSIFSIKLQKELTMSKRD----EENKSHPYMLHGSEALGSYLKVQPRSGEVPQVSKCSF 311
++LQ E + ++ E K + G K + ++ ++++
Sbjct: 1775 ANEALRLRLQAEEVLQQKSLAQAEAEKQKEEAEREARRRG---KAEEQAVRQRELAEQEL 1831
Query: 312 QWYRLSSEGSWREVISGADKSIYAPDPFDVG----RILQVDIVSNGKKLTLTTNPIQTAS 367
+ R +EG+ ++ ++ + I + G ++L+ ++ ++ T Q
Sbjct: 1832 EKQRQLAEGTAQQRLAAEQELIRLRAETEQGEQQRQLLEEELARLQREAAAATQKRQ--- 1888
Query: 368 GLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHSFNVGRMRIKLCR 415
+EA L K + V+++ + SRST + R+ + R
Sbjct: 1889 ----ELEAELAKVRAEMEVLLASKAKAEEESRSTSEKSKQRLEAEAGR 1932
Score = 32.0 bits (71), Expect = 3.7
Identities = 62/309 (20%), Positives = 123/309 (39%), Gaps = 25/309 (8%)
Query: 123 RFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEK 182
R A + + + + ++ VEA K Q Q K + ++ L+Q ++ ++V +E+
Sbjct: 1486 RLAEQQRAEERERLAEVEAALEKQRQLAEAHAQAKAQAEREAKELQQRIQE--EVVRREE 1543
Query: 183 SFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDG-----LVEEVQEARRIKLLHQP 237
+ A+ + RS+ + + + E + QA Q +EE R++L
Sbjct: 1544 AAVDAQ-QQKRSIQEELQQLRQSSEAEIQAKARQAEAAERSRLRIEEEIRVVRLQLEATE 1602
Query: 238 SKVMAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKSHPYMLHGSEALGSYLKVQ 297
+ E EL+ALR + E + Q+E +R +++S L S +K +
Sbjct: 1603 RQRGGAEGELQALRARAEEAEAQKRQAQEEAERLRRQVQDESQ-RKRQAEVELASRVKAE 1661
Query: 298 PRSGEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVGRILQVDIVSNGKKLT 357
+ Q + + + RL +E + R + +V R QV + + +
Sbjct: 1662 AEAAREKQRALQALEELRLQAEEAERWLCQA-----------EVERARQVQVALETAQRS 1710
Query: 358 LTTNPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHSFNVGRMRIKLCR-- 415
+Q+ + A L +S + HV ++Q+ ++ R+ R R + R
Sbjct: 1711 AEAE-LQSKRASFAEKTAQLERSLQEEHVAVAQLR-EEAERRAQQQAEAERAREEAERQL 1768
Query: 416 -GWITKARE 423
W KA E
Sbjct: 1769 ERWQLKANE 1777
Score = 32.0 bits (71), Expect = 3.7
Identities = 61/304 (20%), Positives = 117/304 (38%), Gaps = 63/304 (20%)
Query: 43 LEETVEDNQGPSLKDVIEQ---EATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIR 99
LEET D+Q L + +++ EAT + Q ++ + + + K EA+ R
Sbjct: 2332 LEET--DHQKNLLDEELQRLKAEATEAARQRSQVEEELFSVRVQMEELSKLKARIEAENR 2389
Query: 100 EVASLEGHVLLKKLRDALEYLR------------VRFAGRNKEDVEKAISMVEALAVKLT 147
+ + + L++ E ++ + A R ++ E+ ++ ALA K+
Sbjct: 2390 ALILRDKDNTQRFLQEEAEKMKQVAEEAARLSVAAQEAARLRQLAEEDLAQQRALAEKML 2449
Query: 148 QNEGELIQEKFEVKKLLNFLKQ----ASEDAKKLVNQEKSFA------------------ 185
+ + + +QE +K L+Q A E A++L ++ A
Sbjct: 2450 KEKMQAVQEATRLKAEAELLQQQKELAQEQARRLQEDKEQMAQQLAEETQGFQRTLEAER 2509
Query: 186 ------CAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEV-QEARRIKLLHQPS 238
AE E + V + A E+D+Q + Q EE+ ++ R +L Q
Sbjct: 2510 QRQLEMSAEAERLKLRVAEMSRAQARAEEDAQRFRKQ-----AEEIGEKLHRTELATQEK 2564
Query: 239 KVMAMEYELR---------ALRDQIREKSIFSIKLQKE---LTMSKRDEENKSHPYMLHG 286
+ E++ LR+ I E KLQ+E L + + + +L
Sbjct: 2565 VTLVQTLEIQRQQSDHDAERLREAIAELEREKEKLQQEAKLLQLKSEEMQTVQQEQLLQE 2624
Query: 287 SEAL 290
++AL
Sbjct: 2625 TQAL 2628
>SPOF_SCHPO (Q10411) Sporulation-specific protein 15
Length = 1957
Score = 48.1 bits (113), Expect = 5e-05
Identities = 64/286 (22%), Positives = 128/286 (44%), Gaps = 29/286 (10%)
Query: 20 VSADVSFISDRFPKYKLGADSQILEETVE----DNQGPSLKDVIEQEATNLSDQ--HKRI 73
VS++ + + R + L A+ + L+E +E + + L + ++ N SD HK
Sbjct: 260 VSSNKTVSTLRQTENSLRAECKTLQEKLEKCAINEEDSKLLEELKHNVANYSDAIVHKDK 319
Query: 74 SVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVE 133
+ DL+ + ++ N R+ S++ L K LR+ + L+ N + E
Sbjct: 320 LIEDLSTRI-------SEFDNLKSERDTLSIKNEKLEKLLRNTIGSLKDSRTS-NSQLEE 371
Query: 134 KAISMVEA---LAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAK---KLVNQEKSFACA 187
+ + + E+ + +LT E +L + E K L + + + K+V Q S
Sbjct: 372 EMVELKESNRTIHSQLTDAESKLSSFEQENKSLKGSIDEYQNNLSSKDKMVKQVSS---- 427
Query: 188 EIESARSVVLRIGEALEE--QEQDSQASKPQDVDGLVEEVQ---EARRIKLLHQPSKVMA 242
++E ARS + L E E+D Q K +D + + ++++ + +L + + +
Sbjct: 428 QLEEARSSLAHATGKLAEINSERDFQNKKIKDFEKIEQDLRACLNSSSNELKEKSALIDK 487
Query: 243 MEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKSHPYMLHGSE 288
+ EL LR+QI+E+ S Q L +RD N+ + ++ S+
Sbjct: 488 KDQELNNLREQIKEQKKVSESTQSSLQSLQRDILNEKKKHEVYESQ 533
Score = 32.0 bits (71), Expect = 3.7
Identities = 28/137 (20%), Positives = 64/137 (46%), Gaps = 14/137 (10%)
Query: 145 KLTQNEGELIQEKFEVKKLLN----FLKQASEDAKKLVNQEKSFACAEIESARSVVLRIG 200
+LT + E + +++KL + +L Q + K L + EK+F E E ++
Sbjct: 1310 RLTTTDAEFTKVVADLEKLQHEHDDWLIQRGDLEKALKDSEKNFLRKEAEMTENI----- 1364
Query: 201 EALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIF 260
+LEE +++++ + L + ++K +++ + E+R D ++EK
Sbjct: 1365 HSLEEGKEETKKEIAELSSRLEDNQLATNKLK-----NQLDHLNQEIRLKEDVLKEKESL 1419
Query: 261 SIKLQKELTMSKRDEEN 277
I L++ L+ ++ E +
Sbjct: 1420 IISLEESLSNQRQKESS 1436
>ITN1_HUMAN (Q15811) Intersectin 1 (SH3 domain-containing protein
1A) (SH3P17)
Length = 1721
Score = 46.2 bits (108), Expect = 2e-04
Identities = 57/214 (26%), Positives = 95/214 (43%), Gaps = 26/214 (12%)
Query: 104 LEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNE---GELIQEKFEV 160
LEG L+ +R L R NK E I+ + L +L +++ G LI EK +
Sbjct: 492 LEGK--LQDIRCRLTTQRQEIESTNKSR-ELRIAEITHLQQQLQESQQMLGRLIPEKQIL 548
Query: 161 KKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDG 220
L ++Q S LV +++ E+ AR + + L+E E++++ SK Q++D
Sbjct: 549 NDQLKQVQQNSLHRDSLVTLKRALEAKEL--ARQ---HLRDQLDEVEKETR-SKLQEIDI 602
Query: 221 LVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENK-- 278
+++E R I Q K +ME E L+ + +E+ I ++ QKE + E +K
Sbjct: 603 FNNQLKELREIHNKQQLQKQKSMEAE--RLKQKEQERKIIELEKQKEEAQRRAQERDKQW 660
Query: 279 ----------SHPYMLHGSEALGSYLKVQPRSGE 302
P LH E L V+ + GE
Sbjct: 661 LEHVQQEDEHQRPRKLHEEEKLKREESVKKKDGE 694
>PLE1_RAT (P30427) Plectin 1 (PLTN) (PCN)
Length = 4687
Score = 45.8 bits (107), Expect = 2e-04
Identities = 69/320 (21%), Positives = 136/320 (41%), Gaps = 46/320 (14%)
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVED--------NQG 52
+ K+ E+E+ + +A + + S + K +L A++ E E+ +
Sbjct: 1897 LAKVRAEMEVLLASKAR---AEEESRSTSEKSKQRLEAEAGRFRELAEEAARLRALAEEA 1953
Query: 53 PSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKK 112
+++ E++A + + LA ++ A +L EA+I + L++
Sbjct: 1954 RRHRELAEEDAARQRAEADGVLTEKLAA-----ISEATRLKTEAEIALKEKEAENERLRR 2008
Query: 113 LRDALEYLRVRF---AGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQ 169
L + + R R A ++K D+E E LA +E EL ++K V+ L +Q
Sbjct: 2009 LAEDEAFQRRRLEEQAAQHKADIE------ERLAQLRKASESELERQKGLVEDTLRQRRQ 2062
Query: 170 ASED--AKKLVNQEKSFACAEIE------------SARSVVLRIGEALE------EQEQD 209
E+ A K ++ + AE+E + RS L EA E+EQ
Sbjct: 2063 VEEEIMALKASFEKAAAGKAELELELGRIRSNAEDTMRSKELAEQEAARQRQLAAEEEQR 2122
Query: 210 SQASKPQDVDGLVEEVQEARRIKL-LHQPSKVMAMEYELRALRDQIREKSIFSIKLQKEL 268
+ ++ + L E + AR+ K+ L + ++ A E R LR++ ++S ++L +E
Sbjct: 2123 RREAEERVQRSLAAEEEAARQRKVALEEVERLKAKVEEARRLRERAEQESARQLQLAQEA 2182
Query: 269 TMSKRDEENKSHPYMLHGSE 288
+ E K+H +++ E
Sbjct: 2183 AQKRLQAEEKAHAFVVQQRE 2202
Score = 38.1 bits (87), Expect = 0.052
Identities = 58/269 (21%), Positives = 117/269 (42%), Gaps = 21/269 (7%)
Query: 35 KLGADSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLA-AAAKLS 93
+L A+ +++ E QG + ++E+E L QH+ + + + LA A++
Sbjct: 1848 RLAAEQELIRLRAETEQGEHQRQLLEEELARL--QHEATAATQKRQELEAELAKVRAEME 1905
Query: 94 NEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQN---- 149
+ A E +K + LE AGR +E E+A + + ALA + ++
Sbjct: 1906 VLLASKARAEEESRSTSEKSKQRLE----AEAGRFRELAEEA-ARLRALAEEARRHRELA 1960
Query: 150 EGELIQEKFEVKKLLNFLKQASEDAKKLVNQ-EKSFACAEIESARSVVLRIGEALEEQEQ 208
E + +++ E +L A +A +L + E + E E+ R L EA + +
Sbjct: 1961 EEDAARQRAEADGVLTEKLAAISEATRLKTEAEIALKEKEAENERLRRLAEDEAFQRRRL 2020
Query: 209 DSQASKPQ-DVDGLVEEVQEARRIKLLHQPSKV-------MAMEYELRALRDQIREKSIF 260
+ QA++ + D++ + ++++A +L Q V +E E+ AL+ + +
Sbjct: 2021 EEQAAQHKADIEERLAQLRKASESELERQKGLVEDTLRQRRQVEEEIMALKASFEKAAAG 2080
Query: 261 SIKLQKELTMSKRDEENKSHPYMLHGSEA 289
+L+ EL + + E+ L EA
Sbjct: 2081 KAELELELGRIRSNAEDTMRSKELAEQEA 2109
Score = 38.1 bits (87), Expect = 0.052
Identities = 57/280 (20%), Positives = 119/280 (42%), Gaps = 36/280 (12%)
Query: 35 KLGADS-QILEETVEDNQG-PSLKDVIEQEATNLSDQHKRISVR------------DLAC 80
+L AD Q+ ++ VE+ QG + Q +S + +R+ +R + A
Sbjct: 2485 RLQADKEQMAQQLVEETQGFQRTLEAERQRQLEMSAEAERLKLRMAEMSRAQARAEEDAQ 2544
Query: 81 KFDKNLAAAAK------LSNEAKIREVASLE-----GHVLLKKLRDALEYLRVRFAGRNK 129
+F K + L+ + K+ V +LE ++LR+A+ L R K
Sbjct: 2545 RFRKQAEEIGEKLHRTELATQEKVTLVQTLEIQRQQSDQDAERLREAIAELE-----REK 2599
Query: 130 EDVEKAISMVEALAVKL-TQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAE 188
E +++ +++ + ++ T + +++QE ++K K + ++ + QEK A+
Sbjct: 2600 EKLKQEAKLLQLKSEEMQTVQQEQILQETQALQKSFLSEKDSLLQRERFIEQEK----AK 2655
Query: 189 IESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELR 248
+E + + L+E++Q Q Q+ LV ++EARR + V + EL+
Sbjct: 2656 LEQLFQDEVAKAKQLQEEQQRQQQQMEQEKQELVASMEEARR-RQREAEEGVRRKQEELQ 2714
Query: 249 ALRDQIREKSIFSIKLQKELTMSKRDEENKSHPYMLHGSE 288
L Q +++ + + L + E + + H E
Sbjct: 2715 RLEQQRQQQEKLLAEENQRLRERLQRLEEEHRAALAHSEE 2754
Score = 33.9 bits (76), Expect = 0.98
Identities = 53/302 (17%), Positives = 118/302 (38%), Gaps = 20/302 (6%)
Query: 127 RNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFAC 186
R +D + EA + E E +EK + L+ LK +E+A++ + Q ++
Sbjct: 1641 RQVQDESQRKRQAEAELALRVKAEAEAAREKQRALQALDELKLQAEEAERWLCQAEAERA 1700
Query: 187 AEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEV--------QEARRIKLLHQPS 238
+++ A R E + ++ S A K ++ ++E +EA R +
Sbjct: 1701 RQVQVALETAQRSAEVELQSKRPSFAEKTAQLERTLQEEHVTVTQLREEAERRAQQQAEA 1760
Query: 239 KVMAMEYELRALRDQIREKSIFSIKLQ-KELTMSKRDEENKSHPYMLHGSEALGSYLKVQ 297
+ E E R Q++ ++LQ +E+ K + + K +
Sbjct: 1761 ERAREEAERELERWQLKANEALRLRLQAEEVAQQKSLAQADAEKQKEEAEREARRRGKAE 1820
Query: 298 PRSGEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVG----RILQVDIVSNG 353
++ ++++ + R +EG+ ++ ++ + I + G ++L+ ++
Sbjct: 1821 EQAVRQRELAEQELEKQRQLTEGTAQQRLAAEQELIRLRAETEQGEHQRQLLEEELARLQ 1880
Query: 354 KKLTLTTNPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHSFNVGRMRIKL 413
+ T T Q +EA L K + V+++ + SRST + R+ +
Sbjct: 1881 HEATAATQKRQ-------ELEAELAKVRAEMEVLLASKARAEEESRSTSEKSKQRLEAEA 1933
Query: 414 CR 415
R
Sbjct: 1934 GR 1935
Score = 33.5 bits (75), Expect = 1.3
Identities = 61/287 (21%), Positives = 116/287 (40%), Gaps = 57/287 (19%)
Query: 33 KYKLGADSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKL 92
+ K A+ Q + L+ EQEA +R A K + AA A++
Sbjct: 2256 RLKQSAEEQAQAQAQAQAAAEKLRKEAEQEAA------RRAQAEQAALK--QKQAADAEM 2307
Query: 93 SNEAK-----IREVASLEGHVLLKKLR------------DALEYLR--VRFAGRNKEDVE 133
K +R+ A +E + +L+ + L+ L+ V A R + VE
Sbjct: 2308 EKHKKFAEQTLRQKAQVEQELTTLRLQLEETDHQKSILDEELQRLKAEVTEAARQRSQVE 2367
Query: 134 KAISMVEALAVKL--------TQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFA 185
+ + V +L +N ++++K ++ FL++ +E K++ + +
Sbjct: 2368 EELFSVRVQMEELGKLKARIEAENRALILRDKDNTQR---FLEEEAEKMKQVAEEAARLS 2424
Query: 186 CAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKL---LHQPSKVMA 242
A E+AR + + EE +A + + ++ VQEA R+K L Q K +A
Sbjct: 2425 VAAQEAAR-----LRQLAEEDLAQQRALAEKMLKEKMQAVQEATRLKAEAELLQQQKELA 2479
Query: 243 MEY--ELRALRDQIREKSI---------FSIKLQKELTMSKRDEENK 278
E L+A ++Q+ ++ + + Q++L MS E K
Sbjct: 2480 QEQARRLQADKEQMAQQLVEETQGFQRTLEAERQRQLEMSAEAERLK 2526
Score = 32.3 bits (72), Expect = 2.8
Identities = 49/255 (19%), Positives = 108/255 (42%), Gaps = 49/255 (19%)
Query: 91 KLSNEAKIREVASLEGHVLLKKLRDALEYLR-------------VRFAGRNKEDVEKAIS 137
++ E RE A+++ + +++ L++LR V A R++ +E+ I
Sbjct: 1536 RMQEEVTRREEAAVDAQQQKRSIQEELQHLRQSSEAEIQAKAQQVEAAERSRMRIEEEIR 1595
Query: 138 MVEALAVKLTQN-----EGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESA 192
+V L ++ T+ E EL + ++ +QA E+A++L Q + + + ++
Sbjct: 1596 VVR-LQLETTERQRGGAEDELQALRARAEEAEAQKRQAQEEAERLRRQVQDESQRKRQAE 1654
Query: 193 RSVVLRI---GEALEEQEQ--------------------DSQASKPQDVDGLVEEVQEAR 229
+ LR+ EA E+++ ++A + + V +E Q +
Sbjct: 1655 AELALRVKAEAEAAREKQRALQALDELKLQAEEAERWLCQAEAERARQVQVALETAQRSA 1714
Query: 230 RIKL-LHQPSKVMAMEYELRALRD------QIREKSIFSIKLQKELTMSKRDEENKSHPY 282
++L +PS R L++ Q+RE++ + Q E ++ + E + +
Sbjct: 1715 EVELQSKRPSFAEKTAQLERTLQEEHVTVTQLREEAERRAQQQAEAERAREEAERELERW 1774
Query: 283 MLHGSEALGSYLKVQ 297
L +EAL L+ +
Sbjct: 1775 QLKANEALRLRLQAE 1789
>YH32_AQUAE (O67622) Hypothetical UPF0144 protein AQ_1732
Length = 558
Score = 45.1 bits (105), Expect = 4e-04
Identities = 37/130 (28%), Positives = 68/130 (51%), Gaps = 9/130 (6%)
Query: 152 ELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALE------E 205
E+I+E E +++ LK+A E A+K+V + + A I A+ V RI E +E +
Sbjct: 52 EIIKEAKEKAEVI--LKEAKESAEKIVREAEEKAEKLIREAKEEVERIKEEVERRKKELK 109
Query: 206 QEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQ 265
+ +++ +K + +D E + E R +LLH+ ++ E L RD+IR K ++
Sbjct: 110 EREENVLAKERHLDRRWEAL-EKREEELLHRERELKDFERSLERWRDEIRHKEEELKHMK 168
Query: 266 KELTMSKRDE 275
+E+ K+ E
Sbjct: 169 EEVEELKKKE 178
>KF4A_CHICK (Q90640) Chromosome-associated kinesin KIF4A
(Chromokinesin)
Length = 1225
Score = 44.3 bits (103), Expect = 7e-04
Identities = 71/272 (26%), Positives = 121/272 (44%), Gaps = 48/272 (17%)
Query: 39 DSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNE-AK 97
D Q L ETVED + +VI +++ + +F AAAA+ + E A
Sbjct: 449 DLQKLLETVEDEELKENVEVIR-------------NLQQVLAQFQSESAAAAEAATEMAN 495
Query: 98 IREVASLE---GHVLLKKLRDALEYLRVRFAGRNKEDVE--KAISMVEALAVKLTQNEGE 152
+ A+ E G V + D +R A +KE VE KA+++ EALA K+ QN+ +
Sbjct: 496 AEQDAAGEAETGQVTKRSSDDFTTQHALRQAQMSKELVELNKALALKEALAKKMIQNDSQ 555
Query: 153 L--IQEKFEVK------KLLNFLKQASE------DAKKLVNQEK--SFACAEIESARSVV 196
L IQ +++ ++ N K+ E AKK VNQ K ++ +
Sbjct: 556 LEPIQSQYQTNIKDLELEVSNLQKEKEELILALSMAKKDVNQAKLSERRRKRLQELEGQI 615
Query: 197 LRIGEALEEQEQ--DSQASKPQDVDGLVEEVQEAR--RIKLLHQPSKVMAMEYELRALRD 252
+ + L EQ + + S + V L +E++E + R++L+ Q M + E
Sbjct: 616 NELKKKLNEQAKLLKLKESTERTVSKLNQEIREMKNQRVQLMRQ----MKEDAEKFRQWK 671
Query: 253 QIREKSIFSI-----KLQKELTMSKRDEENKS 279
Q ++K + + K Q EL +RD + ++
Sbjct: 672 QQKDKEVIQLKERDRKRQYELLKLERDFQKQA 703
>CENF_HUMAN (P49454) CENP-F kinetochore protein (Centromere protein F)
(Mitosin) (AH antigen)
Length = 3210
Score = 43.9 bits (102), Expect = 0.001
Identities = 65/277 (23%), Positives = 124/277 (44%), Gaps = 45/277 (16%)
Query: 7 EIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSL-KDVIEQEATN 65
E E+K E + +S+DVS + K L Q LE+ D+Q SL K +E +
Sbjct: 2102 EAEVKEKTELLQTLSSDVSELLK--DKTHLQEKLQSLEK---DSQALSLTKCELENQIAQ 2156
Query: 66 LSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLK-------------- 111
L+ + ++L K ++L A S+ K+ +LE ++ K
Sbjct: 2157 LNKE------KELLVKESESLQARLSESDYEKLNVSKALEAALVEKGEFALRLSSTQEEV 2210
Query: 112 -KLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQA 170
+LR +E LRVR K+ + +A KL + E E K +V+ L L Q
Sbjct: 2211 HQLRRGIEKLRVRIEADEKKQLH--------IAEKLKERERENDSLKDKVENLEREL-QM 2261
Query: 171 SEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARR 230
SE+ ++LV + + AE+E+ ++ + + +L+ E D + + + L +++QE
Sbjct: 2262 SEENQELVILDAENSKAEVETLKTQIEEMARSLKVFELDLVTLR-SEKENLTKQIQE--- 2317
Query: 231 IKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKE 267
+ ++ ++ L + + + EK I++++E
Sbjct: 2318 -----KQGQLSELDKLLSSFKSLLEEKEQAEIQIKEE 2349
>RA50_ARCFU (O29230) DNA double-strand break repair rad50 ATPase
Length = 886
Score = 43.5 bits (101), Expect = 0.001
Identities = 46/218 (21%), Positives = 104/218 (47%), Gaps = 20/218 (9%)
Query: 93 SNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGE 152
S E IR++ +E + K A+ +R R KE +++ +S E + + + + E
Sbjct: 140 SRERIIRQITRIEDYENAWKNLGAV----IRMLEREKERLKEFLSQEEQIKRQKEEKKAE 195
Query: 153 LIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQA 212
+ + E+K + + ++ SE+ + L ++ K E+E +S + E+L +QE +
Sbjct: 196 IERISEEIKSIESLREKLSEEVRNLESRLK-----ELEEHKSRL----ESLRKQE----S 242
Query: 213 SKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSI--KLQKELTM 270
S Q+V GL E+++E + +L ++ +E + + +++ + +SI KL E+
Sbjct: 243 SVLQEVRGLEEKLRELEK-QLKEVVERIEDLEKKAKEVKELKPKAERYSILEKLLSEINQ 301
Query: 271 SKRDEENKSHPYMLHGSEALGSYLKVQPRSGEVPQVSK 308
+ RD E + + K + + ++ +++K
Sbjct: 302 ALRDVEKREGDLTREAAGIQAQLKKAEEDNSKLEEITK 339
>SOJO_XENLA (Q9PW73) Cytoskeletal protein Sojo (p170)
Length = 1335
Score = 43.1 bits (100), Expect = 0.002
Identities = 56/250 (22%), Positives = 109/250 (43%), Gaps = 33/250 (13%)
Query: 40 SQILEETVEDNQGPSLKDVIEQEATNLSDQHKRIS-VRDLACKFDKNLAAAAKLSNEAKI 98
+Q L+ET E+N L+ ++Q+ L RI + D + +K ++ KL E +
Sbjct: 724 TQSLQETSEENV--RLQQTLQQQQHMLQQGTGRIGELEDHHTELEKQVS---KLEFELEK 778
Query: 99 REVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALA-------VKLTQNEG 151
+ S + +L++ +D+L G E+V++ S + + V L Q E
Sbjct: 779 QRSMSED---MLQRTKDSLHAANKEL-GLKTEEVQELCSTLNQVKLELKHTNVTLLQMEE 834
Query: 152 ELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQ 211
EL+ K + +K + LK D +K + ACA +E + + ++ +E ++Q
Sbjct: 835 ELVSLKNKEEKNASMLKLLQMDMQKTQVELDKKACAVLELEEKLHIAEKDSKRTEEMETQ 894
Query: 212 ASKPQ-DVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTM 270
S Q ++DG ++V+E ++ L + EK + L ++L
Sbjct: 895 LSGMQKELDGYTKQVEE---------------LQETLTKTHLSVEEKQVIIQGLTEKLRS 939
Query: 271 SKRDEENKSH 280
K++ E + H
Sbjct: 940 YKQELEERDH 949
>KF4A_HUMAN (O95239) Chromosome-associated kinesin KIF4A
(Chromokinesin)
Length = 1232
Score = 43.1 bits (100), Expect = 0.002
Identities = 60/245 (24%), Positives = 118/245 (47%), Gaps = 38/245 (15%)
Query: 39 DSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKI 98
D Q L ET+ED + LK+ +E NL ++S +AC +AAA + +
Sbjct: 446 DLQKLVETLEDQE---LKENVEI-ICNLQQLITQLSDETVAC-----MAAAI----DTAV 492
Query: 99 REVASLEGHVLLKKLRDALEYLR-VRFAGRNKEDVE--KAISMVEALAVKLTQNEGELIQ 155
+ A +E + DA +R A +KE VE KA+++ EALA K+TQN+ +L
Sbjct: 493 EQEAQVETSPETSRSSDAFTTQHALRQAQMSKELVELNKALALKEALARKMTQNDSQLQP 552
Query: 156 EKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKP 215
+++ + + L+ + +K EK E+++A+ +A + + + + +
Sbjct: 553 IQYQYQDNIKELELEVINLQK----EKEELVLELQTAKK------DANQAKLSERRRKRL 602
Query: 216 QDVDGLVEEVQEARRIKLLHQPSKVMAM----EYELRALRDQIREKSIFSIKLQKELTMS 271
Q+++G + +++ K L++ SK++ + E + L +IR ++L +++
Sbjct: 603 QELEGQIADLK-----KKLNEQSKLLKLKESTERTVSKLNQEIRMMKNQRVQLMRQM--- 654
Query: 272 KRDEE 276
K D E
Sbjct: 655 KEDAE 659
Score = 42.4 bits (98), Expect = 0.003
Identities = 70/311 (22%), Positives = 133/311 (42%), Gaps = 44/311 (14%)
Query: 8 IEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIEQEAT--- 64
I+ + EA S + S SD F +Q+ +E VE N+ +LK+ + ++ T
Sbjct: 488 IDTAVEQEAQVETSPETSRSSDAFTTQHALRQAQMSKELVELNKALALKEALARKMTQND 547
Query: 65 --------NLSDQHKRISVRDLACKFDK-----NLAAAAKLSNEAKIRE-----VASLEG 106
D K + + + + +K L A K +N+AK+ E + LEG
Sbjct: 548 SQLQPIQYQYQDNIKELELEVINLQKEKEELVLELQTAKKDANQAKLSERRRKRLQELEG 607
Query: 107 HV--LLKKLRDALEYLRVR-FAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEV--- 160
+ L KKL + + L+++ R + + I M++ V+L + E EKF
Sbjct: 608 QIADLKKKLNEQSKLLKLKESTERTVSKLNQEIRMMKNQRVQLMRQMKE-DAEKFRQWKQ 666
Query: 161 ---KKLLNFLKQASEDAKKLVNQEKSF------ACAEIESARSVVLRIGEALEEQEQDSQ 211
K+++ ++ + +L+ E++F + E A + R+ +AL++Q + +
Sbjct: 667 KRDKEVIQLKERDRKRQYELLKLERNFQKQSNVLRRKTEEAAAANKRLKDALQKQREVAD 726
Query: 212 ASKPQDVDGLVEEVQEARRIKLLHQPSKVM-AMEYELRALRDQIREKSIFSIKLQKELTM 270
K G+ E AR L +VM + E R L D + ++ I L +++
Sbjct: 727 KRKETQSRGM--EGTAARVKNWLGNEIEVMVSTEEAKRHLNDLLEDRKI----LAQDVAQ 780
Query: 271 SKRDEENKSHP 281
K +E+ +P
Sbjct: 781 LKEKKESGENP 791
Score = 35.4 bits (80), Expect = 0.34
Identities = 49/191 (25%), Positives = 86/191 (44%), Gaps = 27/191 (14%)
Query: 93 SNEAKIREVASLEGHVLLKKLRDA---LEYLRVRFAGRNKEDVEKAISMVEALAVKLTQN 149
S ++ +++ SLE + + + A + L R K+ E +++EA L
Sbjct: 812 SEDSITKQIESLETEMEFRSAQIADLQQKLLDAESEDRPKQRWENIATILEAKCA-LKYL 870
Query: 150 EGELIQEKFEVKKLLNFLKQAS---EDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQ 206
GEL+ K +V KL + LKQ+ D +K++ +E++ AEIE+ L +E+Q
Sbjct: 871 IGELVSSKIQVSKLESSLKQSKTSCADMQKMLFEERNH-FAEIETELQAEL---VRMEQQ 926
Query: 207 EQD------SQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRD-------Q 253
Q+ SQ + Q + +EE + +LL S + + EL +R+
Sbjct: 927 HQEKVLYLLSQLQQSQMAEKQLEESVSEKEQQLL---STLKCQDEELEKMREVCEQNQQL 983
Query: 254 IREKSIFSIKL 264
+RE I KL
Sbjct: 984 LRENEIIKQKL 994
>CEP2_HUMAN (Q9BV73) Centrosomal protein 2 (Centrosomal
Nek2-associated protein 1) (C-NAP1) (Centrosome protein
250) (Centrosome associated protein CEP250)
Length = 2442
Score = 43.1 bits (100), Expect = 0.002
Identities = 59/219 (26%), Positives = 101/219 (45%), Gaps = 28/219 (12%)
Query: 84 KNLAAAAKLSNEAKIREVASLEGHVL-LKKLRD----ALEYLRVRFAGRNKE-------- 130
+NLA + E + EV +L G + L+K R+ ALE L + RN+E
Sbjct: 1421 ENLALLTQTLAERE-EEVETLRGQIQELEKQREMQKAALELLSLDLKKRNQEVDLQQEQI 1479
Query: 131 -DVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLK-QASEDAKKLVNQEKSFACAE 188
++EK S++E L + + + E +L ++ ++++L + Q + +L+ EK
Sbjct: 1480 QELEKCRSVLEHLPMAVQEREQKLTVQREQIRELEKDRETQRNVLEHQLLELEKKDQM-- 1537
Query: 189 IESARSVVLRIGE--------ALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKV 240
IES R V + + ALE +E + Q + +E +E +R+ L H +
Sbjct: 1538 IESQRGQVQDLKKQLVTLECLALELEENHHKMECQQKLIKELEGQRETQRVALTHLTLDL 1597
Query: 241 MAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKS 279
EL+A QI + S L +EL +RD+E KS
Sbjct: 1598 EERSQELQAQSSQIHDLESHSTVLAREL--QERDQEVKS 1634
Score = 38.1 bits (87), Expect = 0.052
Identities = 65/279 (23%), Positives = 124/279 (44%), Gaps = 32/279 (11%)
Query: 43 LEETVEDNQGPSLK-DVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREV 101
LEE ++ Q S + +E +T L+ + + RD K + + E +++
Sbjct: 1597 LEERSQELQAQSSQIHDLESHSTVLA---RELQERDQEVKSQREQIEELQRQKEHLTQDL 1653
Query: 102 ASLEGHVLLKKLR-DALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEV 160
+ ++L+K R LE R R +ED+E+ + +LT + +L+QE+ E
Sbjct: 1654 ERRDQELMLQKERIQVLEDQRTRQTKILEEDLEQIKLSLRERGRELT-TQRQLMQERAEE 1712
Query: 161 KKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQ--------- 211
K + ++ S + KL+ ++K E+E + + + E ++ EQ Q
Sbjct: 1713 GKGPSKAQRGSLEHMKLILRDKE---KEVECQQEHIHELQELKDQLEQQLQGLHRKVGET 1769
Query: 212 ----ASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRAL--RDQIREKSIFSIKLQ 265
+ + Q++ L +++QEAR L + S ++ RAL RDQ E LQ
Sbjct: 1770 SLLLSQREQEIVVLQQQLQEAREQGELKEQSLQSQLDEAQRALAQRDQELE------ALQ 1823
Query: 266 KELTMSKRDEEN-KSHPYMLHGSEALGSYLKVQPRSGEV 303
+E ++ EE K L G+ +++ ++ R GE+
Sbjct: 1824 QEQQQAQGQEERVKEKADALQGA-LEQAHMTLKERHGEL 1861
Score = 37.0 bits (84), Expect = 0.12
Identities = 48/250 (19%), Positives = 109/250 (43%), Gaps = 14/250 (5%)
Query: 35 KLGADSQILEETVEDNQGPSL--KDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKL 92
+L Q+++E E+ +GPS + +E L D+ K + + + L +
Sbjct: 1698 ELTTQRQLMQERAEEGKGPSKAQRGSLEHMKLILRDKEKEVECQQEHIHELQELKDQLEQ 1757
Query: 93 SNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGE 152
+ R+V E +LL + + L+ + +E E+ ++L +L + +
Sbjct: 1758 QLQGLHRKVG--ETSLLLSQREQEIVVLQQQL----QEAREQGELKEQSLQSQLDEAQRA 1811
Query: 153 LIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQA 212
L Q E++ L +QA +++ + + A ++ ++ R GE + +EQ +
Sbjct: 1812 LAQRDQELEALQQEQQQAQGQEERVKEKADALQGALEQAHMTLKERHGELQDHKEQARRL 1871
Query: 213 SKPQDVDG----LVEEVQEARRIKLLHQPSKVMAMEYEL--RALRDQIREKSIFSIKLQK 266
+ V+G +EEV R + Q ++A++ + +A ++ +++ LQ
Sbjct: 1872 EEELAVEGRRVQALEEVLGDLRAESREQEKALLALQQQCAEQAQEHEVETRALQDSWLQA 1931
Query: 267 ELTMSKRDEE 276
+ + +RD+E
Sbjct: 1932 QAVLKERDQE 1941
>MFP1_LYCES (P93203) MAR binding filament-like protein 1
Length = 697
Score = 42.7 bits (99), Expect = 0.002
Identities = 40/172 (23%), Positives = 81/172 (46%), Gaps = 20/172 (11%)
Query: 122 VRFAGRNKEDVEKAISMVEALAVKL---TQNEGELIQE-KFEVKKLLNFLKQASEDAKKL 177
+R G ++ + + + + L QNE +L ++ KFE+K L N L ED KKL
Sbjct: 175 IRREGEERQALVNQLKSAKTTVISLGQELQNEKKLAEDLKFEIKGLQNDLMNTKEDKKKL 234
Query: 178 VNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQP 237
+ K +++ + + +I L + +D + S + L E+ E + ++Q
Sbjct: 235 QEELKE----KLDLIQVLEEKI-TLLTTEIKDKEVSLRSNTSKLAEKESEVNSLSDMYQQ 289
Query: 238 S--KVMAMEYELRALRDQIREK---------SIFSIKLQKELTMSKRDEENK 278
S ++M + E++ L+D+I+++ S ++ +Q + +RDE K
Sbjct: 290 SQDQLMNLTSEIKELKDEIQKRERELELKCVSEDNLNVQLNSLLLERDESKK 341
>EEA1_HUMAN (Q15075) Early endosome antigen 1 (Endosome-associated
protein p162) (Zinc finger FYVE domain containing
protein 2)
Length = 1411
Score = 42.4 bits (98), Expect = 0.003
Identities = 47/237 (19%), Positives = 100/237 (41%), Gaps = 37/237 (15%)
Query: 59 IEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALE 118
+EQ+ L Q K++ L K K A L + ++ L L K+L
Sbjct: 721 LEQKTEELEGQIKKLEADSLEVKASKE-QALQDLQQQRQLNTDLELRATELSKQLE---- 775
Query: 119 YLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGE--LIQEKFEV---------KKLLNFL 167
+ + D++K +E++ KLT+ E E ++++ FE ++L N +
Sbjct: 776 -MEKEIVSSTRLDLQKKSEALESIKQKLTKQEEEKQILKQDFETLSQETKIQHEELNNRI 834
Query: 168 KQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQE 227
+ + +K V EK E+ + + + ++ ++L+ + + + +E Q+
Sbjct: 835 QTTVTELQK-VKMEKEALMTELSTVKDKLSKVSDSLKNSKSEFE-----------KENQK 882
Query: 228 ARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKSHPYML 284
+ + ++ +E + L+ Q++ + ++K QKEL S E+ SH L
Sbjct: 883 GK--------AAILDLEKTCKELKHQLQVQMENTLKEQKELKKSLEKEKEASHQLKL 931
Score = 41.2 bits (95), Expect = 0.006
Identities = 47/207 (22%), Positives = 94/207 (44%), Gaps = 38/207 (18%)
Query: 91 KLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNE 150
K S+E K +++ +L+G + + L+ K ++E + +LTQ
Sbjct: 965 KQSSEQKKKQIEALQGELKIAVLQ--------------KTELENKLQQ------QLTQAA 1004
Query: 151 GELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDS 210
EL EK ++ L N +++ E K+L + +E+ + R + + E L ++D
Sbjct: 1005 QELAAEKEKISVLQNNYEKSQETFKQLQSDFYGRE-SELLATRQDLKSVEEKLSLAQEDL 1063
Query: 211 QASKPQ--DVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKS---------- 258
+++ Q + + L++E++ A+ K ++ +AL+D +EKS
Sbjct: 1064 ISNRNQIGNQNKLIQELKTAKATLEQDSAKKEQQLQERCKALQDIQKEKSLKEKELVNEK 1123
Query: 259 -----IFSIKLQKELTMSKRDEENKSH 280
I IK ++E ++K +EE KSH
Sbjct: 1124 SKLAEIEEIKCRQEKEITKLNEELKSH 1150
Score = 40.8 bits (94), Expect = 0.008
Identities = 82/406 (20%), Positives = 162/406 (39%), Gaps = 66/406 (16%)
Query: 38 ADSQILEETVEDNQGPS-----LKDVIEQEATNLSDQHKRISVRDLACKFD--KNLAAAA 90
A+ Q LE+ +E+ Q + +KD+ EQ+A L+ + + D+ K+D ++L AA
Sbjct: 135 AELQSLEQQLEEAQTENFNIKQMKDLFEQKAAQLATE-----IADIKSKYDEERSLREAA 189
Query: 91 ---------KLSNEAK--------------IREVASLEGH-VLLKKLRDALEYLRVRFAG 126
+L+ EA I +VA L+ V ++ L D + R R +
Sbjct: 190 EQKVTRLTEELNKEATVIQDLKTELLQRPGIEDVAVLKKELVQVQTLMDNMTLERERESE 249
Query: 127 RNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFAC 186
+ K++ +K S + ++Q EL + EV + L++ +L + ++
Sbjct: 250 KLKDECKKLQSQYASSEATISQLRSELAKGPQEVAVYVQELQKLKSSVNELTQKNQTLTE 309
Query: 187 AEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYE 246
++ + LEE+ + SK L ++ + ++++ S++ A E
Sbjct: 310 NLLKKEQDYT-----KLEEKHNEESVSKKNIQATLHQKDLDCQQLQ-----SRLSASETS 359
Query: 247 LRALRDQIREKSIFSIKLQKELT---------------MSKRDEENKSHPYMLHG--SEA 289
L + ++ EK + KL++EL+ + ++ EE + H L ++
Sbjct: 360 LHRIHVELSEKGEATQKLKEELSEVETKYQHLKAEFKQLQQQREEKEQHGLQLQSEINQL 419
Query: 290 LGSYLKVQPRSGEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVGRILQVDI 349
L+ + + GE K + +LSSE + AD + + + +
Sbjct: 420 HSKLLETERQLGEAHGRLK---EQRQLSSEKLMDKEQQVADLQLKLSRLEEQLKEKVTNS 476
Query: 350 VSNGKKLTLTTNPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKD 395
+L T Q L A LR++ D V+ Q+ KD
Sbjct: 477 TELQHQLDKTKQQHQEQQALQQSTTAKLREAQNDLEQVLRQIGDKD 522
>MYHD_HUMAN (Q9UKX3) Myosin heavy chain, skeletal muscle, extraocular
(MyHC-eo)
Length = 1938
Score = 41.6 bits (96), Expect = 0.005
Identities = 66/292 (22%), Positives = 126/292 (42%), Gaps = 21/292 (7%)
Query: 7 EIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSL---KDVIEQEA 63
E ++KM E++ + + I ++ K + L+ ++D Q SL K + E +A
Sbjct: 1060 EGDLKMSQESIMDLENEKQQIEEKLKKKEFELSQ--LQARIDDEQVHSLQFQKKIKELQA 1117
Query: 64 TNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIR-EVAS--LEGHVLLKKLRDALEYL 120
+ + + L K +K + A+ E R E AS + + K R+A E+
Sbjct: 1118 RIEELEEEIEAEHTLRAKIEKQRSDLARELEEISERLEEASGATSAQIEMNKKREA-EFQ 1176
Query: 121 RVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQ 180
++R D+E+A EA A L + + + + E E L +KQ E K +
Sbjct: 1177 KMR------RDLEEATLQHEATAATLRKKQADSVAELGEQIDNLQRVKQKLEKEKSELKM 1230
Query: 181 EKSFACAEIES---ARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQP 237
E + IE+ ++S + R +E+Q + +A Q L+ ++ ++ +L Q
Sbjct: 1231 EIDDMASNIEALSKSKSNIERTCRTVEDQFSEIKAKDEQQTQ-LIHDL-NMQKARLQTQN 1288
Query: 238 SKVMAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKSHPYMLHGSEA 289
++ E +L Q+ KS ++ Q E + +EE K+ M H ++
Sbjct: 1289 GELSHRVEEKESLISQL-TKSKQALTQQLEELKRQMEEETKAKNAMAHALQS 1339
Score = 32.0 bits (71), Expect = 3.7
Identities = 56/225 (24%), Positives = 93/225 (40%), Gaps = 33/225 (14%)
Query: 43 LEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACK-FDKNLAAAAKLSNEAKIR-E 100
LE+T + QG +E +L H + D + FDK LA + +E++ E
Sbjct: 1418 LEKTKQRLQGE-----VEDLMRDLERSHTACATLDKKQRNFDKVLAEWKQKLDESQAELE 1472
Query: 101 VASLEGHVLLKKLRDALEYLRVRFAGRNK-EDVEKAISMVEALAVKLTQNEGELIQEKFE 159
A E L +L F RN E+V + + L + +L ++ E
Sbjct: 1473 AAQKESRSLSTEL----------FKMRNAYEEVVDQLETLRRENKNLQEEISDLTEQIAE 1522
Query: 160 VKKLLNFLKQASEDAKKLVNQEKS---FACAEIESA----RSVVLRIGEALEEQEQDSQA 212
K L Q +E KKLV QEKS A E+E + S +LR+ L + + +
Sbjct: 1523 TGKNL----QEAEKTKKLVEQEKSDLQVALEEVEGSLEHEESKILRVQLELSQVKSELD- 1577
Query: 213 SKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREK 257
+ V EE+++ +R + ++ E+R+ D +R K
Sbjct: 1578 ---RKVIEKDEEIEQLKRNSQRAAEALQSVLDAEIRSRNDALRLK 1619
>RA50_SULSO (Q97WH0) DNA double-strand break repair rad50 ATPase
Length = 864
Score = 41.2 bits (95), Expect = 0.006
Identities = 57/273 (20%), Positives = 120/273 (43%), Gaps = 41/273 (15%)
Query: 32 PKYKLGADSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAK 91
PKY+ Q L+ + + +LK+ +E++A+ LS+ +++ L K ++
Sbjct: 344 PKYQ-----QYLKLKSDLDSKLNLKERLEKDASELSNDIDKVN--SLEQKVEETRKKQLN 396
Query: 92 LSNE-AKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNE 150
L + AK+ + S + ++ + E V ++E +K I ++ ++L N+
Sbjct: 397 LRAQLAKVESLISEKNEIINNISQVEGETCPVCGRPLDEEHKQKIIKEAKSYILQLELNK 456
Query: 151 GELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDS 210
EL +E +KK+ N L + + ++L N + S+ ++ + ++ E +E +
Sbjct: 457 NELEEE---LKKITNELNKIEREYRRLSNNKASY-----DNVMRQLKKLNEEIENLHSEI 508
Query: 211 QASK--PQDVDGLVEEVQEAR--------------------RIKLLHQPSKVMAMEYELR 248
++ K +++ + EEV+E + R+KL K +E E+R
Sbjct: 509 ESLKNIDEEIKKINEEVKELKLYYEEFMRLSKYTKEELDKKRVKLDEMKKKKEEIEKEMR 568
Query: 249 ALRDQIR---EKSIFSIKLQKELTMSKRDEENK 278
L +++ K++ S L E K DE K
Sbjct: 569 GLESELKGLDRKALESKILDLENKRVKLDEMKK 601
>MOEH_DROME (P46150) Moesin/ezrin/radixin homolog 1
(Ezrin-moesin-radixin-1) (D17 protein) (Moesin protein)
(dMoesin)
Length = 578
Score = 41.2 bits (95), Expect = 0.006
Identities = 24/133 (18%), Positives = 71/133 (53%), Gaps = 5/133 (3%)
Query: 148 QNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQE 207
Q E E +Q ++ +Q ED K + ++ + ++ A+ ++ R+ E L++ +
Sbjct: 318 QQEREKLQLALAARERAEKKQQEYEDRLKQMQEDMERSQRDLLEAQDMIRRLEEQLKQLQ 377
Query: 208 --QDSQASKPQDVDGLVEEVQEARRIKLLHQ---PSKVMAMEYELRALRDQIREKSIFSI 262
+D + +++ +++ ++EA+ ++ + + ++MA + E++ ++D++ K +
Sbjct: 378 AAKDELELRQKELQAMLQRLEEAKNMEAVEKLKLEEEIMAKQMEVQRIQDEVNAKDEETK 437
Query: 263 KLQKELTMSKRDE 275
+LQ E+ ++R +
Sbjct: 438 RLQDEVEDARRKQ 450
Score = 34.7 bits (78), Expect = 0.57
Identities = 40/149 (26%), Positives = 70/149 (46%), Gaps = 22/149 (14%)
Query: 86 LAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVK 145
LA AA+ E K +E LK++++ +E R++ D+ +A M+ L +
Sbjct: 326 LALAARERAEKKQQEYEDR-----LKQMQEDME--------RSQRDLLEAQDMIRRLEEQ 372
Query: 146 LTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEE 205
L Q + + + K+L L Q E+AK + EK EI + + V RI
Sbjct: 373 LKQLQAAKDELELRQKELQAML-QRLEEAKNMEAVEKLKLEEEIMAKQMEVQRI------ 425
Query: 206 QEQDSQASKPQDVDGLVEEVQEARRIKLL 234
QD +K ++ L +EV++ARR +++
Sbjct: 426 --QDEVNAKDEETKRLQDEVEDARRKQVI 452
>KF4A_MOUSE (P33174) Chromosome-associated kinesin KIF4A
(Chromokinesin)
Length = 1231
Score = 41.2 bits (95), Expect = 0.006
Identities = 59/245 (24%), Positives = 114/245 (46%), Gaps = 38/245 (15%)
Query: 39 DSQILEETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKI 98
D Q L ET+ED + LK+ IE NL ++S AC + A + EA
Sbjct: 447 DLQKLVETLEDQE---LKENIEI-ICNLQQVIAQLSDEAAAC-----MTATIDTAGEADT 497
Query: 99 REVASLEGHVLLKKLRDALEYLR-VRFAGRNKEDVE--KAISMVEALAVKLTQNEGELIQ 155
+ +S + + D +R A +KE +E KA+++ EALA K+TQN+ +L
Sbjct: 498 QVQSSPD----TSRSSDVFSTQHALRQAQMSKELIELNKALALKEALAKKMTQNDNQLQP 553
Query: 156 EKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKP 215
+F+ + + L E + +EK E+++A+ +A + + + + +
Sbjct: 554 IQFQYQDNIKNL----ESEVLSLQREKEELVLELQTAKK------DANQAKLSERRRKRL 603
Query: 216 QDVDGLVEEVQEARRIKLLHQPSKVMAM----EYELRALRDQIREKSIFSIKLQKELTMS 271
Q+++G + +++ K L + SK++ + E+ + L +IR ++L +++
Sbjct: 604 QELEGQIADLK-----KKLQEQSKLLKLKESTEHTVSKLNQEIRMMKNQRVQLMRQM--- 655
Query: 272 KRDEE 276
K D E
Sbjct: 656 KEDAE 660
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.131 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 52,078,483
Number of Sequences: 164201
Number of extensions: 2075204
Number of successful extensions: 8185
Number of sequences better than 10.0: 323
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 309
Number of HSP's that attempted gapping in prelim test: 7906
Number of HSP's gapped (non-prelim): 567
length of query: 490
length of database: 59,974,054
effective HSP length: 114
effective length of query: 376
effective length of database: 41,255,140
effective search space: 15511932640
effective search space used: 15511932640
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0269a.3


