

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0263.7
(787 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
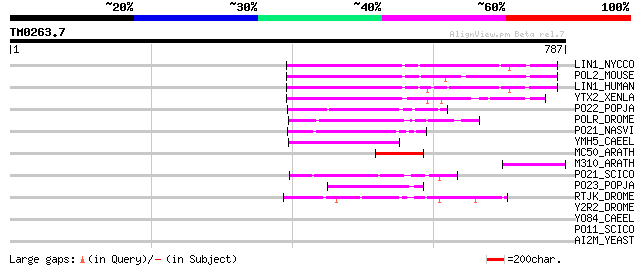
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 142 3e-33
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 135 4e-31
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 124 1e-27
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 109 2e-23
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 81 9e-15
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 81 1e-14
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 71 1e-11
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 66 3e-10
MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250... 65 5e-10
M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310... 65 5e-10
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 60 2e-08
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 52 5e-06
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 45 7e-04
Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I retro... 39 0.053
YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III 37 0.16
PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type... 34 1.7
AI2M_YEAST (P03876) Putative COX1/OXI3 intron 2 protein 33 3.8
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 142 bits (358), Expect = 3e-33
Identities = 104/393 (26%), Positives = 199/393 (50%), Gaps = 19/393 (4%)
Query: 393 ITLIPKV-ENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFML 451
ITLIPK ++P NYRPISL+ KI+ K+L+ R+++ + K+I Q FI G
Sbjct: 513 ITLIPKPGKDPTRKENYRPISLMNIDAKILNKILTNRIQQHIKKIIHHDQVGFIPGSQGW 572
Query: 452 DSVVVANEVVEEAKRRK-KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGC 510
++ + V++ + K K+ + +D EKA+D++ F+ + +IG ++ I+
Sbjct: 573 FNIRKSINVIQHINKLKNKDHMILSIDAEKAFDNIQHPFMIRTLKKIGIEGTFLKLIEAI 632
Query: 511 IQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRV 570
++++++NG + F + G RQG PL+P LF IV E L +R ++K G +
Sbjct: 633 YSKPTANIILNGVKLKSFPLRSGTRQGCPLSPLLFNIVMEVLAIAIR---EEKAIKGIHI 689
Query: 571 GGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSR 630
G +E+++SL FADD + + E + + + +++ + +SG K+N KS + ++
Sbjct: 690 GSEEIKLSL--FADDMIVYLENTRDSTTKLLEVIKEYSNVSGYKINTHKSVAFIYTNNNQ 747
Query: 631 ETQIFAAMLNCKVMGVPFTYLGIPVGANPRK--AATWEPVIKKMQRKLSL*KHKSLSFGG 688
+ + V+ YLG+ + + + +E + K++ ++ K+ S+ G
Sbjct: 748 AEKTVKDSIPFTVVPKKMKYLGVYLTKDVKDLYKENYETLRKEIAEDVNKWKNIPCSWLG 807
Query: 689 RICLIKSVLSSLPLFFFSF----FKAPKCIIDQCNKIQRKFLWGGEEGRGVAWVKWEEIC 744
RI ++K +S LP ++F KAP KI F+W ++ + + +
Sbjct: 808 RINIVK--MSILPKAIYNFNAIPIKAPLSYFKDLEKIILHFIWNQKKPQ----IAKTLLS 861
Query: 745 KKKEEGGLGVKDLESFNKALLVKWRWRLLRERE 777
K + GG+ + DL + K++++K W + RE
Sbjct: 862 NKNKAGGITLPDLRLYYKSIVIKTAWYWHKNRE 894
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 135 bits (340), Expect = 4e-31
Identities = 108/397 (27%), Positives = 192/397 (48%), Gaps = 27/397 (6%)
Query: 393 ITLIPKVE-NPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFML 451
ITLIPK + +P + N+RPISL+ KI+ K+L+ R++E + +I Q FI G
Sbjct: 540 ITLIPKPQKDPTKIENFRPISLMNIDAKILNKILANRIQEHIKAIIHPDQVGFIPGMQGW 599
Query: 452 DSVVVANEVVEEAKRRK-KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGC 510
++ + V+ + K K + +D EKA+D + F+ ++ R G +++ IK
Sbjct: 600 FNIRKSINVIHYINKLKDKNHMIISLDAEKAFDKIQHPFMIKVLERSGIQGPYLNMIKAI 659
Query: 511 IQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRV 570
+++ VNG E + G RQG PL+P+LF IV E L +R QQK G ++
Sbjct: 660 YSKPVANIKVNGEKLEAIPLKSGTRQGCPLSPYLFNIVLEVLARAIR---QQKEIKGIQI 716
Query: 571 GGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVN------FFKSKLAG 624
G +EV+ISLL ADD + + + + +++ F + G K+N F +K
Sbjct: 717 GKEEVKISLL--ADDMIVYISDPKNSTRELLNLINSFGEVVGYKINSNKSMAFLYTKNKQ 774
Query: 625 ISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKA--ATWEPVIKKMQRKLSL*KHK 682
RET F+ + N YLG+ + + ++ + K+++ L K
Sbjct: 775 AEKEIRETTPFSIVTN------NIKYLGVTLTKEVKDLYDKNFKSLKKEIKEDLRRWKDL 828
Query: 683 SLSFGGRICLIKSVLSSLPLFFFSF--FKAPKCIIDQCNKIQRKFLWGGEEGRGVAWVKW 740
S+ GRI ++K + ++ F+ K P ++ KF+W ++ R +
Sbjct: 829 PCSWIGRINIVKMAILPKAIYRFNAIPIKIPTQFFNELEGAICKFVWNNKKPR----IAK 884
Query: 741 EEICKKKEEGGLGVKDLESFNKALLVKWRWRLLRERE 777
+ K+ GG+ + DL+ + +A+++K W R+R+
Sbjct: 885 SLLKDKRTSGGITMPDLKLYYRAIVIKTAWYWYRDRQ 921
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 124 bits (311), Expect = 1e-27
Identities = 104/397 (26%), Positives = 195/397 (48%), Gaps = 27/397 (6%)
Query: 393 ITLIPKV-ENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFML 451
I LIPK + N+RPISL+ KI+ K+L+ ++++ + K+I Q FI
Sbjct: 513 IILIPKPGRDTTKKENFRPISLMNIDAKILNKILANQIQQHIKKLIHHDQVGFIPAMQGW 572
Query: 452 DSVVVANEVVEEAKRRKK-ECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGC 510
++ + +++ R K + +D EKA+D + F+ ++++G ++ I+
Sbjct: 573 FNIRKSINIIQHINRTKDTNHMIISIDAEKAFDKIQQPFMLKPLNKLGIDGTYLKIIRAI 632
Query: 511 IQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRV 570
++++++NG E + G RQG PL+P L IV E L +R Q+K G ++
Sbjct: 633 YDKPTANIILNGQKLEAPPLKTGTRQGCPLSPLLPNIVLEVLARAIR---QEKEIKGIQL 689
Query: 571 GGDEVEISLLQFADDTLFFGE---ASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISM 627
G +EV++SL FADD + + E S QN+L ++ F +SG K+N KS+ A +
Sbjct: 690 GKEEVKLSL--FADDMIVYLENPIVSAQNLL---KLISNFSKVSGYKINVQKSQ-AFLYT 743
Query: 628 VSRETQI-FAAMLNCKVMGVPFTYLGIPVGANPRK--AATWEPVIKKMQRKLSL*KHKSL 684
+R+T+ + L + YLGI + + + ++P++ +++ + K+
Sbjct: 744 NNRQTESQIMSELPFTIASKRIKYLGIQLTRDVKDLFKENYKPLLNEIKEDTNKWKNIPC 803
Query: 685 SFGGRICLIKSVLSSLPLFFFSF----FKAPKCIIDQCNKIQRKFLWGGEEGRGVAWVKW 740
S+ GRI ++K ++ LP + F K P + K KF+W + A +
Sbjct: 804 SWVGRINIVK--MAILPKVIYRFNAIPIKLPMTFFTELEKTTLKFIWNQKR----AHIAK 857
Query: 741 EEICKKKEEGGLGVKDLESFNKALLVKWRWRLLRERE 777
+ +K + GG+ + D + + KA + K W + R+
Sbjct: 858 STLSQKNKAGGITLPDFKLYYKATVTKTAWYWYQNRD 894
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 109 bits (273), Expect = 2e-23
Identities = 98/381 (25%), Positives = 169/381 (43%), Gaps = 37/381 (9%)
Query: 393 ITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFMLD 452
++L+PK + + + N+RP+SL+ YKIVAK +S+RL+ VL++VI Q+ + GR + D
Sbjct: 510 LSLLPKKGDLRLIKNWRPVSLLSTDYKIVAKAISLRLKSVLAEVIHPDQSYTVPGRTIFD 569
Query: 453 SVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCIQ 512
+V + +++ A+R +D EKA+D VD +L + F +++ ++K
Sbjct: 570 NVFLIRDLLHFARRTGLSLAFLSLDQEKAFDRVDHQYLIGTLQAYSFGPQFVGYLKTMYA 629
Query: 513 SASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGG 572
SA V +N S T GRG+RQG PL+ L+ + E L+R K TG +
Sbjct: 630 SAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSLAIEPFLCLLR-----KRLTGLVLKE 684
Query: 573 DEVEISLLQFADDTLFFGE--------ASTQNILVVKSILR--WFEAM----SGLKVNFF 618
++ + L +ADD + + Q + S R W ++ LKV+F
Sbjct: 685 PDMRVVLSAYADDVILVAQDLVDLERAQECQEVYAAASSARINWSKSSGLLEGSLKVDFL 744
Query: 619 KSKLAGISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL 678
IS S+ + L+ + V ++ + E V+ ++ +
Sbjct: 745 PPAFRDISWESKIIKYLGVYLSAEEYPVSQNFIELE-----------ECVLTRLGKWKGF 793
Query: 679 *KHKSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEGRGVAWV 738
K LS GR +I +++S + + I + + FLW G+ WV
Sbjct: 794 --AKVLSMRGRALVINQLVASQIWYRLICLSPTQEFIAKIQRRLLDFLWIGKH-----WV 846
Query: 739 KWEEICKKKEEGGLGVKDLES 759
+EGG GV + S
Sbjct: 847 SAGVSSLPLKEGGQGVVCIRS 867
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 81.3 bits (199), Expect = 9e-15
Identities = 59/228 (25%), Positives = 116/228 (50%), Gaps = 10/228 (4%)
Query: 394 TLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFMLDS 453
TLIPK + + N+RPI++ + +++ ++L+ RL + + A I G + +
Sbjct: 62 TLIPKDGDLENPSNWRPITIASALQRLLHRILAKRLEAAVELHPAQKGYARIDGTLV--N 119
Query: 454 VVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCIQS 513
++ + + + ++K V +D KA+D+V + + + R+G ++I G +
Sbjct: 120 SLLLDTYISSRREQRKTYNVVSLDVRKAFDTVSHSSICRALQRLGIDEGTSNYITGSLSD 179
Query: 514 ASSSVLVN-GSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGG 572
+++++ V GS T + + RG++QGDPL+PFLF V + L ++Q G +G
Sbjct: 180 STTTIRVGPGSQTRKICIRRGVKQGDPLSPFLFNAVLDEL----LCSLQSTPGIGGTIG- 234
Query: 573 DEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKS 620
E +I +L FADD L + + ++ +F + G+ +N KS
Sbjct: 235 -EEKIPVLAFADDLLLLEDNDVLLPTTLATVANFFR-LRGMSLNAKKS 280
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 80.9 bits (198), Expect = 1e-14
Identities = 63/271 (23%), Positives = 124/271 (45%), Gaps = 19/271 (7%)
Query: 396 IPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFMLDSVV 455
IPK + ++RPIS+ + + + +L+ RL ++ D RQ F+ D+
Sbjct: 403 IPKTVTAKRPQDFRPISVPSVLVRQLNAILATRLNSSINW--DPRQRGFLPTDGCADNAT 460
Query: 456 VANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCIQSAS 515
+ + V+ + + + C++ +D KA+DS+ ++ + G ++ +++ +
Sbjct: 461 IVDLVLRHSHKHFRSCYIANLDVSKAFDSLSHASIYDTLRAYGAPKGFVDYVQNTYEGGG 520
Query: 516 SSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGGDEV 575
+S+ +G +EEF RG++QGDPL+P LF +V + L + + G +VG
Sbjct: 521 TSLNGDGWSSEEFVPARGVKQGDPLSPILFNLVMDRLLRTLPSEI------GAKVG--NA 572
Query: 576 EISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSRETQIF 635
+ FADD + F E + + V+ F ++ GLK+N K GI ++
Sbjct: 573 ITNAAAFADDLVLFAE-TRMGLQVLLDKTLDFLSIVGLKLNADKCFTVGIKGQPKQ---- 627
Query: 636 AAMLNCKVMGVPFTYLGIPVGANPRKAATWE 666
C V+ Y+G + ++ W+
Sbjct: 628 ----KCTVLEAQSFYVGSSEIPSLKRTDEWK 654
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 71.2 bits (173), Expect = 1e-11
Identities = 51/197 (25%), Positives = 101/197 (50%), Gaps = 10/197 (5%)
Query: 395 LIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFMLDSV 454
LIPK +RP+S+ + ++L+ R+ E ++D RQ AFI + ++
Sbjct: 376 LIPKEPGTMDPACFRPLSIASVALRHFHRILANRIGE--HGLLDTRQRAFIVADGVAENT 433
Query: 455 VVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCIQSA 514
+ + +++EA+ + K ++ +D +KA+DSV+ + + R + ++I +++
Sbjct: 434 SLLSAMIKEARMKIKGLYIAILDVKKAFDSVEHRSILDALRRKKLPLEMRNYIMWVYRNS 493
Query: 515 SSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGGDE 574
+ + V + RG+RQGDPL+P LF V ++ ++R+ + TG+ +G +
Sbjct: 494 KTRLEVVKTKGRWIRPARGVRQGDPLSPLLFNCV---MDAVLRRLPEN---TGFLMGAE- 546
Query: 575 VEISLLQFADDTLFFGE 591
+I L FADD + E
Sbjct: 547 -KIGALVFADDLVLLAE 562
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 66.2 bits (160), Expect = 3e-10
Identities = 42/158 (26%), Positives = 69/158 (43%), Gaps = 1/158 (0%)
Query: 396 IPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFMLDSVV 455
IPK NP NYRPISL +I+ +++ R+R S ++ Q F+ R S+V
Sbjct: 673 IPKKGNPSSPSNYRPISLTDPFARIMERIICSRIRSEYSHLLSPHQHGFLNFRSCPSSLV 732
Query: 456 VANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCIQSAS 515
+ + + +K + DF KA+D V L ++ G W K + +
Sbjct: 733 RSISLYHSILKNEKSLDILFFDFAKAFDKVSHPILLKKLALFGLDKLTCSWFKEFLHLRT 792
Query: 516 SSVLVNG-SPTEEFNMGRGLRQGDPLAPFLFLIVAEGL 552
SV +N + + + G+ QG P LF++ L
Sbjct: 793 FSVKINKFVSSNAYPISSGVPQGSVSGPLLFILFINDL 830
>MC50_ARATH (P92555) Hypothetical mitochondrial protein AtMg01250
(ORF102)
Length = 122
Score = 65.5 bits (158), Expect = 5e-10
Identities = 32/68 (47%), Positives = 44/68 (64%)
Query: 519 LVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGGDEVEIS 578
++NG+P RGLRQGDPL+P+LF++ E L+GL R+A +Q G RV + I+
Sbjct: 13 IINGAPQGLVTPSRGLRQGDPLSPYLFILCTEVLSGLCRRAQEQGRLPGIRVSNNSPRIN 72
Query: 579 LLQFADDT 586
L FADDT
Sbjct: 73 HLLFADDT 80
>M310_ARATH (P93295) Hypothetical mitochondrial protein AtMg00310
(ORF154)
Length = 154
Score = 65.5 bits (158), Expect = 5e-10
Identities = 32/91 (35%), Positives = 54/91 (59%), Gaps = 2/91 (2%)
Query: 699 SLPLFFFSFFKAPKCIIDQCNKIQRKFLWGG-EEGRGVAWVKWEEICKKKEE-GGLGVKD 756
+LP++ S F+ K + + +F W E R ++WV W+++CK KE+ GGLG +D
Sbjct: 2 ALPVYAMSCFRLSKLLCKKLTSAMTEFWWSSCENKRKISWVAWQKLCKSKEDDGGLGFRD 61
Query: 757 LESFNKALLVKWRWRLLREREWLWCQVLESK 787
L FN+ALL K +R++ + L ++L S+
Sbjct: 62 LGWFNQALLAKQSFRIIHQPHTLLSRLLRSR 92
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 60.5 bits (145), Expect = 2e-08
Identities = 60/242 (24%), Positives = 109/242 (44%), Gaps = 20/242 (8%)
Query: 397 PKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFMLDSVVV 456
PK+E G +RP+S+ + + K+L+ R V D+RQTA++ + +V +
Sbjct: 219 PKIEGGPDPGVFRPLSICSVILREFNKILARRF--VSCYTYDERQTAYLPIDGVCINVSM 276
Query: 457 ANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCIQSASS 516
++ EAKR +KE + +D KA++SV + L ++ G + +I + +
Sbjct: 277 LTAIIAEAKRLRKELHIAILDLVKAFNSVYHSALIDAITEAGCPPGVVDYIADMYNNVIT 336
Query: 517 SVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAE-GLNGLMRQAVQQKLFTGYRVGGDEV 575
+ G E ++ G+ QGDPL+ LF + E L L + R +V
Sbjct: 337 EMQFEGK-CELASILAGVYQGDPLSGPLFTLAYEKALRALNNEG---------RFDIADV 386
Query: 576 EISLLQFADDTLFFGEASTQNILVVKSILRWFE---AMSGLKVNFFKSKLAGISMVSRET 632
++ ++DD L ++ ++ L F A GL++N KSK + RE
Sbjct: 387 RVNASAYSDDGLLL----AMTVIGLQHNLDKFGETLAKIGLRINSRKSKTVSLVPSGREK 442
Query: 633 QI 634
++
Sbjct: 443 KM 444
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 52.4 bits (124), Expect = 5e-06
Identities = 36/137 (26%), Positives = 62/137 (44%), Gaps = 6/137 (4%)
Query: 451 LDSVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGC 510
L ++++ ++ + + K V +D KA+D+V + M G +I
Sbjct: 3 LANIIMLEHYIKLRRLKGKTYNVVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMST 62
Query: 511 IQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRV 570
I A ++++V G T + + G++QGDPL+P LF IV + L + T
Sbjct: 63 ITDAYTNIVVGGRTTNKIYIRNGVKQGDPLSPVLFNIVLDELVTRLNDEQPGASMT---- 118
Query: 571 GGDEVEISLLQFADDTL 587
+I+ L FADD L
Sbjct: 119 --PACKIASLAFADDLL 133
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 45.1 bits (105), Expect = 7e-04
Identities = 74/334 (22%), Positives = 136/334 (40%), Gaps = 36/334 (10%)
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREV--LSKVIDDRQTAFIG 446
+AS I + + P + +YRP SL+ + KI+ +L+ RL ++K I Q F
Sbjct: 496 SASIIMIHKTGKTPTDVDSYRPTSLLPSLGKIMERLILNRLLTCKDVTKAIPKFQFGF-- 553
Query: 447 GRFMLDSVVVANEVVE---EAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKW 503
R + + VV EA K+ +D ++A+D V L Y R+ F +
Sbjct: 554 -RLQHGTPEQLHRVVNFALEAMENKEYAVGAFLDIQQAFDRVWHPGLLYKAKRL-FPPQL 611
Query: 504 IHWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQK 563
+K ++ + V V+G + + G+ QG L P L+ + A + V +
Sbjct: 612 YLVVKSFLEERTFHVSVDGYKSSIKPIAAGVPQGSVLGPTLYSVFASDMP--THTPVTEV 669
Query: 564 LFTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWF-----EAMSGLKVNFF 618
DE ++ + +ADDT +++IL S L+ + + V
Sbjct: 670 ---------DEEDVLIATYADDTAVL--TKSKSILAATSGLQEYLDAFQQWAENWNVRIN 718
Query: 619 KSKLAGISMVSRETQIFAAMLNCKVM--GVPFTYLGIPVGAN---PRKAATWEPVIKKMQ 673
K A ++ +R LN +++ + YLGI + R + +
Sbjct: 719 AEKCANVTFANRTGSCPGVSLNGRLIRHHQAYKYLGITLDRKLTFSRHITNIQQAFRTKV 778
Query: 674 RKLS--L*KHKSLSFGGRICLIKSVLSSLPLFFF 705
++S + LS G ++ + KS+L+ P F+
Sbjct: 779 ARMSWLIAPRNKLSLGCKVNIYKSILA--PCLFY 810
>Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I
retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 38.9 bits (89), Expect = 0.053
Identities = 50/198 (25%), Positives = 86/198 (43%), Gaps = 29/198 (14%)
Query: 407 NYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFMLDSVV-VANEVVEEAK 465
+YR I L+ K++ ++ R+REVL + Q F GR + D+ V + V A
Sbjct: 511 SYRGICLLPVFGKVLEAIMVNRVREVLPEGCR-WQFGFRQGRCVEDAWRHVKSSVGASAA 569
Query: 466 RRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHK--WIHWIKGCIQSASSSVLVNGS 523
+ FV DF+ A+D+V+W+ ++ +G W + G +V+ + S
Sbjct: 570 QYVLGTFV---DFKGAFDNVEWSAALSRLADLGCREMGLWQSFFSG-----RRAVIRSSS 621
Query: 524 PTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGGDEVEISLLQFA 583
T E + RG QG PF++ I+ + L ++ Q L +A
Sbjct: 622 GTVEVPVTRGCPQGSISGPFIWDILMDVLLQRLQPYCQ-----------------LSAYA 664
Query: 584 DDTLFFGEASTQNILVVK 601
DD L E +++ +L K
Sbjct: 665 DDLLLLVEGNSRAVLEEK 682
>YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III
Length = 364
Score = 37.4 bits (85), Expect = 0.16
Identities = 24/90 (26%), Positives = 41/90 (44%)
Query: 476 VDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCIQSASSSVLVNGSPTEEFNMGRGLR 535
+DF KA+D V + L ++ I + I W+ + + S V V + +E G+
Sbjct: 23 LDFSKAFDKVSHDILLDKLTSIKINKHLIRWLDVFLTNRSFKVKVGNTLSEPKKTVCGVP 82
Query: 536 QGDPLAPFLFLIVAEGLNGLMRQAVQQKLF 565
QG ++P LF I ++ + V K F
Sbjct: 83 QGSVISPVLFGIFVNEISANLPVGVYCKQF 112
>PO11_SCICO (Q03277) Retrovirus-related Pol polyprotein from type I
retrotransposable element R1 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1004
Score = 33.9 bits (76), Expect = 1.7
Identities = 24/83 (28%), Positives = 42/83 (49%), Gaps = 3/83 (3%)
Query: 406 GNYRPISLVGCVYKIVAKLLSMRLREVLSKV-IDDRQTAFIGGRFMLDSVVVANEVVEEA 464
G+YRPI L+ + K++ ++ RL + L V + Q AF G+ D+ VE +
Sbjct: 481 GSYRPICLLDNLGKVLEGIMVKRLDQKLMDVEVSPYQFAFTYGKSTEDAWRCVQRHVECS 540
Query: 465 KRRKKECFVFKVDFEKAYDSVDW 487
+ K +DF+ A+D++ W
Sbjct: 541 E--MKYVIGLNIDFQGAFDNLGW 561
>AI2M_YEAST (P03876) Putative COX1/OXI3 intron 2 protein
Length = 789
Score = 32.7 bits (73), Expect = 3.8
Identities = 37/177 (20%), Positives = 72/177 (39%), Gaps = 21/177 (11%)
Query: 396 IPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGRFMLDSVV 455
+ +VE P+ G +RP+S+ KIV + + M L + + F L +++
Sbjct: 278 VRRVEIPKTSGGFRPLSVGNPREKIVQESMRMMLEIIYNNSFSYYSHGFRPNLSCLTAII 337
Query: 456 VANEVVEEAKRRKKEC-FVFKVDFEKAYDSVDWNFLFYMMS-RI---GFSHKWIHWIKGC 510
+ K + C + KVD K +D++ N L +++ RI GF ++
Sbjct: 338 -------QCKNYMQYCNWFIKVDLNKCFDTIPHNMLINVLNERIKDKGFMDLLYKLLRAG 390
Query: 511 IQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTG 567
+++ N G+ QG ++P L I + L+ + + + TG
Sbjct: 391 YVDKNNNY---------HNTTLGIPQGSVVSPILCNIFLDKLDKYLENKFENEFNTG 438
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.341 0.150 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 84,076,379
Number of Sequences: 164201
Number of extensions: 3294122
Number of successful extensions: 8739
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 13
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 8701
Number of HSP's gapped (non-prelim): 20
length of query: 787
length of database: 59,974,054
effective HSP length: 118
effective length of query: 669
effective length of database: 40,598,336
effective search space: 27160286784
effective search space used: 27160286784
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0263.7


