

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
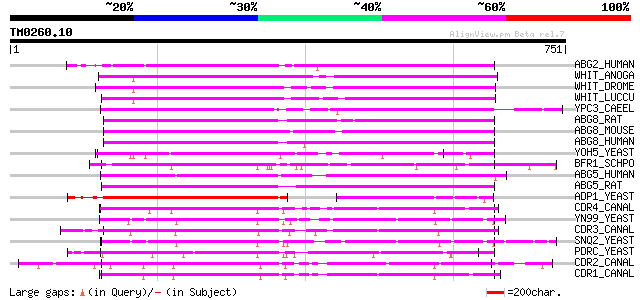
Query= TM0260.10
(751 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
ABG2_HUMAN (Q9UNQ0) ATP-binding cassette, sub-family G, member 2... 258 5e-68
WHIT_ANOGA (Q27256) White protein 249 1e-65
WHIT_DROME (P10090) White protein 242 2e-63
WHIT_LUCCU (Q05360) White protein 242 3e-63
YPC3_CAEEL (Q11180) Putative ABC transporter C05D10.3 in chromos... 238 6e-62
ABG8_RAT (P58428) ATP-binding cassette, sub-family G, member 8 (... 221 4e-57
ABG8_MOUSE (Q9DBM0) ATP-binding cassette, sub-family G, member 8... 219 2e-56
ABG8_HUMAN (Q9H221) ATP-binding cassette, sub-family G, member 8... 211 6e-54
YOH5_YEAST (Q08234) Probable ATP-dependent transporter YOL074C/Y... 208 5e-53
BFR1_SCHPO (P41820) Brefeldin A resistance protein 207 1e-52
ABG5_HUMAN (Q9H222) ATP-binding cassette, sub-family G, member 5... 206 1e-52
ABG5_RAT (Q99PE7) ATP-binding cassette, sub-family G, member 5 (... 202 3e-51
ADP1_YEAST (P25371) Probable ATP-dependent permease precursor 199 2e-50
CDR4_CANAL (O74676) ABC transporter CDR4 199 3e-50
YN99_YEAST (P53756) Probable ATP-dependent transporter YNR070W 191 5e-48
CDR3_CANAL (O42690) Opaque-specific ABC transporter CDR3 186 1e-46
SNQ2_YEAST (P32568) SNQ2 protein 180 1e-44
PDRC_YEAST (Q02785) ATP-dependent permease PDR12 177 1e-43
CDR2_CANAL (P78595) Multidrug resistance protein CDR2 176 2e-43
CDR1_CANAL (P43071) Multidrug resistance protein CDR1 175 3e-43
>ABG2_HUMAN (Q9UNQ0) ATP-binding cassette, sub-family G, member 2
(Placenta-specific ATP-binding cassette transporter)
(Breast cancer resistance protein)
Length = 655
Score = 258 bits (658), Expect = 5e-68
Identities = 175/585 (29%), Positives = 296/585 (49%), Gaps = 41/585 (7%)
Query: 78 VLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLLNDISGEARD 137
VLSF N+ Y VK+K S P R+ VE K +L++I+G +
Sbjct: 36 VLSFHNICYRVKLK-----SGFLPCRK----------PVE-------KEILSNINGIMKP 73
Query: 138 GEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVISAYVMQDDLLF 197
G + A+LG +G GKS+L+D LA R L G + +NG + K S YV+QDD++
Sbjct: 74 G-LNAILGPTGGGKSSLLDVLAARKDPSGLSGDVLINGAPRPANF-KCNSGYVVQDDVVM 131
Query: 198 PMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGER 257
LTV E L F+A RL T++ +K R+ +I++LGL A + +G + RGVSGGER
Sbjct: 132 GTLTVRENLQFSAALRLATTMTNHEKNERINRVIEELGLDKVADSKVGTQFIRGVSGGER 191
Query: 258 RRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPSYRILG 317
+R SIG+++I DP +LFLDEPT+GLDS++A V+ +L+R+++ G +I SIHQP Y I
Sbjct: 192 KRTSIGMELITDPSILFLDEPTTGLDSSTANAVLLLLKRMSKQGRTIIFSIHQPRYSIFK 251
Query: 318 LLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGGTKSL 377
L D + L+ G+ ++ G + +F G+ +N +F LD+I G +
Sbjct: 252 LFDSLTLLASGRLMFHGPAQEALGYFESAGYHCEAYNNPADFFLDII-------NGDSTA 304
Query: 378 VEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEA-ISASI---SRGKL--VSGATNTATTT 431
V N+ + K S+ ++P L E +++S ++ +L +SG T
Sbjct: 305 VALNR--EEDFKATEIIEPSKQDKPLIEKLAEIYVNSSFYKETKAELHQLSGGEKKKKIT 362
Query: 432 PSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPK 491
+ ++ F ++ +SKRSF + P+ ++ +V G ++ +++ L N
Sbjct: 363 VFKEI-SYTTSFCHQLRWVSKRSFKNLLGNPQASIAQIIVTVVLGLVIGAIYFGLKNDST 421
Query: 492 GVQERLGFFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPL 551
G+Q R G F + +++ A+ +F+ E+ +F+ E YR SSY + L L P+
Sbjct: 422 GIQNRAGVLFFLTTNQCFSSVSAVELFVVEKKLFIHEYISGYYRVSSYFLGKLLSDLLPM 481
Query: 552 AFL-SLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIV 610
L S+ F I ++ +GL A F + ++ +S ++ V + ++
Sbjct: 482 RMLPSIIFTCIVYFMLGLKPKADAFFVMMFTLMMVAYSASSMALAIAAGQSVVSVATLLM 541
Query: 611 VAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEF 655
+ ++ SG ++ I S+ W Y S+ +Y + A+ NEF
Sbjct: 542 TICFVFMMIFSGLLVNLTTIASWLSWLQYFSIPRYGFTALQHNEF 586
>WHIT_ANOGA (Q27256) White protein
Length = 695
Score = 249 bits (637), Expect = 1e-65
Identities = 159/544 (29%), Positives = 270/544 (49%), Gaps = 22/544 (4%)
Query: 121 FSRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKG---RLKGTLALNGEA 177
F+ K LL +++G A+ GE++AV+G+SG+GK+TL++ALA R G ALNG
Sbjct: 109 FNPRKHLLKNVTGVAKSGELLAVMGSSGAGKTTLLNALAFRSPPGVKISPNAVRALNGVP 168
Query: 178 LESRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLR 237
+ + L+ AYV QDDL P LT E L F A R+ R + S K+ RVQ ++ +L L
Sbjct: 169 VNAEQLRARCAYVQQDDLFIPSLTTREHLLFQAMLRMGRDVPASVKQHRVQEVLQELSLV 228
Query: 238 NAAKTVIGDEGH-RGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQR 296
A T+IG G +G+SGGER+R++ + + DP LL DEPTSGLDS A V++VL+
Sbjct: 229 KCADTIIGAPGRIKGLSGGERKRLAFASETLTDPHLLLCDEPTSGLDSFMAHSVLQVLKG 288
Query: 297 IAQSGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNR 356
+A G +I++IHQPS + L D+++ ++ G+ + GSP Q FF++ G P P + N
Sbjct: 289 MAMKGKTIILTIHQPSSELYCLFDKILLVAEGRVAFLGSPYQSAEFFSQLGIPCPPNYNP 348
Query: 357 TEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASIS 416
+F + ++ + +++ +++ + + + GM +
Sbjct: 349 ADFYVQMLAIAPAKEAECRDMIKKICDSFAVSPIAREVLETASVAGKGMDEPYML----- 403
Query: 417 RGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTG 476
+ V G +T + + +W + + RS+ + P L +RL +
Sbjct: 404 --QQVEGVGSTG----------YRSSWWTQFYCILWRSWLSVLKDPMLVKVRLLQTAMVA 451
Query: 477 FILATMFWNLDNSPKGVQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYR 535
++ ++++ GV G F F + TF + VF E +F+RE YR
Sbjct: 452 TLIGSIYFGQVLDQDGVMNINGSLFLFLTNMTFQNVFAVINVFSAELPVFLREKRSRLYR 511
Query: 536 RSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTF 595
+Y + + LP + F IT+ +GL GA+ +L I+ SF
Sbjct: 512 VDTYFLGKTIAELPLFIAVPFVFTSITYPMIGLRTGATHYLTTLFIVTLVANVSTSFGYL 571
Query: 596 LSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEF 655
+S + + ++ ++ FL+ GFF++ +P+Y+ + YLS +Y EA+L N++
Sbjct: 572 ISCASSSISMALSVGPPVVIPFLIFGGFFLNSASVPAYFKYLSYLSWFRYANEALLINQW 631
Query: 656 DDPV 659
V
Sbjct: 632 STVV 635
>WHIT_DROME (P10090) White protein
Length = 687
Score = 242 bits (618), Expect = 2e-63
Identities = 161/546 (29%), Positives = 269/546 (48%), Gaps = 28/546 (5%)
Query: 117 EESVFSRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKG---RLKGTLAL 173
E + + K LL ++ G A GE++AV+G+SG+GK+TL++ALA R +G G L
Sbjct: 102 ERHIPAPRKHLLKNVCGVAYPGELLAVMGSSGAGKTTLLNALAFRSPQGIQVSPSGMRLL 161
Query: 174 NGEALESRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQ 233
NG+ ++++ ++ AYV QDDL LT E L F A R+PR L+ ++ ARV +I +
Sbjct: 162 NGQPVDAKEMQARCAYVQQDDLFIGSLTAREHLIFQAMVRMPRHLTYRQRVARVDQVIQE 221
Query: 234 LGLRNAAKTVIGDEGH-RGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVK 292
L L T+IG G +G+SGGER+R++ + + DP LL DEPTSGLDS +A VV+
Sbjct: 222 LSLSKCQHTIIGVPGRVKGLSGGERKRLAFASEALTDPPLLICDEPTSGLDSFTAHSVVQ 281
Query: 293 VLQRIAQSGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPD 352
VL++++Q G VI++IHQPS + L D+++ ++ G+ + G+P++ FF+ G P
Sbjct: 282 VLKKLSQKGKTVILTIHQPSSELFELFDKILLMAEGRVAFLGTPSEAVDFFSYVGAQCPT 341
Query: 353 SDNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAIS 412
+ N +F + ++ + G +S + K+ + S+ R E +
Sbjct: 342 NYNPADFYVQVLAVVPGRE---------IESRDRIAKICDNFAISKVARD-----MEQLL 387
Query: 413 ASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAV 472
A+ + K + N T+ ++++ + RS+ ++P L +RL
Sbjct: 388 ATKNLEKPLEQPENGY---------TYKATWFMQFRAVLWRSWLSVLKEPLLVKVRLIQT 438
Query: 473 MVTGFILATMFWNLDNSPKGVQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAY 531
+ ++ +F + GV G F F + TF + VF E +FMRE
Sbjct: 439 TMVAILIGLIFLGQQLTQVGVMNINGAIFLFLTNMTFQNVFATINVFTSELPVFMREARS 498
Query: 532 NAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNS 591
YR +Y + + LP + L F I + +GL G F ++ S
Sbjct: 499 RLYRCDTYFLGKTIAELPLFLTVPLVFTAIAYPMIGLRAGVLHFFNCLALVTLVANVSTS 558
Query: 592 FVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVL 651
F +S + ++ ++ FLL GFF++ +P Y W YLS +Y E +L
Sbjct: 559 FGYLISCASSSTSMALSVGPPVIIPFLLFGGFFLNSGSVPVYLKWLSYLSWFRYANEGLL 618
Query: 652 QNEFDD 657
N++ D
Sbjct: 619 INQWAD 624
>WHIT_LUCCU (Q05360) White protein
Length = 677
Score = 242 bits (617), Expect = 3e-63
Identities = 159/538 (29%), Positives = 261/538 (47%), Gaps = 26/538 (4%)
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKG---RLKGTLALNGEALESR 181
K L+ ++ G A GE++AV+G+SG+GK+TL++ALA R A+G LNG ++++
Sbjct: 99 KHLIKNVCGVAYPGELLAVMGSSGAGKTTLLNALAFRSARGVQISPSSVRMLNGHPVDAK 158
Query: 182 LLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAK 241
++ AYV QDDL LT E L F A R+PRT+++ +K RV +I L L
Sbjct: 159 EMQARCAYVQQDDLFIGSLTAREHLIFQATVRMPRTMTQKQKLQRVDQVIQDLSLIKCQN 218
Query: 242 TVIGDEGH-RGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQS 300
T+IG G +G+SGGER+R++ + + DP LL DEPTSGLDS A VV+VL++++Q
Sbjct: 219 TIIGVPGRVKGLSGGERKRLAFASEALTDPPLLICDEPTSGLDSFMAASVVQVLKKLSQR 278
Query: 301 GSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFA 360
G VI++IHQPS + L D+++ ++ G+ + G+P + FF+ G P + N +F
Sbjct: 279 GKTVILTIHQPSSELFELFDKILLMAEGRVAFLGTPVEAVDFFSFIGAQCPTNYNPADFY 338
Query: 361 LDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGKL 420
+ ++ + G +S ++K+ + + R + ++ + + K
Sbjct: 339 VQVLAVVPGRE---------IESRDRISKICDNFAVGKVSREMEQNFQKIAAKTDGLQK- 388
Query: 421 VSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILA 480
T +S F W RS+ + ++P L +RL + ++
Sbjct: 389 ---DDETTILYKASWFTQFRAIMW--------RSWISTLKEPLLVKVRLIQTTMVAVLIG 437
Query: 481 TMFWNLDNSPKGVQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSY 539
+F N + GV G F F + TF + VF E +FMRET YR +Y
Sbjct: 438 LIFLNQPMTQVGVMNINGAIFLFLTNMTFQNVFAVINVFTSELPVFMRETRSRLYRCDTY 497
Query: 540 LISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGV 599
+ L LP + F I + +GL G + FL ++ SF +S
Sbjct: 498 FLGKTLAELPLFLVVPFLFIAIAYPMIGLRPGITHFLSALALVTLVANVSTSFGYLISCA 557
Query: 600 VPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDD 657
+ ++ + FLL G F++ +P Y+ W Y S +Y E +L N++ D
Sbjct: 558 STSTSMALSVGPPLTIPFLLFGGVFLNSGSVPVYFKWLSYFSWFRYANEGLLINQWAD 615
>YPC3_CAEEL (Q11180) Putative ABC transporter C05D10.3 in chromosome
III
Length = 598
Score = 238 bits (606), Expect = 6e-62
Identities = 177/639 (27%), Positives = 307/639 (47%), Gaps = 60/639 (9%)
Query: 123 RTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANR-IAKGRLKGTLALNGEALESR 181
R K +L+++SG A G+++A+LG+SG+GK+TL++ L +R + ++G++ ++G
Sbjct: 5 RVKEILHNVSGMAESGKLLAILGSSGAGKTTLMNVLTSRNLTNLDVQGSILIDGRRANKW 64
Query: 182 LLKVISAYVMQDDLLFPMLTVEETLTFAAEFRL-PRTLSKSKKKARVQALIDQLGLRNAA 240
++ +SA+V Q D+ +T E L F A R+ + S +++ RV+ ++ Q+GL+ A
Sbjct: 65 KIREMSAFVQQHDMFVGTMTAREHLQFMARLRMGDQYYSDHERQLRVEQVLTQMGLKKCA 124
Query: 241 KTVIGDEGH-RGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQ 299
TVIG +G+S GE++R+S +I+ P +LF DEPTSGLD+ A VV+ L+ +A
Sbjct: 125 DTVIGIPNQLKGLSCGEKKRLSFASEILTCPKILFCDEPTSGLDAFMAGHVVQALRSLAD 184
Query: 300 SGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEF 359
+G VI++IHQPS + L + + ++ G+ +Y G Q F + G+P P N +
Sbjct: 185 NGMTVIITIHQPSSHVYSLFNNVCLMACGRVIYLGPGDQAVPLFEKCGYPCPAYYNPAD- 243
Query: 360 ALDLIRDLEGSPGGTKSLVEFNK--SWQSMTKVHSHSVSSQPERPNGMSLKEAISASISR 417
LIR T ++++ ++ S ++++K+ +S+ L +++ A +
Sbjct: 244 --HLIR--------TLAVIDSDRATSMKTISKIRQGFLST--------DLGQSVLAIGNA 285
Query: 418 GKLVSGATNTATTTPSSMVPTFAN-----PFWIEMVTLSKRSFTDSRRKPELFGIRLGAV 472
KL + + T + T S TF N FW + + L RS+ R P L +RL +
Sbjct: 286 NKLRAASFVTGSDT-SEKTKTFFNQDYNASFWTQFLALFWRSWLTVIRDPNLLSVRLLQI 344
Query: 473 MVTGFILATMFWNLDNSPKGVQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAY 531
++T FI +F+ +P + G F + F +PV E I +RE A
Sbjct: 345 LITAFITGIVFFQTPVTPATIISINGIMFNHIRNMNFMLQFPNVPVITAELPIVLRENAN 404
Query: 532 NAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNS 591
YR S+Y ++ + LP L + + I +W GL + F L+ S
Sbjct: 405 GVYRTSAYFLAKNIAELPQYIILPILYNTIVYWMSGLYPNFWNYCFASLVTILITNVAIS 464
Query: 592 FVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVL 651
++ + + + TI+ + + GFFI D IPSY+ W LS KY YEA+
Sbjct: 465 ISYAVATIFANTDVAMTILPIFVVPIMAFGGFFITFDAIPSYFKWLSSLSYFKYGYEALA 524
Query: 652 QNEFDDPVKCFVRGVQIFDNSPLRSVPYELKLKLLGSMSGTLGTNITATTCLTTGADILQ 711
NE+D +K++ + T +C G +L+
Sbjct: 525 INEWD-------------------------SIKVIPECFNSSMTAFALDSCPKNGHQVLE 559
Query: 712 QNGVTELSKWNCLWITVAWGFF--FRFLFYLALLVGSKN 748
+ K I++ +G F R + Y+ALL+ S N
Sbjct: 560 SIDFSASHK--IFDISILFGMFIGIRIIAYVALLIRSYN 596
>ABG8_RAT (P58428) ATP-binding cassette, sub-family G, member 8
(Sterolin-2)
Length = 694
Score = 221 bits (564), Expect = 4e-57
Identities = 150/537 (27%), Positives = 263/537 (48%), Gaps = 24/537 (4%)
Query: 128 LNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLK-GTLALNGEALESRLLKVI 186
+ ++S + R G+++A++G++G G++TL+D + R G++K G + +NG+ +L++
Sbjct: 110 IRNLSFKVRSGQMLAIIGSAGCGRATLLDVITGRDHGGKMKSGQIWINGQPSTPQLIQKC 169
Query: 187 SAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGD 246
A+V Q D L P LTV ETLTF A+ RLP+T S++++ RV+ +I +L LR A T +G+
Sbjct: 170 VAHVRQQDQLLPNLTVRETLTFIAQMRLPKTFSQAQRDKRVEDVIAELRLRQCANTRVGN 229
Query: 247 EGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIM 306
RGVSGGERRRVSIG+ ++ +P +L LDEPTSGLDS +A +V+ L R+A+ +V++
Sbjct: 230 TYVRGVSGGERRRVSIGVQLLWNPGILILDEPTSGLDSFTAHNLVRTLSRLAKGNRLVLI 289
Query: 307 SIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRD 366
S+HQP I L D ++ ++ G +Y G + +F G+P P N +F +DL
Sbjct: 290 SLHQPRSDIFRLFDLVLLMTSGTPIYLGVAQHMVQYFTSIGYPCPRYSNPADFYVDL--- 346
Query: 367 LEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQ-PERPNGMS--LKEAISASISRGKLVSG 423
S+ +K + T + +++ E+ G L +A + S+ G
Sbjct: 347 --------TSIDRRSKEQEVATMEKARLLAALFLEKVQGFDDFLWKAEAKSLDTGTYAVS 398
Query: 424 ATNTATTT--PSSMVPTFANPFWIEMVTLSKRSFT-DSRRKPELFGIRLGAVMVTGFILA 480
T T T ++ +P F TL +R + D R P LF I + I+
Sbjct: 399 QTLTQDTNCGTAAELPGMIQQF----TTLIRRQISNDFRDLPTLF-IHGAEACLMSLIIG 453
Query: 481 TMFWNLDNSPKGVQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSY 539
+++ + P + F F D + ER + E Y Y
Sbjct: 454 FLYYGHADKPLSFMDMAALLFMIGALIPFNVILDVVSKCHSERSLLYYELEDGLYTAGPY 513
Query: 540 LISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGV 599
+ L LP + + + +W L G FL +F++++ + + S +
Sbjct: 514 FFAKVLGELPEHCAYVIIYGMPIYWLTNLRPGPELFLLHFMLLWLVVFCCRTMALAASAM 573
Query: 600 VPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFD 656
+P + A+ F L +GF I+ + + W +S +++ + ++Q +F+
Sbjct: 574 LPTFHMSSFCCNALYNSFYLTAGFMINLNNLWIVPAWISKMSFLRWCFSGLMQIQFN 630
>ABG8_MOUSE (Q9DBM0) ATP-binding cassette, sub-family G, member 8
(Sterolin-2)
Length = 673
Score = 219 bits (558), Expect = 2e-56
Identities = 148/532 (27%), Positives = 255/532 (47%), Gaps = 14/532 (2%)
Query: 128 LNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLK-GTLALNGEALESRLLKVI 186
+ ++S + R G+++A++G+SG G+++L+D + R G++K G + +NG+ +L++
Sbjct: 89 IRNLSFKVRSGQMLAIIGSSGCGRASLLDVITGRGHGGKMKSGQIWINGQPSTPQLVRKC 148
Query: 187 SAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGD 246
A+V Q D L P LTV ETL F A+ RLPRT S++++ RV+ +I +L LR A T +G+
Sbjct: 149 VAHVRQHDQLLPNLTVRETLAFIAQMRLPRTFSQAQRDKRVEDVIAELRLRQCANTRVGN 208
Query: 247 EGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIM 306
RGVSGGERRRVSIG+ ++ +P +L LDEPTSGLDS +A +V L R+A+ +V++
Sbjct: 209 TYVRGVSGGERRRVSIGVQLLWNPGILILDEPTSGLDSFTAHNLVTTLSRLAKGNRLVLI 268
Query: 307 SIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRD 366
S+HQP I L D ++ ++ G +Y G+ Q+ +F GHP P N +F +DL
Sbjct: 269 SLHQPRSDIFRLFDLVLLMTSGTPIYLGAAQQMVQYFTSIGHPCPRYSNPADFYVDLTSI 328
Query: 367 LEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGKLVSGATN 426
S + VE QS+ + V + KE +++ + ++ T+
Sbjct: 329 DRRSKEREVATVE---KAQSLAALFLEKVQGFDDFLWKAEAKELNTSTHTVSLTLTQDTD 385
Query: 427 TATTTP-SSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWN 485
T M+ F+ TL +R ++ R I + I+ +++
Sbjct: 386 CGTAVELPGMIEQFS--------TLIRRQISNDFRDLPTLLIHGSEACLMSLIIGFLYYG 437
Query: 486 LDNSPKGVQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHA 544
+ F F D + ER + E Y Y +
Sbjct: 438 HGAKQLSFMDTAALLFMIGALIPFNVILDVVSKCHSERSMLYYELEDGLYTAGPYFFAKI 497
Query: 545 LVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVM 604
L LP + +A+ +W L FL +FL+++ + + S ++P
Sbjct: 498 LGELPEHCAYVIIYAMPIYWLTNLRPVPELFLLHFLLVWLVVFCCRTMALAASAMLPTFH 557
Query: 605 LGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFD 656
+ A+ F L +GF I+ D + W LS +++ + ++Q +F+
Sbjct: 558 MSSFFCNALYNSFYLTAGFMINLDNLWIVPAWISKLSFLRWCFSGLMQIQFN 609
>ABG8_HUMAN (Q9H221) ATP-binding cassette, sub-family G, member 8
(Sterolin-2)
Length = 673
Score = 211 bits (537), Expect = 6e-54
Identities = 146/533 (27%), Positives = 257/533 (47%), Gaps = 17/533 (3%)
Query: 128 LNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLK-GTLALNGEALESRLLKVI 186
+ ++S + R G+++A++G+SG G+++L+D + R G++K G + +NG+ +L++
Sbjct: 88 IQNLSFKVRSGQMLAIIGSSGCGRASLLDVITGRGHGGKIKSGQIWINGQPSSPQLVRKC 147
Query: 187 SAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGD 246
A+V Q + L P LTV ETL F A+ RLPRT S++++ RV+ +I +L LR A T +G+
Sbjct: 148 VAHVRQHNQLLPNLTVRETLAFIAQMRLPRTFSQAQRDKRVEDVIAELRLRQCADTRVGN 207
Query: 247 EGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIM 306
RG+SGGERRRVSIG+ ++ +P +L LDEPTSGLDS +A +VK L R+A+ +V++
Sbjct: 208 MYVRGLSGGERRRVSIGVQLLWNPGILILDEPTSGLDSFTAHNLVKTLSRLAKGNRLVLI 267
Query: 307 SIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRD 366
S+HQP I L D ++ ++ G +Y G+ + +F G+P P N +F +DL
Sbjct: 268 SLHQPRSDIFRLFDLVLLMTSGTPIYLGAAQHMVQYFTAIGYPCPRYSNPADFYVDL--- 324
Query: 367 LEGSPGGTKSLVEFNKSWQSMTKVHSHSVSS---QPERPNGMSLKEAISASISRGKLVSG 423
S+ ++ + T+ + S+++ + R L +A + + V
Sbjct: 325 --------TSIDRRSREQELATREKAQSLAALFLEKVRDLDDFLWKAETKDLDEDTCVES 376
Query: 424 ATNTATTTPSSMVPTFANPFWIEMVTLSKRSFT-DSRRKPELFGIRLGAVMVTGFILATM 482
+ T T PT + TL +R + D R P L A +++ I
Sbjct: 377 SV-TPLDTNCLPSPTKMPGAVQQFTTLIRRQISNDFRDLPTLLIHGAEACLMSMTIGFLY 435
Query: 483 FWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLIS 542
F + + F F D + ER + E Y Y +
Sbjct: 436 FGHGSIQLSFMDTAALLFMIGALIPFNVILDVISKCYSERAMLYYELEDGLYTTGPYFFA 495
Query: 543 HALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPH 602
L LP + + + T+W L G FL +FL+++ + + ++P
Sbjct: 496 KILGELPEHCAYIIIYGMPTYWLANLRPGLQPFLLHFLLVWLVVFCCRIMALAAAALLPT 555
Query: 603 VMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEF 655
+ A+ F L GF I+ + + W +S +++ +E +++ +F
Sbjct: 556 FHMASFFSNALYNSFYLAGGFMINLSSLWTVPAWISKVSFLRWCFEGLMKIQF 608
>YOH5_YEAST (Q08234) Probable ATP-dependent transporter
YOL074C/YOL075C
Length = 1294
Score = 208 bits (529), Expect = 5e-53
Identities = 162/571 (28%), Positives = 272/571 (47%), Gaps = 69/571 (12%)
Query: 120 VFSRTKT-LLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGR---------LKG 169
V S+T T L+N S + G +MAV+G SGSGK+TL++ LA++I+ G L+
Sbjct: 36 VASKTNTTLVNTFSMDLPSGSVMAVMGGSGSGKTTLLNVLASKISGGLTHNGSIRYVLED 95
Query: 170 TLALNGEALESRL----------LKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLS 219
T + E R VI AY+ Q D+L P LT ETL FAA+ +L S
Sbjct: 96 TGSEPNETEPKRAHLDGQDHPIQKHVIMAYLPQQDVLSPRLTCRETLKFAADLKL--NSS 153
Query: 220 KSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPT 279
+ KK V+ LI++LGL++ A T++GD HRG+SGGE+RR+SIG +I +P ++FLDEPT
Sbjct: 154 ERTKKLMVEQLIEELGLKDCADTLVGDNSHRGLSGGEKRRLSIGTQMISNPSIMFLDEPT 213
Query: 280 SGLDSTSAFMVVKVLQRIA-QSGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQ 338
+GLD+ SAF+V+K L+++A + G IMSIHQP IL LLD++ LS+G VY
Sbjct: 214 TGLDAYSAFLVIKTLKKLAKEDGRTFIMSIHQPRSDILFLLDQVCILSKGNVVYCDKMDN 273
Query: 339 LPSFFAEFGHPLPDSDNRTEFALDLI-------RDLEGSPGGTKSLVEFNKSWQSMTKVH 391
+F G+ +P N ++ +DL ++ + SL++ W + H
Sbjct: 274 TIPYFESIGYHVPQLVNPADYFIDLSSVDSRSDKEEAATQSRLNSLID---HWHDYERTH 330
Query: 392 SHSVSSQPERPNGMSLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLS 451
+ +S AT ++ + PFW ++ L+
Sbjct: 331 ---------------------LQLQAESYISNATEIQIQNMTTRL-----PFWKQVTVLT 364
Query: 452 KRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFA---MSTTF 508
+R+F + ++ G + +++ D S G +A +
Sbjct: 365 RRNFKLNFSDYVTLISTFAEPLIIGTVCGWIYYKPDKSSIGGLRTTTACLYASTILQCYL 424
Query: 509 YTTADALPVFLQERYIFMRETAYNAYRRSSYLISHAL-VSLPPLAFLSLAFAVITFWAVG 567
Y D + Q+ ++ RE A + +++++ + + L +++ F IT++ G
Sbjct: 425 YLLFDTYRLCEQDIALYDRERAEGSVTPLAFIVARKISLFLSDDFAMTMIFVSITYFMFG 484
Query: 568 LDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLS---GFF 624
L+ A F + F ++F S ++ LS V ++V + F +LS GFF
Sbjct: 485 LEADARKFFYQFAVVFLC-QLSCSGLSMLSVAVSRDFSKASLVGNMT--FTVLSMGCGFF 541
Query: 625 IDRDRIPSYWIWFHYLSLVKYPYEAVLQNEF 655
++ +P Y W Y++ Y + ++ + F
Sbjct: 542 VNAKVMPVYVRWIKYIAFTWYSFGTLMSSTF 572
Score = 199 bits (505), Expect = 3e-50
Identities = 137/482 (28%), Positives = 243/482 (49%), Gaps = 36/482 (7%)
Query: 117 EESVFSRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRI-----AKGRLKGTL 171
E + TK +L ++ + G I A++G SGSGKS+L++ ++ R+ AK G++
Sbjct: 699 EGNFHHETKEILQSVNAIFKPGMINAIMGPSGSGKSSLLNLISGRLKSSVFAKFDTSGSI 758
Query: 172 ALNGEALESRLLKVISAYVMQDD-LLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQAL 230
N + + K + +YV QDD L LTV+ETL +AA RL L+++++ R L
Sbjct: 759 MFNDIQVSELMFKNVCSYVSQDDDHLLAALTVKETLKYAAALRLHH-LTEAERMERTDNL 817
Query: 231 IDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMV 290
I LGL++ +IG+E +G+SGGE+RRV++G+ +++DP +L LDEPTSGLDS ++ +
Sbjct: 818 IRSLGLKHCENNIIGNEFVKGISGGEKRRVTMGVQLLNDPPILLLDEPTSGLDSFTSATI 877
Query: 291 VKVLQRIA-QSGSIVIMSIHQPSYRILGLLDRMIFLSR-GQTVYSGSPTQLPSFFAEFGH 348
+++L+++ + G +I++IHQP + ++ L++ G+T ++GSP ++ ++F E G+
Sbjct: 878 LEILEKLCREQGKTIIITIHQPRSELFKRFGNVLLLAKSGRTAFNGSPDEMIAYFTELGY 937
Query: 349 PLPDSDNRTEFALDLI----RDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNG 404
P N +F LDLI ++ + + + +W++ + + S+S P
Sbjct: 938 NCPSFTNVADFFLDLISVNTQNEQNEISSRARVEKILSAWKA--NMDNESLSPTPISEKQ 995
Query: 405 MSLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPEL 464
+E+ S P+++V + + KR FT +RR +
Sbjct: 996 QYSQESFFTEYSE----------FVRKPANLV--------LAYIVNVKRQFTTTRRSFDS 1037
Query: 465 FGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERYI 524
R+ + G I A F + ++ + RLG + + F L + ER
Sbjct: 1038 LMARIAQIPGLGVIFALFFAPVKHNYTSISNRLGLAQESTALYFVGMLGNLACYPTERDY 1097
Query: 525 FMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFA 584
F E N Y + + +++ + LP A S+ +AV T A GL A F F ++
Sbjct: 1098 FYEEYNDNVYGIAPFFLAYMTLELPLSALASVLYAVFTVLACGLPRTAGNF---FATVYC 1154
Query: 585 SF 586
SF
Sbjct: 1155 SF 1156
>BFR1_SCHPO (P41820) Brefeldin A resistance protein
Length = 1530
Score = 207 bits (526), Expect = 1e-52
Identities = 171/643 (26%), Positives = 295/643 (45%), Gaps = 56/643 (8%)
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLK 184
+ LLN + G G++ A++G SG+GK+TL++ LA R+ G + G + +NG L+S +
Sbjct: 898 RRLLNGVQGFVVPGKLTALMGESGAGKTTLLNVLAQRVDTGVVTGDMLVNGRGLDSTFQR 957
Query: 185 VISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVI 244
+ YV Q D+ TV E L F+A R P ++ S+K V+++I L + + A+ +I
Sbjct: 958 R-TGYVQQQDVHIGESTVREALRFSAALRQPASVPLSEKYEYVESVIKLLEMESYAEAII 1016
Query: 245 GDEGHRGVSGGERRRVSIGIDIIHDP-ILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSI 303
G G G++ +R+R +IG+++ P +LLFLDEPTSGLDS SA+ +V L+++A +G
Sbjct: 1017 GTPGS-GLNVEQRKRATIGVELAAKPALLLFLDEPTSGLDSQSAWSIVCFLRKLADAGQA 1075
Query: 304 VIMSIHQPSYRILGLLDRMIFLSR-GQTVYSG-----SPTQLPSFFAEFGHPLPDSDNRT 357
++ +IHQPS + DR++ L + G+TVY G S T L F + PD N
Sbjct: 1076 ILCTIHQPSAVLFDQFDRLLLLQKGGKTVYFGDIGEHSKTLLNYFESHGAVHCPDDGNPA 1135
Query: 358 EFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISR 417
E+ LD+I G + N+ W H V + E ++AISA + +
Sbjct: 1136 EYILDVI--------GAGATATTNRDW--------HEVWNNSEE------RKAISAELDK 1173
Query: 418 GKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGF 477
+ T T+A P W ++ + R+F R+P + +L + G
Sbjct: 1174 INASFSNSEDKKTLSKEDRSTYAMPLWFQVKMVMTRNFQSYWREPSILMSKLALDIFAGL 1233
Query: 478 ILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTA--DALPVFLQERYIF-MRETAYNAY 534
+ F+N + +Q +L FA M+T P F++ R +F +RE N Y
Sbjct: 1234 FIGFTFYNQGLGVQNIQNKL--FAVFMATVLAVPLINGLQPKFIELRNVFEVREKPSNIY 1291
Query: 535 RRSSYLISHALVSLP------PLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWA 588
+++ S +V +P L FL + + + + G +G+ + + F ++
Sbjct: 1292 SWVAFVFSAIIVEIPFNLVFGTLFFLCWFYPIKFYKHIHHPGDKTGYAWLLYMFFQMYF- 1350
Query: 589 GNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYE 648
++F ++ P+ + + + + +G + +W W H L+ Y E
Sbjct: 1351 -STFGQAVASACPNAQTASVVNSLLFTFVITFNGVLQPNSNLVGFWHWMHSLTPFTYLIE 1409
Query: 649 AVLQNEFDD-PVKCFVRGVQIFDNSPLRSVPYELKLKLL--GSMSGTLGTNITATTC--- 702
+L + PV+C + + N P E L + +G L T+C
Sbjct: 1410 GLLSDLVHGLPVECKSHEM-LTINPPSGQTCGEYMSAFLTNNTAAGNLLNPNATTSCSYC 1468
Query: 703 -LTTGADILQQNGVTELSKWNCLWITVAWGFFFRF----LFYL 740
T L++ + +W L I V + FF F LFY+
Sbjct: 1469 PYQTADQFLERFSMRYTHRWRNLGIFVGYVFFNIFAVLLLFYV 1511
Score = 159 bits (403), Expect = 2e-38
Identities = 155/589 (26%), Positives = 267/589 (45%), Gaps = 64/589 (10%)
Query: 109 AVEDAPAVEESVFSRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDAL-ANRIAKGRL 167
A+ + VE+++ S L N GE++ VLG GSG ST + ++ ++ + R+
Sbjct: 165 AITEKQVVEKAILSHCHALANA-------GELVMVLGQPGSGCSTFLRSVTSDTVHYKRV 217
Query: 168 KGTLALNG--EALESRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRT----LSKS 221
+GT +G +A + Y ++D+ FP LT ETL FAA+ R P L++
Sbjct: 218 EGTTHYDGIDKADMKKFFPGDLLYSGENDVHFPSLTTAETLDFAAKCRTPNNRPCNLTRQ 277
Query: 222 KKKARVQALI-DQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTS 280
+ +R + LI GL + T +G++ RGVSGGER+RV+I P + D T
Sbjct: 278 EYVSRERHLIATAFGLTHTFNTKVGNDFVRGVSGGERKRVTISEGFATRPTIACWDNSTR 337
Query: 281 GLDSTSAFMVVKVLQRIAQSGSIV-IMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQL 339
GLDS++AF V VL+ A + ++ +Q S +I L DR+ L G+ +Y G +
Sbjct: 338 GLDSSTAFEFVNVLRTCANELKMTSFVTAYQASEKIYKLFDRICVLYAGRQIYYGPADKA 397
Query: 340 PSFFAEFG---HP---LPD-----SDNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQS-- 386
+F + G HP PD SD + F + E T EF + W++
Sbjct: 398 KQYFLDMGFDCHPRETTPDFLTAISDPKARFPR---KGFENRVPRTPD--EFEQMWRNSS 452
Query: 387 -----MTKVHSH---------SVSSQPERPN-GMSLKEAISASISRGKLVSGATNTATTT 431
M ++ S+ + S PE+ N G + + R V+ + T
Sbjct: 453 VYADLMAEMESYDKRWTETTPASSEAPEKDNFGSDISATTKHELYRQSAVAEKSKRVKDT 512
Query: 432 PSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPK 491
S TF+ W + +R D P G A + I+ ++F+++ +
Sbjct: 513 -SPYTVTFSQQLWYCLARSWERYIND----PAYIGSMAFAFLFQSLIIGSIFYDMKLNTV 567
Query: 492 GVQERLGFFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPL 551
V R G F++ + + +R I + A Y ++ +IS +V L P
Sbjct: 568 DVFSRGGVLFFSILFCALQSLSEIANMFSQRPIIAKHRASALYHPAADVISSLIVDL-PF 626
Query: 552 AFLSLA-FAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHV----MLG 606
F++++ F+++ ++ L A GF YFL +F ++F L+G++P+V LG
Sbjct: 627 RFINISVFSIVLYFLTNLKRTAGGFWTYFLFLFIGATCMSAFFRSLAGIMPNVESASALG 686
Query: 607 YTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEF 655
V+AI Y +G+ I + ++ W YL +++ +E+++ NEF
Sbjct: 687 GIGVLAIAIY----TGYAIPNIDVGWWFRWIAYLDPLQFGFESLMINEF 731
>ABG5_HUMAN (Q9H222) ATP-binding cassette, sub-family G, member 5
(Sterolin-1)
Length = 651
Score = 206 bits (525), Expect = 1e-52
Identities = 156/570 (27%), Positives = 276/570 (48%), Gaps = 43/570 (7%)
Query: 124 TKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAK-GRLKGTLALNGEALESRL 182
T+ +L D+S G+IM +LG+SGSGK+TL+DA++ R+ + G G + +NG AL
Sbjct: 65 TRQILKDVSLYVESGQIMCILGSSGSGKTTLLDAMSGRLGRAGTFLGEVYVNGRALRREQ 124
Query: 183 LKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKT 242
+ +YV+Q D L LTV ETL + A + R S +K +V+A++ +L L + A
Sbjct: 125 FQDCFSYVLQSDTLLSSLTVRETLHYTALLAIRRGNPGSFQK-KVEAVMAELSLSHVADR 183
Query: 243 VIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGS 302
+IG+ G+S GERRRVSI ++ DP ++ DEPT+GLD +A +V +L +A+
Sbjct: 184 LIGNYSLGGISTGERRRVSIAAQLLQDPKVMLFDEPTTGLDCMTANQIVVLLVELARRNR 243
Query: 303 IVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALD 362
IV+++IHQP + L D++ LS G+ ++ G+P ++ FF + G+P P+ N +F +D
Sbjct: 244 IVVLTIHQPRSELFQLFDKIAILSFGELIFCGTPAEMLDFFNDCGYPCPEHSNPFDFYMD 303
Query: 363 LIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGKLVS 422
L + +E +K Q + + S N +K
Sbjct: 304 L---TSVDTQSKEREIETSKRVQMIESAYKKSAICHKTLKNIERMKH------------- 347
Query: 423 GATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGI-RLGAVMVTG-FILA 480
T P T +P + + R T + + +L I RL ++ G F+L
Sbjct: 348 -----LKTLPMVPFKTKDSPGVFSKLGVLLRRVTRNLVRNKLAVITRLLQNLIMGLFLLF 402
Query: 481 TMFWNLDNSPKG-VQERLG-FFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSS 538
+ N KG +Q+R+G + F +T + +A+ +F R + +E+ Y++
Sbjct: 403 FVLRVRSNVLKGAIQDRVGLLYQFVGATPYTGMLNAVNLFPVLRAVSDQESQDGLYQKWQ 462
Query: 539 YLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSG 598
++++AL LP ++ F+ + +W +GL + F ++ + A G L G
Sbjct: 463 MMLAYALHVLPFSVVATMIFSSVCYWTLGLHPEVARFGYFSAALLAPHLIGEFLTLVLLG 522
Query: 599 VVPHVMLGYTIVVAI-LAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEF-- 655
+V + + ++V + +A L+ SGF + +P + Y + KY E ++ NEF
Sbjct: 523 IVQNPNIVNSVVALLSIAGVLVGSGFLRNIQEMPIPFKIISYFTFQKYCSEILVVNEFYG 582
Query: 656 -------------DDPVKCFVRGVQIFDNS 672
+P+ F +G+Q + +
Sbjct: 583 LNFTCGSSNVSVTTNPMCAFTQGIQFIEKT 612
>ABG5_RAT (Q99PE7) ATP-binding cassette, sub-family G, member 5
(Sterolin-1)
Length = 652
Score = 202 bits (513), Expect = 3e-51
Identities = 153/540 (28%), Positives = 266/540 (48%), Gaps = 34/540 (6%)
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAK-GRLKGTLALNGEALESRLL 183
+ +L D+S G+ M +LG+SGSGK+TL+DA++ R+ + G L+G + +NG L
Sbjct: 67 RKILKDVSLYIESGQTMCILGSSGSGKTTLLDAISGRLRRTGTLEGEVFVNGCELRRDQF 126
Query: 184 KVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTV 243
+ +Y++Q D+ LTV ETL + A L R+ S +V+A++ +L L + A +
Sbjct: 127 QDCVSYLLQSDVFLSSLTVRETLRYTAMLAL-RSSSADFYDKKVEAVLTELSLSHVADQM 185
Query: 244 IGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSI 303
IG+ G+S GERRRVSI ++ DP ++ LDEPT+GLD +A +V +L +A+ I
Sbjct: 186 IGNYNFGGISSGERRRVSIAAQLLQDPKVMMLDEPTTGLDCMTANHIVLLLVELARRNRI 245
Query: 304 VIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDL 363
VI++IHQP + D++ L+ G+ V+ G+P ++ FF G+P P+ N +F +DL
Sbjct: 246 VIVTIHQPRSELFHHFDKIAILTYGELVFCGTPEEMLGFFNNCGYPCPEHSNPFDFYMDL 305
Query: 364 IRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMS-LKEAISASISRGKLVS 422
T V + S + E + L+ A S K++
Sbjct: 306 ------------------------TSVDTQSREREIETYKRVQMLESAFRQSDICHKILE 341
Query: 423 GATNTATTTPSSMVP-TFANP--FWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFIL 479
T MVP NP + ++ L +R + R ++ +RL ++ G L
Sbjct: 342 NIERTRHLKTLPMVPFKTKNPPGMFCKLGVLLRRVTRNLMRNKQVVIMRLVQNLIMGLFL 401
Query: 480 ATMFWNLDNSP-KG-VQERLGFFAFAMSTTFYT-TADALPVFLQERYIFMRETAYNAYRR 536
+ N+ KG VQ+R+G + T YT +A+ +F R + +E+ Y++
Sbjct: 402 IFYLLRVQNNMLKGAVQDRVGLLYQLVGATPYTGMLNAVNLFPMLRAVSDQESQDGLYQK 461
Query: 537 SSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFL 596
L+++ L +LP ++ F+ + +W +GL + F ++ + A G L
Sbjct: 462 WQMLLAYVLHALPFSIVATVIFSSVCYWTLGLYPEVARFGYFSAALLAPHLIGEFLTLVL 521
Query: 597 SGVVPHVMLGYTIVVAI-LAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEF 655
G+V + + +IV + ++ L+ SGF + + +P Y + KY E ++ NEF
Sbjct: 522 LGMVQNPNIVNSIVALLSISGLLIGSGFIRNIEEMPIPLKILGYFTFQKYCCEILVVNEF 581
>ADP1_YEAST (P25371) Probable ATP-dependent permease precursor
Length = 1049
Score = 199 bits (506), Expect = 2e-50
Identities = 114/298 (38%), Positives = 180/298 (60%), Gaps = 30/298 (10%)
Query: 79 LSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLLNDISGEARDG 138
LSF N+TYSV SI ++ VEE T+LN+ISG + G
Sbjct: 384 LSFENITYSV--------PSI------------NSDGVEE-------TVLNEISGIVKPG 416
Query: 139 EIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVISAYVMQDDLLFP 198
+I+A++G SG+GK+TL+D LA + G + G++ +NG +++ + I +V QDD L P
Sbjct: 417 QILAIMGGSGAGKTTLLDILAMKRKTGHVSGSIKVNGISMDRKSFSKIIGFVDQDDFLLP 476
Query: 199 MLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERR 258
LTV ET+ +A RLP+ LS KKARV ++++L + + +IG+E RG+SGGE+R
Sbjct: 477 TLTVFETVLNSALLRLPKALSFEAKKARVYKVLEELRIIDIKDRIIGNEFDRGISGGEKR 536
Query: 259 RVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQS-GSIVIMSIHQPSYRILG 317
RVSI +++ P++LFLDEPTSGLD+++A V++ L R++ +++SIHQP I
Sbjct: 537 RVSIACELVTSPLVLFLDEPTSGLDASNANNVIECLVRLSSDYNRTLVLSIHQPRSNIFY 596
Query: 318 LLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDLIRDLEGSPGGTK 375
L D+++ LS+G+ VYSG+ ++ F G+ PD+ N ++ +D+ E P G +
Sbjct: 597 LFDKLVLLSKGEMVYSGNAKKVSEFLRNEGYICPDNYNIADYLIDI--TFEAGPQGKR 652
Score = 76.3 bits (186), Expect = 3e-13
Identities = 63/218 (28%), Positives = 105/218 (47%), Gaps = 10/218 (4%)
Query: 443 FWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAF 502
F ++ L+ RSF + R P+L ++ L T+++N+ N G Q R+G F F
Sbjct: 773 FLQQLSILNSRSFKNMYRNPKLLLGNYLLTILLSLFLGTLYYNVSNDISGFQNRMGLFFF 832
Query: 503 AMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFL-SLAFAVI 561
++ + T L F ER IF++E + N Y +Y IS + + PL + + ++I
Sbjct: 833 ILTYFGFVTFTGLSSFALERIIFIKERSNNYYSPLAYYISKIMSEVVPLRVVPPILLSLI 892
Query: 562 TFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYF--LL 619
+ GL+ + F F + I F G S G++ L +I++++L LL
Sbjct: 893 VYPMTGLNMKDNAF-FKCIGILILFNLGISLEILTIGII-FEDLNNSIILSVLVLLGSLL 950
Query: 620 LSGFFIDRDRIPSYWIWFHYL---SLVKYPYEAVLQNE 654
SG FI+ I + + F YL S+ Y YE++L NE
Sbjct: 951 FSGLFINTKNITN--VAFKYLKNFSVFYYAYESLLINE 986
>CDR4_CANAL (O74676) ABC transporter CDR4
Length = 1490
Score = 199 bits (505), Expect = 3e-50
Identities = 154/562 (27%), Positives = 281/562 (49%), Gaps = 53/562 (9%)
Query: 122 SRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRL-KGTLALNGEALES 180
S + +L+ +SG + G++ A++GASG+GK+TL++AL++R+ G + +G +NG L+S
Sbjct: 859 SEDRVILDHVSGWVKPGQVTALMGASGAGKTTLLNALSDRLTTGVVTEGIRLVNGRPLDS 918
Query: 181 RLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAA 240
+ I YV Q DL TV E L FAA R P+++S+ +K V +I L + A
Sbjct: 919 SFQRSIG-YVQQQDLHLETSTVREALEFAAYLRQPKSVSRKEKNEYVDYIIRLLEMEQYA 977
Query: 241 KTVIGDEGHRGVSGGERRRVSIGIDIIHDP-ILLFLDEPTSGLDSTSAFMVVKVLQRIAQ 299
V+G G G++ +R+R+SIG++++ P +L+FLDEPTSGLDS +A+ + K+++++A
Sbjct: 978 DAVVGVSGE-GLNVEQRKRLSIGVELVAKPKLLVFLDEPTSGLDSQTAWSICKLIRKLAD 1036
Query: 300 SGSIVIMSIHQPSYRILGLLDRMIFLSR-GQTVYSG----SPTQLPSFFAEFGHP-LPDS 353
+G ++ +IHQPS +L DR++FL R GQTVY G + T L ++F ++G P P
Sbjct: 1037 NGQAILCTIHQPSAILLAEFDRLLFLQRGGQTVYFGDLGKNFTTLINYFEKYGAPKCPPE 1096
Query: 354 DNRTEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISA 413
N E+ L++I G+ G+K+ ++ W SS+ + N S + +S
Sbjct: 1097 ANPAEWMLEVI----GAAPGSKANQDYYDVWLK---------SSEFQEMN--SELDLMSE 1141
Query: 414 SISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVM 473
+ + L P + P +A P+W + + ++KR F + R P + V+
Sbjct: 1142 ELVKKPL--------DDDPDRLKP-YAAPYWEQYLFVTKRVFEQNWRTPSYLYSKFLLVV 1192
Query: 474 VTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADA-LPVFLQERYIF-MRETAY 531
+ F+ D S +G+Q ++ F F +T LP F+ +R ++ +RE
Sbjct: 1193 TSSLFNGFSFYKADRSLQGLQNQM-FSVFMFLVILHTLIQQYLPTFVSQRDLYEVRERPS 1251
Query: 532 NAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGAS---------GFLFYFLII 582
+ +++ + +P ++ VGL A+ F+++ +++
Sbjct: 1252 KTFSWITFIAAQVTAEIPWNIICGTLGYFCWYYPVGLYQNATYTNTVHQRGAFMWFAIVL 1311
Query: 583 FASFWA--GNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYL 640
F + + ++FL L + LA+ G + ++++P +W++ +
Sbjct: 1312 FFIYTSTLAQLCISFLEIDDNAANLSVLLFTMCLAF----CGVLVTKEQLPGFWVFMYRC 1367
Query: 641 SLVKYPYEAVLQ-NEFDDPVKC 661
S Y +L D PV C
Sbjct: 1368 SPFTYLVSVMLSVGLVDAPVTC 1389
Score = 146 bits (369), Expect = 2e-34
Identities = 124/550 (22%), Positives = 238/550 (42%), Gaps = 28/550 (5%)
Query: 124 TKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLL 183
T +L + G + GE+ VLG G+G ST + +A++ + + +L +
Sbjct: 169 TFDILKPMDGLIKPGELTVVLGRPGAGCSTFLKTIASQTYGYHIDKDSVIRYNSLTPHEI 228
Query: 184 KVIS----AYVMQDDLLFPMLTVEETLTFAAEFRLPRT----LSKSKKKARVQALIDQL- 234
K Y + + FP LTV +TL FAA+ R P+ +S+ + A++ +
Sbjct: 229 KKHYRGEVVYCAETENHFPQLTVGDTLEFAAKMRTPQNRPLGVSRDAYARHLAAVVMAVY 288
Query: 235 GLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVL 294
GL + T +G++ RGVSGGER+RVSI +++ ++ D T GLDS +A ++ L
Sbjct: 289 GLSHTRNTKVGNDFIRGVSGGERKRVSIAEITLNNAMVQCWDNSTRGLDSATALEFIRAL 348
Query: 295 QRIAQS-GSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDS 353
+ A + +++I+Q S L D+++ + +G +Y GS + +F + G+ P
Sbjct: 349 KASADIVHTTPLVAIYQCSQDAYDLFDKVVLMYQGYQIYFGSAKKAKQYFIDMGYECPQR 408
Query: 354 DNRTEFALDLIRDLE-----GSPGGT-KSLVEFNKSWQSMTKVHS--HSVSSQPERPNGM 405
+F L E G G ++ EF + W+ + V +
Sbjct: 409 QTTADFLTSLTNPAERIVRQGFEGKVPQTPQEFYEYWKKSPEGQQIVADVDQYLTEHSSA 468
Query: 406 SLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELF 465
+ KEAI K A + P+S + F++++ ++ R+ + P +
Sbjct: 469 AEKEAI-------KEAHQARQSDHLKPAS---PYTVSFFMQVRYIAHRNILRIKGNPSIH 518
Query: 466 GIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERYIF 525
++ + FIL+++F+NL + R FA+ ++ + + R I
Sbjct: 519 LFQIFGNIGMSFILSSIFYNLPTATSSFYHRTAALFFAVLFNAFSCLLEIFSLYEARSIV 578
Query: 526 MRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFAS 585
+ Y Y ++ + + LP +++ F ++ ++ V F FY LI F++
Sbjct: 579 EKHKKYALYHPAADAFASIVTELPTKFIIAIGFNLVYYFMVNFRRTPGNFFFYLLINFSA 638
Query: 586 FWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKY 645
A + + + T +L + +GF I + + W +YL + Y
Sbjct: 639 TLAMSHIFRTIGAATKTLQEAMTPAAILLLALTIFTGFVIPTPNMHGWCRWINYLDPLAY 698
Query: 646 PYEAVLQNEF 655
+E+++ NEF
Sbjct: 699 AFESLIANEF 708
>YN99_YEAST (P53756) Probable ATP-dependent transporter YNR070W
Length = 1333
Score = 191 bits (486), Expect = 5e-48
Identities = 158/566 (27%), Positives = 279/566 (48%), Gaps = 51/566 (9%)
Query: 122 SRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESR 181
S + LL+ +SG G + A++G SG+GK+TL++ LA R G + G + ++G +++
Sbjct: 742 SGQRKLLDSVSGYCVPGTLTALIGESGAGKTTLLNTLAQRNV-GTITGDMLVDGLPMDAS 800
Query: 182 LLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAK 241
K + YV Q DL LTV+E+L F+A R P+++ ++K V+ +I L ++ ++
Sbjct: 801 F-KRRTGYVQQQDLHVAELTVKESLQFSARMRRPQSIPDAEKMEYVEKIISILEMQEFSE 859
Query: 242 TVIGDEGHRGVSGGERRRVSIGIDIIHDP-ILLFLDEPTSGLDSTSAFMVVKVLQRIAQS 300
++G+ G+ G++ +R+++SIG++++ P +LLFLDEPTSGLDS SA+ VVK+L+R+A +
Sbjct: 860 ALVGEIGY-GLNVEQRKKLSIGVELVGKPDLLLFLDEPTSGLDSQSAWAVVKMLKRLALA 918
Query: 301 GSIVIMSIHQPSYRILGLLDRMIFLSR-GQTVYSG-----SPTQLPSFFAEFGHPLPDSD 354
G ++ +IHQPS + DR++ L + GQT+Y G S + + F ++
Sbjct: 919 GQSILCTIHQPSATLFEQFDRLLLLGKGGQTIYFGEIGKNSSSVIKYFEKNGARKCQQNE 978
Query: 355 NRTEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISAS 414
N E+ L+ I G + ++W + + SH ++ E+ N M
Sbjct: 979 NPAEYILEAI--------GAGATASVQQNWPDIWQ-KSHEYANINEKINDMI-------- 1021
Query: 415 ISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMV 474
K +S T T T +S +A + + + KRS R ++ +M+
Sbjct: 1022 ----KDLSSTTLHKTATRAS---KYATSYSYQFHHVLKRSSLTFWRNLNYIMAKMMLLMI 1074
Query: 475 TGFILATMFWNLDNSPKGVQERL--GFFAFAMSTTFYTTADALPVFLQERYIFMRETAYN 532
+G + F+++ + G+Q L F A +S +E Y +RE+ N
Sbjct: 1075 SGLFIGFTFFHVGVNAIGLQNSLFACFMAIVISAPATNQIQERATVAKELY-EVRESKSN 1133
Query: 533 AYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASG---FLFYFLIIFASFWAG 589
+ S LI+H L LP S F V +++ +G+ AS F + I+F ++ G
Sbjct: 1134 MFHWSLLLITHYLNELPYHLLFSTIFFVSSYFPLGVFTEASRSSVFYLNYAILFQLYYIG 1193
Query: 590 NSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEA 649
+ + + P++ IV IL++ L G +P +W + LS PY
Sbjct: 1194 LALMILY--MSPNLQSANVIVGFILSFLLSFCGAVQPASLMPGFWTFMWKLS----PYTY 1247
Query: 650 VLQN-----EFDDPVKCFVRGVQIFD 670
LQN D PV+C + + +F+
Sbjct: 1248 FLQNLVGLLMHDKPVRCSKKELSLFN 1273
Score = 135 bits (341), Expect = 3e-31
Identities = 126/555 (22%), Positives = 237/555 (42%), Gaps = 34/555 (6%)
Query: 122 SRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALA---NRIAKGRLKGTLALNGEAL 178
++ K +L ++S A+ GE++ VLG G+G ++ + + A ++ A G G ++ +G
Sbjct: 40 NKMKIILKNVSLLAKSGEMVLVLGRPGAGCTSFLKSAAGETSQFAGGVTTGHISYDGIPQ 99
Query: 179 ESRL--LKVISAYVMQDDLLFPMLTVEETLTFAAEFRLP--RTLSKSKKK---ARVQALI 231
+ + K Y + D+ FP LTV++TL FA ++P R + +K++ A +
Sbjct: 100 KEMMQHYKPDVIYNGEQDVHFPHLTVKQTLDFAISCKMPAKRVNNVTKEEYITANREFYA 159
Query: 232 DQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVV 291
GL + T +G++ GVSGGER+RVSI + + D T GLDS++A
Sbjct: 160 KIFGLTHTFDTKVGNDFISGVSGGERKRVSIAEALAAKGSIYCWDNATRGLDSSTALEFA 219
Query: 292 KVLQRIAQ-SGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPL 350
+ ++ + G+ +++++Q S I D++ L G+ ++ G T+ +F G+
Sbjct: 220 RAIRTMTNLLGTTALVTVYQASENIYETFDKVTVLYAGRQIFCGKTTEAKDYFENMGYLC 279
Query: 351 PDSDNRTEFALDLIRDLEG----SPGGTKSLV----EFNKSWQSMTKVH--SHSVSSQPE 400
P + E+ L I D G PG + EF K W + +
Sbjct: 280 PPRQSTAEY-LTAITDPNGLHEIKPGFEYQVPHTADEFEKYWLDSPEYARLKGEIQKYKH 338
Query: 401 RPNGMSLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRR 460
N K+ + S+++ K G + T S +W ++ + R F
Sbjct: 339 EVNTEWTKKTYNESMAQEK-SKGTRKKSYYTVS---------YWEQIRLCTIRGFLRIYG 388
Query: 461 KPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQ 520
I A + FI ++F+ +S G R G F S +Y+ + +
Sbjct: 389 DKSYTVINTCAAIAQAFITGSLFYQAPSSTLGAFSRSGVLFF--SLLYYSLMGLANISFE 446
Query: 521 ERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFL 580
R I + Y+ Y S+ ++ + S P F +I ++ GL A F +L
Sbjct: 447 HRPILQKHKVYSLYHPSAEALASTISSFPFRMIGLTFFIIILYFLAGLHRSAGAFFTMYL 506
Query: 581 IIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYL 640
++ A S +S + + +I ++ + S + I + ++ W Y+
Sbjct: 507 LLTMCSEAITSLFQMVSSLCDTLSQANSIAGVVMLSIAMYSTYMIQLPSMHPWFKWISYI 566
Query: 641 SLVKYPYEAVLQNEF 655
++Y +E++L EF
Sbjct: 567 LPIRYAFESMLNAEF 581
>CDR3_CANAL (O42690) Opaque-specific ABC transporter CDR3
Length = 1501
Score = 186 bits (473), Expect = 1e-46
Identities = 153/610 (25%), Positives = 282/610 (46%), Gaps = 53/610 (8%)
Query: 69 GMEPRSLPFVLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRTKTLL 128
GM P S+ +Y ++ L S+IF RN V+ + S + +L
Sbjct: 809 GMAPLDFSGSTEISDYSYDYMDRKLLDTSNIF-HWRNLTYTVK--------IKSEERVIL 859
Query: 129 NDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRL-KGTLALNGEALESRLLKVIS 187
N+I G + GE+ A++GASG+GK+TL++AL+ R+ G + GT +NG L+S + I
Sbjct: 860 NNIDGWVKPGEVTALMGASGAGKTTLLNALSERLTTGVITSGTRMVNGGELDSSFQRSIG 919
Query: 188 AYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDE 247
YV Q DL TV E L F+A R P ++S ++K + V+ +ID L +R ++G
Sbjct: 920 -YVQQQDLHLETSTVREALKFSARLRQPNSVSIAEKDSYVEKIIDLLEMRTYVDAIVGVP 978
Query: 248 GHRGVSGGERRRVSIGIDIIHDP-ILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIM 306
G G++ +R+R++I ++++ P +L+FLDEPTSGLDS +A+ + K+++++A G ++
Sbjct: 979 GE-GLNVEQRKRLTIAVELVARPKLLVFLDEPTSGLDSQTAWSICKLIRKLANHGQAILC 1037
Query: 307 SIHQPSYRILGLLDRMIFLSRGQTVYSGS-----PTQLPSFFAEFGHPLPDSDNRTEFAL 361
+IHQPS +L DR++ L +G+TVY G T + F P N E+ L
Sbjct: 1038 TIHQPSAILLEEFDRLLLLQKGETVYFGEFGANCHTLIEYFERNGASKCPQHANPAEWML 1097
Query: 362 DLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASISRGKLV 421
+I G+ GT++ ++ ++W++ P +++ + L
Sbjct: 1098 GVI----GAAPGTQANQDYFETWRN--------------SPEYRAVQNELHRLEEMPGLA 1139
Query: 422 SGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILAT 481
SG T +A FW + + + R F R P + ++
Sbjct: 1140 SGEKEPDTN------QAYAASFWKQYIFVVHRLFQQYWRTPSYIYSKFAMAVLCSLFNGF 1193
Query: 482 MFWNLDNSPKGVQERLGFFAFAMSTTFYTTADA-LPVFLQERYIF-MRETAYNAYRRSSY 539
++ NS +G++ ++ F+M T A +P+F+ +R ++ RE + ++
Sbjct: 1194 TYYKSQNSMQGLKNQM-LSIFSMFVVLTTLAQQYVPLFVTQRDLYEARERPSKTFSWLAF 1252
Query: 540 LISHALVSLPPLAFLSLAFAVITFWAVGLDGGA--SGFLFY-----FLIIFASFWAGNSF 592
+ + +P + ++ VGL A SG + + +LI+ F ++
Sbjct: 1253 IAAQITAEIPYQVLAATISFFSWYYPVGLYRNAVYSGAVTHRGVLMWLIMTLMFIYSSTL 1312
Query: 593 VTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQ 652
F + +L ++ G +D +P +W++ + + + Y A++
Sbjct: 1313 AQFCISWNQLADYAANWISLLLTISMIFCGVIATKDSMPKFWVFLYRCTPLTYLTSAMMS 1372
Query: 653 NEFDDP-VKC 661
D VKC
Sbjct: 1373 IGLGDSFVKC 1382
Score = 122 bits (305), Expect = 5e-27
Identities = 127/555 (22%), Positives = 232/555 (40%), Gaps = 44/555 (7%)
Query: 127 LLNDISGEARDGEIMAVLGASGSGKSTLIDALANR-----IAKGRLKGTLALNGEALESR 181
+L + G + GE+ VLG G+G ST + +A R +A G + + + + +
Sbjct: 160 ILKPMEGLIKPGEVTVVLGRPGAGCSTFLKTIACRTEGFHVADGSVISYDGITQDEIRNH 219
Query: 182 LLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLP--RTLSKSKK---KARVQALIDQLGL 236
L + Y + + FP LTV ETL FAA + P R + S++ K V ++ GL
Sbjct: 220 LRGEV-VYCAETETHFPNLTVGETLEFAALMKTPQNRPMGVSREEYAKHVVDVVMATYGL 278
Query: 237 RNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQR 296
+ T +G++ RG+SGGER+R+SI + + D T GLD+ +A + L+
Sbjct: 279 SHTKNTKVGNDFIRGISGGERKRLSIAEVTLVQASIQCWDNSTRGLDAATALEFISSLKT 338
Query: 297 IAQ-SGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDN 355
A +++I+Q S L D++I + G ++ GS + ++F + G D
Sbjct: 339 SASILNDTPLIAIYQCSQNAYDLFDKVIVMYEGYQIFFGSSQRAAAYFKKMGFVCQDRQT 398
Query: 356 RTEFALDLIRDLEG--SPGGTKSL----VEFNKSWQSMTKVHSHSVSSQPERPNGMSLKE 409
+F + E PG + + EF + W+ PER +L E
Sbjct: 399 TPDFLTSITSPAERIIKPGYERLVPRTPKEFYRYWR-----------RSPER---QALLE 444
Query: 410 AIS------ASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPE 463
I + + + + A N + ++ +++ + KR + R
Sbjct: 445 EIDEYLDNCENYDQKQKIFEANNAKKAKHTYNKSSYTVSLPMQVRYIMKRYWDRMRGDII 504
Query: 464 LFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERY 523
+ + + IL+++F+NL + R +A+ Y++ + + R
Sbjct: 505 VPLSTVAGNIAMALILSSVFYNLQPNSSSFYYRTSVMYYALLFNAYSSVLEIYNMYEGRA 564
Query: 524 IFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIF 583
I + Y Y + I + P S+ F +I ++ V F FY LI F
Sbjct: 565 IVQKHREYALYPPMADAIGSIISDFPLKVVCSVLFNLILYFMVNFKREPGAFFFYLLISF 624
Query: 584 AS-FWAGNSFVTF--LSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYL 640
S + + F T + + M ++++ L+ F SGF I + + W ++
Sbjct: 625 CSTLFMSHLFRTIGAFTNSLAEAMTPSSLLLFALSTF---SGFAIPVTYMLGWCKWIRWV 681
Query: 641 SLVKYPYEAVLQNEF 655
+ + Y YEA++ NEF
Sbjct: 682 NPLAYAYEALISNEF 696
>SNQ2_YEAST (P32568) SNQ2 protein
Length = 1501
Score = 180 bits (456), Expect = 1e-44
Identities = 169/635 (26%), Positives = 301/635 (46%), Gaps = 59/635 (9%)
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLK 184
+ LL+++SG G + A++G SG+GK+TL++ LA R G + G + +NG +++ +
Sbjct: 869 RMLLDNVSGYCIPGTMTALMGESGAGKTTLLNTLAQRNV-GIITGDMLVNGRPIDASFER 927
Query: 185 VISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVI 244
+ YV Q D+ LTV E+L F+A R P+ L S+K V+ +I LG+ A+ ++
Sbjct: 928 R-TGYVQQQDIHIAELTVRESLQFSARMRRPQHLPDSEKMDYVEKIIRVLGMEEYAEALV 986
Query: 245 GDEGHRGVSGGERRRVSIGIDIIHDP-ILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSI 303
G+ G G++ +R+++SIG++++ P +LLFLDEPTSGLDS S++ ++++L++++++G
Sbjct: 987 GEVGC-GLNVEQRKKLSIGVELVAKPDLLLFLDEPTSGLDSQSSWAIIQLLRKLSKAGQS 1045
Query: 304 VIMSIHQPSYRILGLLDRMIFLSR-GQTVYSG-----SPTQLPSFFAEFGHPLPDSDNRT 357
++ +IHQPS + DR++ L + GQTVY G S T L F S+N
Sbjct: 1046 ILCTIHQPSATLFEEFDRLLLLRKGGQTVYFGDIGKNSATILNYFERNGARKCDSSENPA 1105
Query: 358 EFALDLIRDLEGSPGGTKSLVE-FNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASIS 416
E+ L+ I G T S+ E +++ W + + Q + N +S +E S
Sbjct: 1106 EYILEAI-----GAGATASVKEDWHEKWLNSVEFEQTKEKVQ-DLINDLSKQETKS---- 1155
Query: 417 RGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTG 476
PS ++A F ++ R+ T R ++ ++V G
Sbjct: 1156 ----------EVGDKPSKYATSYAYQFRYVLI----RTSTSFWRSLNYIMSKMMLMLVGG 1201
Query: 477 FILATMFWNLDNSPKGVQERL--GFFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAY 534
+ F+N+ S G+Q + F + +S + +E + +RE+ N +
Sbjct: 1202 LYIGFTFFNVGKSYVGLQNAMFAAFISIILSAPAMNQIQGRAIASRELF-EVRESQSNMF 1260
Query: 535 RRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFL---IIFASFWAG-N 590
S LI+ L LP F S F V +++ + + AS YFL I+F ++ G
Sbjct: 1261 HWSLVLITQYLSELPYHLFFSTIFFVSSYFPLRIFFEASRSAVYFLNYCIMFQLYYVGLG 1320
Query: 591 SFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAV 650
+ ++S P++ I+ L++ L G +P +W + S PY
Sbjct: 1321 LMILYMS---PNLPSANVILGLCLSFMLSFCGVTQPVSLMPGFWTFMWKAS----PYTYF 1373
Query: 651 LQN-----EFDDPVKCFVRGVQIFDNSPLRSVPYELKLKLLGSMSGTLGTNITATTCLTT 705
+QN PV C + + F N P S E L +G + + C
Sbjct: 1374 VQNLVGIMLHKKPVVCKKKELNYF-NPPNGSTCGEYMKPFLEKATGYIENPDATSDCAYC 1432
Query: 706 GADILQQNGVTEL-SKWNCLWITVAWGFFFRFLFY 739
++ N +T + SK++ LW +G F+ ++F+
Sbjct: 1433 IYEV-GDNYLTHISSKYSYLWRN--FGIFWIYIFF 1464
Score = 150 bits (379), Expect = 1e-35
Identities = 136/553 (24%), Positives = 241/553 (42%), Gaps = 33/553 (5%)
Query: 123 RTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAK--GRLKGTLALNGEALES 180
+ + ++++++ A GE++ VLG G+G S+ + A I + G + G +A +G E
Sbjct: 171 KMRQIISNVNALAEAGEMILVLGRPGAGCSSFLKVTAGEIDQFAGGVSGEVAYDGIPQEE 230
Query: 181 RL--LKVISAYVMQDDLLFPMLTVEETLTFAAEFRLP--RTLSKSKKK---ARVQALIDQ 233
+ K Y + D+ FP LTV++TL FA + P R + SKK+ +R
Sbjct: 231 MMKRYKADVIYNGELDVHFPYLTVKQTLDFAIACKTPALRVNNVSKKEYIASRRDLYATI 290
Query: 234 LGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKV 293
GLR+ T +G++ RGVSGGER+RVSI + + D T GLD+++A K
Sbjct: 291 FGLRHTYNTKVGNDFVRGVSGGERKRVSIAEALAAKGSIYCWDNATRGLDASTALEYAKA 350
Query: 294 LQRIAQS-GSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPD 352
++ + S ++I+Q S I D++ L G+ +Y G + +FA+ G+ P
Sbjct: 351 IRIMTNLLKSTAFVTIYQASENIYETFDKVTVLYSGKQIYFGLIHEAKPYFAKMGYLCPP 410
Query: 353 SDNRTEFALDLIRDLEG----SPGGT----KSLVEFNKSWQSMTKVHS--HSVSSQPERP 402
EF L + D G PG ++ EF W + + +++ E+
Sbjct: 411 RQATAEF-LTALTDPNGFHLIKPGYENKVPRTAEEFETYWLNSPEFAQMKKDIAAYKEKV 469
Query: 403 NGMSLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKP 462
N KE S+++ K + T S + +W ++ ++R F
Sbjct: 470 NTEKTKEVYDESMAQEK-------SKYTRKKSY---YTVSYWEQVKLCTQRGFQRIYGNK 519
Query: 463 ELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQER 522
I + + ++ FI ++F+N +S G R G FA+ +Y+ + + R
Sbjct: 520 SYTVINVCSAIIQSFITGSLFYNTPSSTSGAFSRGGVLYFAL--LYYSLMGLANISFEHR 577
Query: 523 YIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLII 582
I + Y+ Y S+ I L S P F +I F+ GL A F +L +
Sbjct: 578 PILQKHKGYSLYHPSAEAIGSTLASFPFRMIGLTCFFIILFFLSGLHRTAGSFFTIYLFL 637
Query: 583 FASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSL 642
A N +S V + +I ++ + S + I + ++ W Y+
Sbjct: 638 TMCSEAINGLFEMVSSVCDTLSQANSISGILMMSISMYSTYMIQLPSMHPWFKWISYVLP 697
Query: 643 VKYPYEAVLQNEF 655
++Y +E++L EF
Sbjct: 698 IRYAFESMLNAEF 710
>PDRC_YEAST (Q02785) ATP-dependent permease PDR12
Length = 1511
Score = 177 bits (448), Expect = 1e-43
Identities = 142/525 (27%), Positives = 244/525 (46%), Gaps = 45/525 (8%)
Query: 124 TKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLL 183
T+ LL+D+ G + G++ A++G SG+GK+TL++ LA RI G + G + +N + L +
Sbjct: 857 TRKLLSDVFGYVKPGKMTALMGESGAGKTTLLNVLAQRINMGVITGDMLVNAKPLPASFN 916
Query: 184 KVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTV 243
+ YV Q D L+V E+L FAAE R ++ +K V+ +I LG++N A+ +
Sbjct: 917 RSCG-YVAQADNHMAELSVRESLRFAAELRQQSSVPLEEKYEYVEKIITLLGMQNYAEAL 975
Query: 244 IGDEGHRGVSGGERRRVSIGIDIIHDP-ILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGS 302
+G G RG++ +R+++SIG++++ P +LLFLDEPTSGLDS SA+ +V+ ++ +A SG
Sbjct: 976 VGKTG-RGLNVEQRKKLSIGVELVAKPSLLLFLDEPTSGLDSQSAWSIVQFMRALADSGQ 1034
Query: 303 IVIMSIHQPSYRILGLLDRMIFLSR-GQTVYSG-----SPTQLPSFFAEFGHPLPDSDNR 356
++ +IHQPS + DR++ L + G+ VY G S T L F + G S+N
Sbjct: 1035 SILCTIHQPSATLFEQFDRLLLLKKGGKMVYFGDIGPNSETLLKYFERQSGMKCGVSENP 1094
Query: 357 TEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASIS 416
E+ L+ I G + N W + ++ E +A
Sbjct: 1095 AEYILNCI--------GAGATASVNSDWHDL----------------WLASPECAAARAE 1130
Query: 417 RGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTG 476
+L A + FA + ++ + +R+ R P + +
Sbjct: 1131 VEELHRTLPGRAVNDDPELATRFAASYMTQIKCVLRRTALQFWRSPVYIRAKFFECVACA 1190
Query: 477 FILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQE-RYIF-MRETAYNAY 534
+ + +++S G E F + + L VF + R ++ +RE A N +
Sbjct: 1191 LFVGLSYVGVNHSVGGAIEAFSSI-FMLLLIALAMINQLHVFAYDSRELYEVREAASNTF 1249
Query: 535 RRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGAS--GFLFYFLIIFASFWAGNSF 592
S L+ HA V + +W G AS GF F+F ++ + F
Sbjct: 1250 HWSVLLLCHAAVENFWSTLCQFMCFICYYWPAQFSGRASHAGFFFFFYVLIFPLY----F 1305
Query: 593 VTF---LSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYW 634
VT+ + + P V I + A LL G R+++P++W
Sbjct: 1306 VTYGLWILYMSPDVPSASMINSNLFAAMLLFCGILQPREKMPAFW 1350
Score = 134 bits (336), Expect = 1e-30
Identities = 135/618 (21%), Positives = 255/618 (40%), Gaps = 67/618 (10%)
Query: 79 LSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDAPAVEESVFSRT-----KTLLNDISG 133
++F NLT +V V ++ P + PA S F++ + ++ + +G
Sbjct: 114 IAFKNLT-AVGVDASAAYG---PSVEEMFRNIASIPAHLISKFTKKSDVPLRNIIQNCTG 169
Query: 134 EARDGEIMAVLGASGSGKSTLIDALANRIAK-GRLKGTLALNGEALESRLLKVISAYVM- 191
GE++ V+G G+G ST + L+ ++ ++G + +G +S ++ YV+
Sbjct: 170 VVESGEMLFVVGRPGAGCSTFLKCLSGETSELVDVQGEFSYDGLD-QSEMMSKYKGYVIY 228
Query: 192 --QDDLLFPMLTVEETLTFAAEFRLPRT-LSKSKKKARVQALIDQ----LGLRNAAKTVI 244
+ D FP +TV+ET+ FA + + PR + K +K V + D GLR+ T +
Sbjct: 229 CPELDFHFPKITVKETIDFALKCKTPRVRIDKMTRKQYVDNIRDMWCTVFGLRHTYATKV 288
Query: 245 GDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQS-GSI 303
G++ RGVSGGER+RVS+ + + D T GLD+++A + ++ +
Sbjct: 289 GNDFVRGVSGGERKRVSLVEAQAMNASIYSWDNATRGLDASTALEFAQAIRTATNMVNNS 348
Query: 304 VIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDL 363
I++I+Q I L D+ L G+ +Y G + +F G P+ EF +
Sbjct: 349 AIVAIYQAGENIYELFDKTTVLYNGRQIYFGPADKAVGYFQRMGWVKPNRMTSAEFLTSV 408
Query: 364 IRDLEG-----SPG----GTKSLVEFNKSW------QSMTKVHSHSVSSQPERPNGMSLK 408
D E PG KS EF + W Q + + + S P L
Sbjct: 409 TVDFENRTLDIKPGYEDKVPKSSSEFEEYWLNSEDYQELLRTYDDYQSRHPVNETRDRLD 468
Query: 409 EAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIR 468
A + +G+ + + +W ++ R F + +
Sbjct: 469 VAKKQRLQQGQRENS--------------QYVVNYWTQVYYCMIRGFQRVKGDSTYTKVY 514
Query: 469 LGAVMVTGFILATMFWNLDNSPK----GVQERLGFFAFAMSTTFYTTADALPVFLQERYI 524
L + ++ I+ +MF +D+ + G R G + + T+ + R +
Sbjct: 515 LSSFLIKALIIGSMFHKIDDKSQSTTAGAYSRGGMLFYVLLFASVTSLAEIGNSFSSRPV 574
Query: 525 FMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFA 584
++ +Y+ Y S+ + + P + +IT+W + A F Y L +
Sbjct: 575 IVKHKSYSMYHLSAESLQEIITEFPTKFVAIVILCLITYWIPFMKYEAGAFFQYILYLLT 634
Query: 585 SFWAGNSFVTFL-----SGVVPHVMLG-YTIVVAILAYFLLLSGFFIDRDRIPSYWI-WF 637
+ F+ SGV H + G + +++ + A F+L G +WI W
Sbjct: 635 VQQCTSFIFKFVATMSKSGVDAHAVGGLWVLMLCVYAGFVLPIGEM-------HHWIRWL 687
Query: 638 HYLSLVKYPYEAVLQNEF 655
H+++ + Y +E+++ EF
Sbjct: 688 HFINPLTYAFESLVSTEF 705
>CDR2_CANAL (P78595) Multidrug resistance protein CDR2
Length = 1499
Score = 176 bits (447), Expect = 2e-43
Identities = 142/547 (25%), Positives = 267/547 (47%), Gaps = 50/547 (9%)
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLK-GTLALNGEALESRLL 183
+ +L+ + G + G+I A++GASG+GK+TL++ L+ R+ G + G +NG AL+S
Sbjct: 873 RVILDHVDGWVKPGQITALMGASGAGKTTLLNCLSERVTTGIITDGERLVNGHALDSSFQ 932
Query: 184 KVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTV 243
+ I YV Q D+ TV E L F+A R +SK +K V +ID L + + A +
Sbjct: 933 RSIG-YVQQQDVHLETTTVREALQFSAYLRQSNKISKKEKDDYVDYVIDLLEMTDYADAL 991
Query: 244 IGDEGHRGVSGGERRRVSIGIDIIHDP-ILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGS 302
+G G G++ +R+R++IG++++ P +LLFLDEPTSGLDS +A+ + K+++++A G
Sbjct: 992 VGVAGE-GLNVEQRKRLTIGVELVAKPKLLLFLDEPTSGLDSQTAWSICKLMRKLADHGQ 1050
Query: 303 IVIMSIHQPSYRILGLLDRMIFLSR-GQTVYSGSPTQ----LPSFFAEFG-HPLPDSDNR 356
++ +IHQPS I+ D+++FL + G+T Y G + + ++F ++G P P N
Sbjct: 1051 AILCTIHQPSALIMAEFDKLLFLQKGGRTAYFGELGENCQTMINYFEKYGADPCPKEANP 1110
Query: 357 TEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASIS 416
E+ L ++ G+ G+ + ++ + W+ +S + E N M EA + +
Sbjct: 1111 AEWMLQVV----GAAPGSHAKQDYFEVWR-----NSSEYQAVREEINRM---EAELSKLP 1158
Query: 417 RGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTG 476
R P +++ +A P W + + +S R+ R P +L V+ +
Sbjct: 1159 R-----------DNDPEALL-KYAAPLWKQYLLVSWRTIVQDWRSPGYIYSKLILVISSS 1206
Query: 477 FILATMFWNLDNSPKGVQERL--GFFAFAMSTTFYTTADALPVFLQERYIF-MRETAYNA 533
+ F+ N+ +G+Q ++ F F TTF LP F++ R ++ +RE
Sbjct: 1207 LFIGFSFFKSKNNLQGLQSQMLAVFMFFVPFTTFID--QMLPYFVKHRAVYEVREAPSRT 1264
Query: 534 YRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGAS--------GFLFYFLIIFAS 585
+ +++ +P + ++ VGL A G L + L+ +
Sbjct: 1265 FSWFAFIAGQITSEIPFQIVVGTISYFCWYYPVGLYANAEPTDSVNSRGVLMWMLL--TA 1322
Query: 586 FWAGNSFVTFLSGVVPHVM-LGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVK 644
F+ S + L+ + ++ + + L+ G + IP +WI+ + +
Sbjct: 1323 FYVYTSTMGQLAISLNELIDNAANLATTLFTLCLMFCGVLAGPNVIPGFWIFMYRCNPFT 1382
Query: 645 YPYEAVL 651
Y +A+L
Sbjct: 1383 YLIQAIL 1389
Score = 151 bits (381), Expect = 7e-36
Identities = 171/767 (22%), Positives = 312/767 (40%), Gaps = 71/767 (9%)
Query: 13 DATSYLDLMELS-DLTRRPASGDLPT---LGQLLKHVGDARKEAAGDGSETPLHHALDVV 68
DAT+ ++ +L+ LT +GD + L + L H+ D + +G+ + H LD
Sbjct: 33 DATASHNIQDLARKLTHGSTNGDHHSANDLARYLSHMSDIPGVSPFNGNIS--HEQLDPD 90
Query: 69 GMEPRSLPFVLSFSNLTYSVKVKRKLSFSSIFPRRRNRLGAVEDA---PAVEESVFSRTK 125
+ +V + L S K S + R G D+ P V +++ T
Sbjct: 91 SENFNAKYWVKNLKKLFESDSDYYKPSKLGVAYRNLRAYGIANDSDYQPTVTNALWKFTT 150
Query: 126 TLLNDISGE---------------ARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGT 170
+N + R GE+ VLG G+G STL+ +A +
Sbjct: 151 EAINKLKKPDDSKYFDILKSMDAIMRPGELTVVLGRPGAGCSTLLKTIAVNTYGFHIGKE 210
Query: 171 LALNGEALE----SRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSK-----S 221
+ + L R + Y + D+ FP L+V +TL FAA R P+ + +
Sbjct: 211 SQITYDGLSPHDIERHYRGDVIYSAETDVHFPHLSVGDTLEFAARLRTPQNRGEGIDRET 270
Query: 222 KKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSG 281
K + GL + T +G++ RGVSGGER+RVSI + + D T G
Sbjct: 271 YAKHMASVYMATYGLSHTRNTNVGNDFVRGVSGGERKRVSIAEASLSGANIQCWDNATRG 330
Query: 282 LDSTSAFMVVKVLQRIAQS-GSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLP 340
LDS +A ++ L+ A + +++I+Q S L D ++ L G ++ G ++
Sbjct: 331 LDSATALEFIRALKTSATILDTTPLIAIYQCSQDAYELFDNVVVLYEGYQIFFGKASKAK 390
Query: 341 SFFAEFGHPLPDSDNRTEFALDLIRDLEGSP------GGTKSLVEFNKSWQSMTKVHSHS 394
+F G P +F L E P ++ EF W++ + ++
Sbjct: 391 EYFENMGWKCPQRQTTADFLTSLTNPAEREPLPGYEDKVPRTAQEFETFWKNSPE---YA 447
Query: 395 VSSQPERPNGMSLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRS 454
++ + + + + R V +N T PSS + F++++ + R+
Sbjct: 448 ELTKEIDEYFVECERSNTGETYRESHVGKQSNN--TRPSS---PYTVSFFMQVRYVIARN 502
Query: 455 FTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADA 514
F + P + I + + +V G ILA++F+NL S R G F++ +++
Sbjct: 503 FLRMKGDPSIPLISILSQLVMGLILASVFFNLRKSTDTFYFRGGALFFSVLFNAFSSLLE 562
Query: 515 LPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASG 574
+ + R I + Y YR S+ ++ + LP ++++F ++ ++ V L A
Sbjct: 563 ILSLYEARPIVEKHRKYALYRPSADALASIISELPVKLLMTMSFNIVYYFMVNLRRTAGN 622
Query: 575 FLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYW 634
F FY+L+ + + + V + ++ L ++ +GF + I +
Sbjct: 623 FFFYWLMCASCTLVMSHMFRSIGAVTTTIATAMSLSTVFLLAMIIYAGFVLPIPYILGWS 682
Query: 635 IWFHYLSLVKYPYEAVLQNEFDDPVKCFVRGVQIFDNSPLRSVPYELKLKLLGSMSGTLG 694
W Y++ V Y +E+++ NEF + F G I ++P E K
Sbjct: 683 RWIRYINPVTYIFESLMVNEFHG--REFECGQYIPSGPGFENLPVENK------------ 728
Query: 695 TNITATTCLTTGADILQQNGVTELS-------KWNCLWITVAWGFFF 734
+ T T G+ ++Q +L+ KW ITVA+ FF
Sbjct: 729 --VCTTVGSTPGSTVVQGTEYIKLAYQFYSSHKWRNFGITVAFAVFF 773
>CDR1_CANAL (P43071) Multidrug resistance protein CDR1
Length = 1501
Score = 175 bits (444), Expect = 3e-43
Identities = 148/562 (26%), Positives = 272/562 (48%), Gaps = 53/562 (9%)
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLK-GTLALNGEALESRLL 183
+ +L+ + G + G+I A++GASG+GK+TL++ L+ R+ G + G +NG AL+S
Sbjct: 875 RVILDHVDGWVKPGQITALMGASGAGKTTLLNCLSERVTTGIITDGERLVNGHALDSSFQ 934
Query: 184 KVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTV 243
+ I YV Q D+ P TV E L F+A R +SK +K V +ID L + + A +
Sbjct: 935 RSIG-YVQQQDVHLPTSTVREALQFSAYLRQSNKISKKEKDDYVDYVIDLLEMTDYADAL 993
Query: 244 IGDEGHRGVSGGERRRVSIGIDIIHDP-ILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGS 302
+G G G++ +R+R++IG++++ P +LLFLDEPTSGLDS +A+ + K+++++A G
Sbjct: 994 VGVAGE-GLNVEQRKRLTIGVELVAKPKLLLFLDEPTSGLDSQTAWSICKLMRKLADHGQ 1052
Query: 303 IVIMSIHQPSYRILGLLDRMIFLSR-GQTVYSGSPTQ----LPSFFAEFG-HPLPDSDNR 356
++ +IHQPS I+ DR++FL + G+T Y G + + ++F ++G P P N
Sbjct: 1053 AILCTIHQPSALIMAEFDRLLFLQKGGRTAYFGELGENCQTMINYFEKYGADPCPKEANP 1112
Query: 357 TEFALDLIRDLEGSPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAISASIS 416
E+ L ++ G+ G+ + ++ + W+ +S + E N M EA + +
Sbjct: 1113 AEWMLQVV----GAAPGSHAKQDYFEVWR-----NSSEYQAVREEINRM---EAELSKLP 1160
Query: 417 RGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTG 476
R P +++ +A P W + + +S R+ R P ++ V+
Sbjct: 1161 R-----------DNDPEALL-KYAAPLWKQYLLVSWRTIVQDWRSPGYIYSKIFLVVSAA 1208
Query: 477 FILATMFWNLDNSPKGVQERLGFFAFAMSTTFYT-TADALPVFLQERYIF-MRETAYNAY 534
F+ N+ +G+Q ++ F F F T LP F+++R ++ +RE +
Sbjct: 1209 LFNGFSFFKAKNNMQGLQNQM-FSVFMFFIPFNTLVQQMLPYFVKQRDVYEVREAPSRTF 1267
Query: 535 RRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGAS--------GFLFYFLI-IFAS 585
+++ +P + ++ +GL A+ G L + L+ F
Sbjct: 1268 SWFAFIAGQITSEIPYQVAVGTIAFFCWYYPLGLYNNATPTDSVNPRGVLMWMLVTAFYV 1327
Query: 586 FWA--GNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLV 643
+ A G ++F L + L + +L+G D +P +WI+ + +
Sbjct: 1328 YTATMGQLCMSFSELADNAANLATLLFTMCLNFCGVLAG----PDVLPGFWIFMYRCNPF 1383
Query: 644 KYPYEAVLQNEFDDP-VKCFVR 664
Y +A+L + VKC R
Sbjct: 1384 TYLVQAMLSTGLANTFVKCAER 1405
Score = 138 bits (348), Expect = 5e-32
Identities = 124/550 (22%), Positives = 234/550 (42%), Gaps = 24/550 (4%)
Query: 122 SRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALE-- 179
SR +L + R GE+ VLG G+G STL+ +A + + + L
Sbjct: 164 SRYFDILKSMDAIMRPGELTVVLGRPGAGCSTLLKTIAVNTYGFHIGKESQITYDGLSPH 223
Query: 180 --SRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSK-----SKKKARVQALID 232
R + Y + D+ FP L+V +TL FAA R P+ + + K +
Sbjct: 224 DIERHYRGDVIYSAETDVHFPHLSVGDTLEFAARLRTPQNRGEGIDRETYAKHMASVYMA 283
Query: 233 QLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVK 292
GL + T +G++ RGVSGGER+RVSI + + D T GLDS +A ++
Sbjct: 284 TYGLSHTRNTNVGNDFVRGVSGGERKRVSIAEASLSGANIQCWDNATRGLDSATALEFIR 343
Query: 293 VLQRIAQS-GSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLP 351
L+ A + +++I+Q S L D+++ L G ++ G T+ +F + G P
Sbjct: 344 ALKTSAVILDTTPLIAIYQCSQDAYDLFDKVVVLYEGYQIFFGKATKAKEYFEKMGWKCP 403
Query: 352 DSDNRTEFALDLIRDLEGSP------GGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGM 405
+F L E P ++ EF W++ + ++ ++ +
Sbjct: 404 QRQTTADFLTSLTNPAEREPLPGYEDKVPRTAQEFETYWKNSPE---YAELTKEIDEYFV 460
Query: 406 SLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELF 465
+ + + R V+ +N T P+S + F++++ R+F + P +
Sbjct: 461 ECERSNTRETYRESHVAKQSNN--TRPAS---PYTVSFFMQVRYGVARNFLRMKGDPSIP 515
Query: 466 GIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAFAMSTTFYTTADALPVFLQERYIF 525
+ +V G IL+++F+NL + R FA+ +++ + + R I
Sbjct: 516 IFSVFGQLVMGLILSSVFYNLSQTTGSFYYRGAAMFFAVLFNAFSSLLEIMSLFEARPIV 575
Query: 526 MRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFAS 585
+ Y YR S+ ++ + LP +S++F + ++ V F FY+L+
Sbjct: 576 EKHKKYALYRPSADALASIISELPVKLAMSMSFNFVFYFMVNFRRNPGRFFFYWLMCIWC 635
Query: 586 FWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKY 645
+ + + V + T +L ++ +GF I + + W +Y++ V Y
Sbjct: 636 TFVMSHLFRSIGAVSTSISGAMTPATVLLLAMVIYTGFVIPTPSMLGWSRWINYINPVGY 695
Query: 646 PYEAVLQNEF 655
+E+++ NEF
Sbjct: 696 VFESLMVNEF 705
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.137 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 84,004,397
Number of Sequences: 164201
Number of extensions: 3594464
Number of successful extensions: 13187
Number of sequences better than 10.0: 983
Number of HSP's better than 10.0 without gapping: 838
Number of HSP's successfully gapped in prelim test: 145
Number of HSP's that attempted gapping in prelim test: 10798
Number of HSP's gapped (non-prelim): 1454
length of query: 751
length of database: 59,974,054
effective HSP length: 118
effective length of query: 633
effective length of database: 40,598,336
effective search space: 25698746688
effective search space used: 25698746688
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0260.10


