

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0252a.5
(150 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
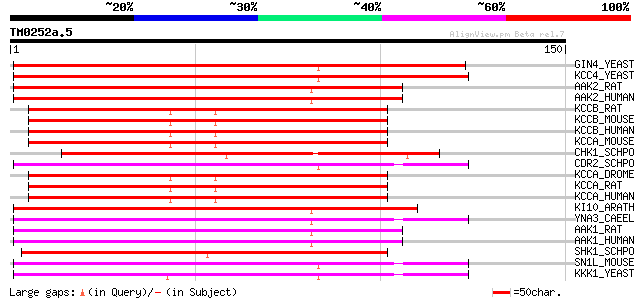
Score E
Sequences producing significant alignments: (bits) Value
GIN4_YEAST (Q12263) Serine/threonine-protein kinase GIN4 (EC 2.7... 100 2e-21
KCC4_YEAST (P25389) Probable serine/threonine-protein kinase KCC... 99 3e-21
AAK2_RAT (Q09137) 5'-AMP-activated protein kinase, catalytic alp... 96 3e-20
AAK2_HUMAN (P54646) 5'-AMP-activated protein kinase, catalytic a... 95 7e-20
KCCB_RAT (P08413) Calcium/calmodulin-dependent protein kinase ty... 91 9e-19
KCCB_MOUSE (P28652) Calcium/calmodulin-dependent protein kinase ... 91 9e-19
KCCB_HUMAN (Q13554) Calcium/calmodulin-dependent protein kinase ... 91 9e-19
KCCA_MOUSE (P11798) Calcium/calmodulin-dependent protein kinase ... 91 1e-18
CHK1_SCHPO (P34208) Serine/threonine-protein kinase chk1 (EC 2.7... 91 1e-18
CDR2_SCHPO (P87050) Mitosis inducer protein kinase cdr2 (EC 2.7.... 89 3e-18
KCCA_DROME (Q00168) Calcium/calmodulin-dependent protein kinase ... 89 5e-18
KCCA_RAT (P11275) Calcium/calmodulin-dependent protein kinase ty... 88 6e-18
KCCA_HUMAN (Q9UQM7) Calcium/calmodulin-dependent protein kinase ... 88 6e-18
KI10_ARATH (Q38997) SNF1-related protein kinase KIN10 (EC 2.7.1.... 88 8e-18
YNA3_CAEEL (P45894) Putative serine/threonine-protein kinase PAR... 87 1e-17
AAK1_RAT (P54645) 5'-AMP-activated protein kinase, catalytic alp... 87 1e-17
AAK1_HUMAN (Q13131) 5'-AMP-activated protein kinase, catalytic a... 87 1e-17
SHK1_SCHPO (P50527) Serine/threonine-protein kinase pak1/shk1 (E... 86 3e-17
SN1L_MOUSE (Q60670) Probable serine/threonine-protein kinase SNF... 86 4e-17
KKK1_YEAST (P34244) Probable serine/threonine-protein kinase YKL... 86 4e-17
>GIN4_YEAST (Q12263) Serine/threonine-protein kinase GIN4 (EC
2.7.1.37)
Length = 1142
Score = 99.8 bits (247), Expect = 2e-21
Identities = 48/123 (39%), Positives = 74/123 (60%), Gaps = 1/123 (0%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
GP+PE E +Q++ +++CH LG+ H D+KP+N+L ++K+ADFG A +G+
Sbjct: 126 GPLPEHEAIRFFRQIIIGVSYCHALGIVHRDLKPENLLLDHKYNIKIADFGMAALETEGK 185
Query: 62 RMSGVVGTPYYVAPEVLMGWE-NGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLLLLRI 120
+ G+P+Y APE++ G G DVWSCGVI A++ G L F E + LLL++
Sbjct: 186 LLETSCGSPHYAAPEIVSGIPYQGFASDVWSCGVILFALLTGRLPFDEEDGNIRTLLLKV 245
Query: 121 S*G 123
G
Sbjct: 246 QKG 248
>KCC4_YEAST (P25389) Probable serine/threonine-protein kinase KCC4
(EC 2.7.1.37)
Length = 1037
Score = 99.0 bits (245), Expect = 3e-21
Identities = 49/124 (39%), Positives = 76/124 (60%), Gaps = 1/124 (0%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
GP+PE E + +Q++ I++CH LG+ H D+KP+N+L + ++K+ADFG A D
Sbjct: 122 GPLPEREAINCFRQIIIGISYCHALGIVHRDLKPENLLLDSFYNIKIADFGMAALQTDAD 181
Query: 62 RMSGVVGTPYYVAPEVLMGWE-NGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLLLLRI 120
+ G+P+Y APE++ G G DVWSCGVI A++ G L F E+ + LLL++
Sbjct: 182 LLETSCGSPHYAAPEIVSGLPYEGFASDVWSCGVILFALLTGRLPFDEENGNVRDLLLKV 241
Query: 121 S*GR 124
G+
Sbjct: 242 QKGQ 245
>AAK2_RAT (Q09137) 5'-AMP-activated protein kinase, catalytic
alpha-2 chain (EC 2.7.1.-) (AMPK alpha-2 chain)
Length = 552
Score = 95.9 bits (237), Expect = 3e-20
Identities = 48/106 (45%), Positives = 65/106 (61%), Gaps = 1/106 (0%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + EVE L +Q+L A+ +CHR V H D+KP+NVL A + K+ADFG + DG
Sbjct: 109 GRVEEVEARRLFQQILSAVDYCHRHMVVHRDLKPENVLLDAQMNAKIADFGLSNMMSDGE 168
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRF 106
+ G+P Y APEV+ G G +VD+WSCGVI A++ G L F
Sbjct: 169 FLRTSCGSPNYAAPEVISGRLYAGPEVDIWSCGVILYALLCGTLPF 214
>AAK2_HUMAN (P54646) 5'-AMP-activated protein kinase, catalytic
alpha-2 chain (EC 2.7.1.-) (AMPK alpha-2 chain)
Length = 552
Score = 94.7 bits (234), Expect = 7e-20
Identities = 47/106 (44%), Positives = 65/106 (60%), Gaps = 1/106 (0%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E+E L +Q+L A+ +CHR V H D+KP+NVL A + K+ADFG + DG
Sbjct: 109 GRVEEMEARRLFQQILSAVDYCHRHMVVHRDLKPENVLLDAHMNAKIADFGLSNMMSDGE 168
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRF 106
+ G+P Y APEV+ G G +VD+WSCGVI A++ G L F
Sbjct: 169 FLRTSCGSPNYAAPEVISGRLYAGPEVDIWSCGVILYALLCGTLPF 214
>KCCB_RAT (P08413) Calcium/calmodulin-dependent protein kinase type
II beta chain (EC 2.7.1.123) (CaM-kinase II beta chain)
(CaM kinase II beta subunit) (CaMK-II beta subunit)
Length = 542
Score = 90.9 bits (224), Expect = 9e-19
Identities = 45/101 (44%), Positives = 66/101 (64%), Gaps = 4/101 (3%)
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGA---GGDLKLADFGSA-EWFGDGR 61
E + + ++Q+LEA+ HCH++GV H D+KP+N+L + G +KLADFG A E GD +
Sbjct: 110 EADASHCIQQILEAVLHCHQMGVVHRDLKPENLLLASKCKGAAVKLADFGLAIEVQGDQQ 169
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G GTP Y++PEVL G+ VD+W+CGVI ++ G
Sbjct: 170 AWFGFAGTPGYLSPEVLRKEAYGKPVDIWACGVILYILLVG 210
>KCCB_MOUSE (P28652) Calcium/calmodulin-dependent protein kinase
type II beta chain (EC 2.7.1.123) (CaM-kinase II beta
chain) (CaM kinase II beta subunit) (CaMK-II beta
subunit)
Length = 542
Score = 90.9 bits (224), Expect = 9e-19
Identities = 45/101 (44%), Positives = 66/101 (64%), Gaps = 4/101 (3%)
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGA---GGDLKLADFGSA-EWFGDGR 61
E + + ++Q+LEA+ HCH++GV H D+KP+N+L + G +KLADFG A E GD +
Sbjct: 110 EADASHCIQQILEAVLHCHQMGVVHRDLKPENLLLASKCKGAAVKLADFGLAIEVQGDQQ 169
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G GTP Y++PEVL G+ VD+W+CGVI ++ G
Sbjct: 170 AWFGFAGTPGYLSPEVLRKEAYGKPVDIWACGVILYILLVG 210
>KCCB_HUMAN (Q13554) Calcium/calmodulin-dependent protein kinase
type II beta chain (EC 2.7.1.123) (CaM-kinase II beta
chain) (CaM kinase II beta subunit) (CaMK-II beta
subunit)
Length = 664
Score = 90.9 bits (224), Expect = 9e-19
Identities = 45/101 (44%), Positives = 66/101 (64%), Gaps = 4/101 (3%)
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGA---GGDLKLADFGSA-EWFGDGR 61
E + + ++Q+LEA+ HCH++GV H D+KP+N+L + G +KLADFG A E GD +
Sbjct: 110 EADASHCIQQILEAVLHCHQMGVVHRDLKPENLLLASKCKGAAVKLADFGLAIEVQGDQQ 169
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G GTP Y++PEVL G+ VD+W+CGVI ++ G
Sbjct: 170 AWFGFAGTPGYLSPEVLRKEAYGKPVDIWACGVILYILLVG 210
>KCCA_MOUSE (P11798) Calcium/calmodulin-dependent protein kinase
type II alpha chain (EC 2.7.1.123) (CaM-kinase II alpha
chain) (CaM kinase II alpha subunit) (CaMK-II alpha
subunit)
Length = 478
Score = 90.5 bits (223), Expect = 1e-18
Identities = 45/101 (44%), Positives = 67/101 (65%), Gaps = 4/101 (3%)
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGA---GGDLKLADFGSA-EWFGDGR 61
E + + ++Q+LEA+ HCH++GV H D+KP+N+L + G +KLADFG A E G+ +
Sbjct: 109 EADASHCIQQILEAVLHCHQMGVVHRDLKPENLLLASKLKGAAVKLADFGLAIEVEGEQQ 168
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
R G GTP Y++PEVL G+ VD+W+CGVI ++ G
Sbjct: 169 RWFGFAGTPGYLSPEVLRKDPYGKPVDLWACGVILYILLVG 209
>CHK1_SCHPO (P34208) Serine/threonine-protein kinase chk1 (EC
2.7.1.37) (Checkpoint kinase 1)
Length = 496
Score = 90.5 bits (223), Expect = 1e-18
Identities = 47/107 (43%), Positives = 69/107 (63%), Gaps = 6/107 (5%)
Query: 15 QLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWF---GDGRRMSGVVGTPY 71
QL+E I+ H GVAH D+KP+N+L G+LK++DFG A F G R ++ VG+P
Sbjct: 120 QLMEGISFMHSKGVAHRDLKPENILLDYNGNLKISDFGFASLFSYKGKSRLLNSPVGSPP 179
Query: 72 YVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRF--PLESSGMFLL 116
Y APE+ ++ G KVDVWSCG+I A++ GN + + ++G +LL
Sbjct: 180 YAAPEITQQYD-GSKVDVWSCGIILFALLLGNTPWDEAISNTGDYLL 225
>CDR2_SCHPO (P87050) Mitosis inducer protein kinase cdr2 (EC
2.7.1.-)
Length = 775
Score = 89.4 bits (220), Expect = 3e-18
Identities = 43/124 (34%), Positives = 73/124 (58%), Gaps = 3/124 (2%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G E + A + Q+L + +CH+L + H D+KP+N+ A G +K+ +FG A G+
Sbjct: 103 GSFTEQDTAKFLWQILCGLEYCHKLHICHRDLKPENLYLDAHGSIKIGEFGMASIQQPGK 162
Query: 62 RMSGVVGTPYYVAPEVLMGWE-NGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLLLLRI 120
+ G+P+Y +PE++MG +G D+WSCG+IF A++ G L P + + LLL++
Sbjct: 163 LLQTSCGSPHYASPEIIMGRSYDGCASDIWSCGIIFFALLTGKL--PFDDDNIRSLLLKV 220
Query: 121 S*GR 124
G+
Sbjct: 221 CQGQ 224
>KCCA_DROME (Q00168) Calcium/calmodulin-dependent protein kinase
type II alpha chain (EC 2.7.1.123) (CaM-kinase II alpha
chain)
Length = 530
Score = 88.6 bits (218), Expect = 5e-18
Identities = 44/101 (43%), Positives = 65/101 (63%), Gaps = 4/101 (3%)
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGA---GGDLKLADFGSA-EWFGDGR 61
E + + ++Q+LE++ HCH+ GV H D+KP+N+L + G +KLADFG A E GD +
Sbjct: 110 EADASHCIQQILESVNHCHQNGVVHRDLKPENLLLASKAKGAAVKLADFGLAIEVQGDHQ 169
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G GTP Y++PEVL G+ VD+W+CGVI ++ G
Sbjct: 170 AWFGFAGTPGYLSPEVLKKEPYGKSVDIWACGVILYILLVG 210
>KCCA_RAT (P11275) Calcium/calmodulin-dependent protein kinase type
II alpha chain (EC 2.7.1.123) (CaM-kinase II alpha
chain) (CaM kinase II alpha subunit) (CaMK-II alpha
subunit)
Length = 478
Score = 88.2 bits (217), Expect = 6e-18
Identities = 44/101 (43%), Positives = 66/101 (64%), Gaps = 4/101 (3%)
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGA---GGDLKLADFGSA-EWFGDGR 61
E + + ++Q+LEA+ HCH++GV H D+KP+N+L + G +KLADFG A E G+ +
Sbjct: 109 EADASHCIQQILEAVLHCHQMGVVHRDLKPENLLLASKLKGAAVKLADFGLAIEVEGEQQ 168
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G GTP Y++PEVL G+ VD+W+CGVI ++ G
Sbjct: 169 AWFGFAGTPGYLSPEVLRKDPYGKPVDLWACGVILYILLVG 209
>KCCA_HUMAN (Q9UQM7) Calcium/calmodulin-dependent protein kinase
type II alpha chain (EC 2.7.1.123) (CaM-kinase II alpha
chain) (CaM kinase II alpha subunit) (CaMK-II alpha
subunit)
Length = 478
Score = 88.2 bits (217), Expect = 6e-18
Identities = 44/101 (43%), Positives = 66/101 (64%), Gaps = 4/101 (3%)
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGA---GGDLKLADFGSA-EWFGDGR 61
E + + ++Q+LEA+ HCH++GV H D+KP+N+L + G +KLADFG A E G+ +
Sbjct: 109 EADASHCIQQILEAVLHCHQMGVVHRDLKPENLLLASKLKGAAVKLADFGLAIEVEGEQQ 168
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G GTP Y++PEVL G+ VD+W+CGVI ++ G
Sbjct: 169 AWFGFAGTPGYLSPEVLRKDPYGKPVDLWACGVILYILLVG 209
>KI10_ARATH (Q38997) SNF1-related protein kinase KIN10 (EC 2.7.1.-)
(AKIN10)
Length = 535
Score = 87.8 bits (216), Expect = 8e-18
Identities = 44/110 (40%), Positives = 66/110 (60%), Gaps = 1/110 (0%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E E + +Q++ + +CHR V H D+KP+N+L + ++K+ADFG + DG
Sbjct: 135 GRLQEDEARNFFQQIISGVEYCHRNMVVHRDLKPENLLLDSKCNVKIADFGLSNIMRDGH 194
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
+ G+P Y APEV+ G G +VDVWSCGVI A++ G L F E+
Sbjct: 195 FLKTSCGSPNYAAPEVISGKLYAGPEVDVWSCGVILYALLCGTLPFDDEN 244
>YNA3_CAEEL (P45894) Putative serine/threonine-protein kinase PAR2.3
(EC 2.7.1.37)
Length = 622
Score = 87.4 bits (215), Expect = 1e-17
Identities = 44/124 (35%), Positives = 71/124 (56%), Gaps = 3/124 (2%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G +P E +Q++ +++CH + H D+KP+N+L A ++K+ADFG + + DG
Sbjct: 117 GALPIRESRRYFQQIISGVSYCHNHMIVHRDLKPENLLLDANKNIKIADFGLSNYMTDGD 176
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLLLLRI 120
+S G+P Y APE++ G +VD+WSCGVI A++ G L P + + L +I
Sbjct: 177 LLSTACGSPNYAAPELISNKLYVGPEVDLWSCGVILYAMLCGTL--PFDDQNVPTLFAKI 234
Query: 121 S*GR 124
GR
Sbjct: 235 KSGR 238
>AAK1_RAT (P54645) 5'-AMP-activated protein kinase, catalytic
alpha-1 chain (EC 2.7.1.-) (AMPK alpha-1 chain)
Length = 548
Score = 87.0 bits (214), Expect = 1e-17
Identities = 45/106 (42%), Positives = 62/106 (58%), Gaps = 1/106 (0%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E E L +Q+L + +CHR V H D+KP+NVL A + K+ADFG + DG
Sbjct: 109 GRLDEKESRRLFQQILSGVDYCHRHMVVHRDLKPENVLLDAHMNAKIADFGLSNMMSDGE 168
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRF 106
+ G+P Y APEV+ G G +VD+WS GVI A++ G L F
Sbjct: 169 FLRTSCGSPNYAAPEVISGRLYAGPEVDIWSSGVILYALLCGTLPF 214
>AAK1_HUMAN (Q13131) 5'-AMP-activated protein kinase, catalytic
alpha-1 chain (EC 2.7.1.-) (AMPK alpha-1 chain)
Length = 550
Score = 87.0 bits (214), Expect = 1e-17
Identities = 45/106 (42%), Positives = 62/106 (58%), Gaps = 1/106 (0%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E E L +Q+L + +CHR V H D+KP+NVL A + K+ADFG + DG
Sbjct: 111 GRLDEKESRRLFQQILSGVDYCHRHMVVHRDLKPENVLLDAHMNAKIADFGLSNMMSDGE 170
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRF 106
+ G+P Y APEV+ G G +VD+WS GVI A++ G L F
Sbjct: 171 FLRTSCGSPNYAAPEVISGRLYAGPEVDIWSSGVILYALLCGTLPF 216
>SHK1_SCHPO (P50527) Serine/threonine-protein kinase pak1/shk1 (EC
2.7.1.37)
Length = 658
Score = 85.9 bits (211), Expect = 3e-17
Identities = 41/100 (41%), Positives = 64/100 (64%), Gaps = 1/100 (1%)
Query: 4 IPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFG-SAEWFGDGRR 62
+ E ++A++ K+ LE + H H G+ H D+K DN+L GD+KL DFG A+ + +
Sbjct: 477 LSEGQIAAICKETLEGLQHLHENGIVHRDIKSDNILLSLQGDIKLTDFGFCAQIDSNMTK 536
Query: 63 MSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
+ +VGTPY++APEV+ E G KVDVWS G++ ++ G
Sbjct: 537 RTTMVGTPYWMAPEVVTRKEYGFKVDVWSLGIMAIEMVEG 576
>SN1L_MOUSE (Q60670) Probable serine/threonine-protein kinase SNF1LK
(EC 2.7.1.37) (HRT-20) (Myocardial SNF1-like kinase)
Length = 779
Score = 85.5 bits (210), Expect = 4e-17
Identities = 46/124 (37%), Positives = 69/124 (55%), Gaps = 3/124 (2%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E E Q+L A+ +CH + H D+K +N+L + D+KLADFG ++ G
Sbjct: 119 GHLSENEARQKFWQILSAVEYCHNHHIVHRDLKTENLLLDSNMDIKLADFGFGNFYKPGE 178
Query: 62 RMSGVVGTPYYVAPEVLMGWE-NGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLLLLRI 120
+S VG+P Y APEV G E G ++DVWS GV+ ++ G+L P + + L R+
Sbjct: 179 PLSTCVGSPPYAAPEVFEGKEYEGPQLDVWSLGVVLYVLVCGSL--PFDGPNLPTLRQRV 236
Query: 121 S*GR 124
GR
Sbjct: 237 LEGR 240
>KKK1_YEAST (P34244) Probable serine/threonine-protein kinase
YKL101W (EC 2.7.1.37)
Length = 1518
Score = 85.5 bits (210), Expect = 4e-17
Identities = 43/125 (34%), Positives = 72/125 (57%), Gaps = 4/125 (3%)
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFG-AGGDLKLADFGSAEWFGDG 60
G +PE E KQ++E +++CH + H D+KP+N+L +K+ADFG A
Sbjct: 209 GKLPEREAIHYFKQIVEGVSYCHSFNICHRDLKPENLLLDKKNRRIKIADFGMAALELPN 268
Query: 61 RRMSGVVGTPYYVAPEVLMGWE-NGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLLLLR 119
+ + G+P+Y +PE++MG +G DVWSCG++ A++ G+L P + LLL+
Sbjct: 269 KLLKTSCGSPHYASPEIVMGRPYHGGPSDVWSCGIVLFALLTGHL--PFNDDNIKKLLLK 326
Query: 120 IS*GR 124
+ G+
Sbjct: 327 VQSGK 331
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.334 0.150 0.505
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,746,822
Number of Sequences: 164201
Number of extensions: 751623
Number of successful extensions: 5479
Number of sequences better than 10.0: 1623
Number of HSP's better than 10.0 without gapping: 1334
Number of HSP's successfully gapped in prelim test: 289
Number of HSP's that attempted gapping in prelim test: 2588
Number of HSP's gapped (non-prelim): 1713
length of query: 150
length of database: 59,974,054
effective HSP length: 100
effective length of query: 50
effective length of database: 43,553,954
effective search space: 2177697700
effective search space used: 2177697700
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0252a.5


