

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0243.9
(310 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
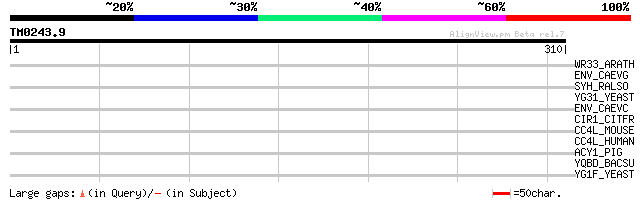
Score E
Sequences producing significant alignments: (bits) Value
WR33_ARATH (Q8S8P5) Probable WRKY transcription factor 33 (WRKY ... 34 0.41
ENV_CAEVG (P31627) Env polyprotein precursor (Coat polyprotein) ... 33 0.90
SYH_RALSO (Q8Y029) Histidyl-tRNA synthetase (EC 6.1.1.21) (Histi... 33 1.2
YG31_YEAST (P53269) Hypothetical 27.2 kDa protein in SHY1-SPT6 i... 32 2.6
ENV_CAEVC (P31626) ENV polyprotein precursor (Coat polyprotein) ... 32 2.6
CIR1_CITFR (P23182) Citrolysin protein 1 30 5.8
CC4L_MOUSE (Q8K3P5) Carbon catabolite repressor protein 4 homolo... 30 5.8
CC4L_HUMAN (Q9ULM6) Carbon catabolite repressor protein 4 homolo... 30 5.8
ACY1_PIG (P37111) Aminoacylase-1 (EC 3.5.1.14) (N-acyl-L-amino-a... 30 5.8
YQBD_BACSU (P45920) Hypothetical protein yqbD 30 7.6
YG1F_YEAST (P53214) Hypothetical 57.5 kDa protein in VMA7-RPS25A... 30 10.0
>WR33_ARATH (Q8S8P5) Probable WRKY transcription factor 33 (WRKY
DNA-binding protein 33)
Length = 512
Score = 34.3 bits (77), Expect = 0.41
Identities = 29/103 (28%), Positives = 52/103 (50%), Gaps = 10/103 (9%)
Query: 157 EKGVLEGAFQTLHSGYFRLFPFSFF--SLPLSPPLTSTISSSTGTISSSSTTKKKTSRGV 214
+KG+ EG ++ F LF FSF S +S P T+T +++T T ++SS + + +
Sbjct: 91 QKGINEG--DKSNNNNFNLFDFSFHTQSSGVSAPTTTTTTTTTTTTTNSSIFQSQEQQKK 148
Query: 215 GRSEELQIWCESKGRVPDLVRQWLYGSDPALNGEEQW-WRNLG 256
+SE+ W +++ R + + Y GE+ + WR G
Sbjct: 149 NQSEQ---WSQTETRPNN--QAVSYNGREQRKGEDGYNWRKYG 186
>ENV_CAEVG (P31627) Env polyprotein precursor (Coat polyprotein)
[Contains: Surface protein; Transmembrane protein]
Length = 942
Score = 33.1 bits (74), Expect = 0.90
Identities = 20/66 (30%), Positives = 37/66 (55%), Gaps = 8/66 (12%)
Query: 229 RVPDLVRQWLYGSDPA--LNGEEQWWRNL-----GVSGSVSGSLLHLLHRLCFSLSESGA 281
++P VR W+ ++ + +NG +W+N G++G++ G L + H L F+LS++G
Sbjct: 318 KLPLTVRVWVKLANVSTWVNGTPPYWQNRINGSKGINGTLWGQLSGM-HHLGFNLSQTGK 376
Query: 282 WLRKEG 287
W G
Sbjct: 377 WCNYTG 382
>SYH_RALSO (Q8Y029) Histidyl-tRNA synthetase (EC 6.1.1.21)
(Histidine--tRNA ligase) (HisRS)
Length = 432
Score = 32.7 bits (73), Expect = 1.2
Identities = 17/52 (32%), Positives = 27/52 (51%)
Query: 218 EELQIWCESKGRVPDLVRQWLYGSDPALNGEEQWWRNLGVSGSVSGSLLHLL 269
E+ Q++ G V D+V + +Y ALNGE+ R G + +V + H L
Sbjct: 46 EQTQLFVRGIGEVTDIVEKEMYSFTDALNGEQLTMRPEGTAAAVRAVIEHNL 97
>YG31_YEAST (P53269) Hypothetical 27.2 kDa protein in SHY1-SPT6
intergenic region
Length = 259
Score = 31.6 bits (70), Expect = 2.6
Identities = 17/34 (50%), Positives = 22/34 (64%)
Query: 178 FSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTS 211
FS SL + P T SSS+ +ISSSS+ + KTS
Sbjct: 181 FSSSSLTMKPSCTFLASSSSSSISSSSSEESKTS 214
>ENV_CAEVC (P31626) ENV polyprotein precursor (Coat polyprotein)
[Contains: Surface protein; Transmembrane protein]
Length = 966
Score = 31.6 bits (70), Expect = 2.6
Identities = 17/57 (29%), Positives = 29/57 (50%), Gaps = 9/57 (15%)
Query: 234 VRQWLYGSDPALNGEEQWWRNLGVSGSVSGSL---LHLLHRLCFSLSESGAWLRKEG 287
V +W+ G+ P W + S ++G+L L+ +H L F+LS++G W G
Sbjct: 335 VSEWVNGTPP------DWQDRINGSKGINGTLWGELNSMHHLGFALSQNGKWCNYTG 385
>CIR1_CITFR (P23182) Citrolysin protein 1
Length = 618
Score = 30.4 bits (67), Expect = 5.8
Identities = 17/46 (36%), Positives = 22/46 (46%), Gaps = 4/46 (8%)
Query: 2 GVGGGVWDSVTGRG--WR--AAQRKVVSGVWWVGFGTVKEYYQKLD 43
GVGG VW V R WR + +V+ +WW YY +LD
Sbjct: 547 GVGGDVWRRVLVRAACWRQISYSAAIVNLMWWRSHDDEYSYYARLD 592
>CC4L_MOUSE (Q8K3P5) Carbon catabolite repressor protein 4 homolog
(EC 3.1.-.-) (Cytoplasmic deadenylase) (CCR4 carbon
catabolite repression 4-like) (Nocturnin homolog)
Length = 557
Score = 30.4 bits (67), Expect = 5.8
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 4/59 (6%)
Query: 225 ESKGRVPDLVRQW-LYGSDPALNGEEQWWRNLGVSG---SVSGSLLHLLHRLCFSLSES 279
+ K PD R + + S+ A NG++ W L +SG S+S SL L H LS++
Sbjct: 3 KEKYEPPDPRRMYTIMSSEEAANGKKSHWAELEISGKVRSLSSSLWSLTHLTALHLSDN 61
>CC4L_HUMAN (Q9ULM6) Carbon catabolite repressor protein 4 homolog
(EC 3.1.-.-) (Cytoplasmic deadenylase) (CCR4 carbon
catabolite repression 4-like) (Nocturnin homolog)
Length = 557
Score = 30.4 bits (67), Expect = 5.8
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 4/59 (6%)
Query: 225 ESKGRVPDLVRQW-LYGSDPALNGEEQWWRNLGVSG---SVSGSLLHLLHRLCFSLSES 279
+ K PD R + + S+ A NG++ W L +SG S+S SL L H LS++
Sbjct: 3 KEKYEPPDPRRMYTIMSSEEAANGKKSHWAELEISGKVRSLSASLWSLTHLTALHLSDN 61
>ACY1_PIG (P37111) Aminoacylase-1 (EC 3.5.1.14) (N-acyl-L-amino-acid
amidohydrolase) (ACY-1)
Length = 406
Score = 30.4 bits (67), Expect = 5.8
Identities = 20/78 (25%), Positives = 33/78 (41%), Gaps = 5/78 (6%)
Query: 218 EELQIWCESKGR--VPDLVRQWLYGSDPALNGEEQWWRNLGVSGSVSGSLLHLLHRLCFS 275
E+LQ WC++ G + V++W+ + + + WW SG L L +C
Sbjct: 286 EQLQSWCQAAGEGVTFEFVQKWMETQVTSTDDSDPWW--AAFSGVFKDMKLALELEIC-P 342
Query: 276 LSESGAWLRKEGCDGDGF 293
S ++R G GF
Sbjct: 343 ASTDARYIRAAGVPALGF 360
>YQBD_BACSU (P45920) Hypothetical protein yqbD
Length = 322
Score = 30.0 bits (66), Expect = 7.6
Identities = 18/55 (32%), Positives = 25/55 (44%)
Query: 83 FDCLSIFFTGDVQGKVEEADSRARRSGRSSRWFASHQFHIAVGDAIKDVMVLEGE 137
F+ L FF G Q EE ++A R +S H A+G+ + V EGE
Sbjct: 173 FNLLKNFFVGKQQQSYEEPVAKAGRKFSASNLQEIKNAHAALGNLLSQVETKEGE 227
>YG1F_YEAST (P53214) Hypothetical 57.5 kDa protein in VMA7-RPS25A
intergenic region
Length = 551
Score = 29.6 bits (65), Expect = 10.0
Identities = 17/38 (44%), Positives = 23/38 (59%)
Query: 173 FRLFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKT 210
F FP S L LS L+S +SSS+ +SSSST+ +
Sbjct: 83 FTSFPQSSSLLTLSSTLSSELSSSSMQVSSSSTSSSSS 120
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.323 0.139 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,324,660
Number of Sequences: 164201
Number of extensions: 1660696
Number of successful extensions: 4408
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 4393
Number of HSP's gapped (non-prelim): 18
length of query: 310
length of database: 59,974,054
effective HSP length: 110
effective length of query: 200
effective length of database: 41,911,944
effective search space: 8382388800
effective search space used: 8382388800
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0243.9


