

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
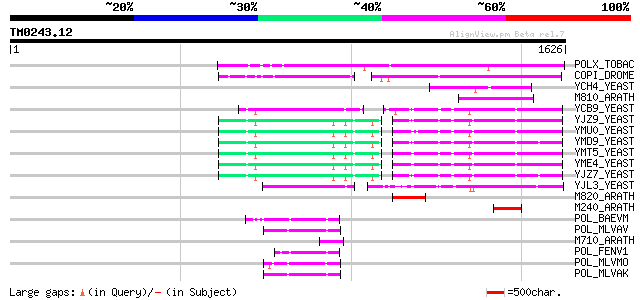
Query= TM0243.12
(1626 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 545 e-154
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 382 e-105
YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein 164 2e-39
M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810... 157 2e-37
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 120 4e-26
YJZ9_YEAST (P47100) Transposon Ty1 protein B 117 2e-25
YMU0_YEAST (Q04670) Transposon Ty1 protein B 115 1e-24
YMD9_YEAST (Q03434) Transposon Ty1 protein B 115 1e-24
YMT5_YEAST (Q04214) Transposon Ty1 protein B 114 2e-24
YME4_YEAST (Q04711) Transposon Ty1 protein B 114 2e-24
YJZ7_YEAST (P47098) Transposon Ty1 protein B 113 5e-24
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 110 2e-23
M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820... 99 1e-19
M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240... 69 8e-11
POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.2... 60 4e-08
POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.2... 57 5e-07
M710_ARATH (P92512) Hypothetical mitochondrial protein AtMg00710... 56 7e-07
POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse transcript... 55 1e-06
POL_MLVMO (P03355) Pol polyprotein [Contains: Protease (EC 3.4.2... 53 6e-06
POL_MLVAK (P03357) Pol polyprotein [Contains: Reverse transcript... 53 6e-06
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 545 bits (1404), Expect = e-154
Identities = 341/1048 (32%), Positives = 555/1048 (52%), Gaps = 49/1048 (4%)
Query: 610 WYLDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTICVDSSP----CIDNV 665
W +D+ S H T + +F G V G KI G G IC+ ++ + +V
Sbjct: 294 WVVDTAASHHATPVRDLFCRYVAGDFGTVKMGNTSYSKIAGIGDICIKTNVGCTLVLKDV 353
Query: 666 LLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGS-VLFNSKRKNNIYKIR--LSE 722
V L NL+S L GY+ F + R GS V+ + +Y+ + +
Sbjct: 354 RHVPDLRMNLISGIALDRDGYESYFANQKWRLTK---GSLVIAKGVARGTLYRTNAEICQ 410
Query: 723 LEAQNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKG 782
E + +SV+ +WH+R+GH S + + L+K +L+ K + C+ C G
Sbjct: 411 GELNAAQDEISVD----LWHKRMGHMSEKGLQILAKKSLIS---YAKGTTVKPCDYCLFG 463
Query: 783 KFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRK 842
K +V F+ + + L+L++ D+ GP++ ES+GG +Y + +DD SR WV L K
Sbjct: 464 KQHRVSFQTSSERKLNI-LDLVYSDVCGPMEIESMGGNKYFVTFIDDASRKLWVYILKTK 522
Query: 843 DESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQN 902
D+ VF F A V+ E ++ R+RSD GGE+ + +FE S+GI H+ + P TPQ N
Sbjct: 523 DQVFQVFQKFHALVERETGRKLKRLRSDNGGEYTSREFEEYCSSHGIRHEKTVPGTPQHN 582
Query: 903 GVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIK 962
GV ER NRT+ E R+ML+ + K FW E V TACY+ NR P+ + P +W N +
Sbjct: 583 GVAERMNRTIVEKVRSMLRMAKLPKSFWGEAVQTACYLINRSPSVPLAFEIPERVWTNKE 642
Query: 963 PNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHV 1022
+ S+ FGC + K++ K D KS C+ +GY D G+R ++ K + S V
Sbjct: 643 VSYSHLKVFGCRAFAHVPKEQRTKLDDKSIPCIFIGYGDEEFGYRLWDPVKKKVIRSRDV 702
Query: 1023 RFDD---KLDSDQSKLVEK--------FADLSINVSDKGKAPEEVEPEEDEPEE--EAGP 1069
F + + +D S+ V+ S N + +EV + ++P E E G
Sbjct: 703 VFRESEVRTAADMSEKVKNGIIPNFVTIPSTSNNPTSAESTTDEVSEQGEQPGEVIEQGE 762
Query: 1070 SNSQTLKKSRITAAHPKELILGNKDEPVRTRSAFRPSEETLLSLKGLVSLIEPKSIDEAL 1129
+ +++ ++ + E R S PS E +L + EP+S+ E L
Sbjct: 763 QLDEGVEEVEHPTQGEEQHQPLRRSERPRVESRRYPSTEYVL----ISDDREPESLKEVL 818
Query: 1130 ---QDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVVRNKARL 1186
+ + + AM+EE+ KN + LV+ P+ + KWVF+ K + +VR KARL
Sbjct: 819 SHPEKNQLMKAMQEEMESLQKNGTYKLVELPKGKRPLKCKWVFKLKKDGDCKLVRYKARL 878
Query: 1187 VAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYV 1246
V +G+ Q++GID+ E F+PV ++ +IR ++S + + ++ + Q+DVK+AFL+G + EE+Y+
Sbjct: 879 VVKGFEQKKGIDFDEIFSPVVKMTSIRTILSLAASLDLEVEQLDVKTAFLHGDLEEEIYM 938
Query: 1247 HQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTY- 1305
QP GFE K V KL KSLYGLKQAPR WY + SF+ +++ D ++ K +
Sbjct: 939 EQPEGFEVAGKKHMVCKLNKSLYGLKQAPRQWYMKFDSFMKSQTYLKTYSDPCVYFKRFS 998
Query: 1306 KDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQV--DQTPEGT 1363
+++ +I+ +YVDD++ ++ L + + F+M +G + LG+++ ++T
Sbjct: 999 ENNFIILLLYVDDMLIVGKDKGLIAKLKGDLSKSFDMKDLGPAQQILGMKIVRERTSRKL 1058
Query: 1364 YIHQSKYTKELLKKFNMLESTVAKTPMHPTCILEKE------DKSGKVCQKLYRGMIGSL 1417
++ Q KY + +L++FNM + TP+ L K+ ++ G + + Y +GSL
Sbjct: 1059 WLSQEKYIERVLERFNMKNAKPVSTPLAGHLKLSKKMCPTTVEEKGNMAKVPYSSAVGSL 1118
Query: 1418 LY-LTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSG 1476
+Y + +RPDI +V + +RF +P + H AVK ILRYL+GTT L + S+ L G
Sbjct: 1119 MYAMVCTRPDIAHAVGVVSRFLENPGKEHWEAVKWILRYLRGTTGDCLCF-GGSDPILKG 1177
Query: 1477 YCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLWM 1536
Y DAD AGD RKS++G +SW SK Q +ALST EAEYI+A +M+W+
Sbjct: 1178 YTDADMAGDIDNRKSSTGYLFTFSGGAISWQSKLQKCVALSTTEAEYIAATETGKEMIWL 1237
Query: 1537 KHQLEDYQILESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFV 1596
K L++ + + +YCD+ +AI LSKN + H+R KHI+V+YH+IR+ V L + +
Sbjct: 1238 KRFLQELGLHQKEYVVYCDSQSAIDLSKNSMYHARTKHIDVRYHWIREMVDDESLKVLKI 1297
Query: 1597 DTDHQWADIFTKSLAEDRFNFILKNLNM 1624
T+ AD+ TK + ++F + + M
Sbjct: 1298 STNENPADMLTKVVPRNKFELCKELVGM 1325
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 382 bits (982), Expect = e-105
Identities = 219/584 (37%), Positives = 340/584 (57%), Gaps = 33/584 (5%)
Query: 1060 EDEPEEEAGPSNSQTLKKSRITAAHPKEL------------ILGNKDEPVRTRSAFRPSE 1107
+D E G N ++S TA H KE+ I+ + E ++T+ +E
Sbjct: 813 DDHLNESKGSGNPNESRESE-TAEHLKEIGIDNPTKNDGIEIINRRSERLKTKPQISYNE 871
Query: 1108 E------TLLSLKGLVSLIEPKSIDEAL--QDKE-WILAMEEELNQFSKNDVWSLVKKPE 1158
E +L+ + + + P S DE DK W A+ ELN N+ W++ K+PE
Sbjct: 872 EDNSLNKVVLNAHTIFNDV-PNSFDEIQYRDDKSSWEEAINTELNAHKINNTWTITKRPE 930
Query: 1159 SVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISF 1218
+ +++ ++WVF K NE G+ +R KARLVA+G++Q+ IDY ETFAPVAR+ + R ++S
Sbjct: 931 NKNIVDSRWVFSVKYNELGNPIRYKARLVARGFTQKYQIDYEETFAPVARISSFRFILSL 990
Query: 1219 SVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAW 1278
+ +N+ +HQMDVK+AFLNG + EE+Y+ P G D+V KL K++YGLKQA R W
Sbjct: 991 VIQYNLKVHQMDVKTAFLNGTLKEEIYMRLPQGISCNS--DNVCKLNKAIYGLKQAARCW 1048
Query: 1279 YERLSSFLLENEFVRGKVDTTLFC--KTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMM 1336
+E L E EFV VD ++ K ++ + V +YVDD++ + + + F +
Sbjct: 1049 FEVFEQALKECEFVNSSVDRCIYILDKGNINENIYVLLYVDDVVIATGDMTRMNNFKRYL 1108
Query: 1337 QAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKELLKKFNMLESTVAKTPMHPTCIL 1396
+F M+ + E+K+F+GI+++ + Y+ QS Y K++L KFNM TP+ P+ I
Sbjct: 1109 MEKFRMTDLNEIKHFIGIRIEMQEDKIYLSQSAYVKKILSKFNMENCNAVSTPL-PSKIN 1167
Query: 1397 EKEDKSGKVCQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTAVKRILRY 1455
+ S + C R +IG L+Y + +RPD+ +V++ +R+ S +KR+LRY
Sbjct: 1168 YELLNSDEDCNTPCRSLIGCLMYIMLCTRPDLTTAVNILSRYSSKNNSELWQNLKRVLRY 1227
Query: 1456 LKGTTNLGLMYKK--TSEYKLSGYCDADYAGDRTERKSTSGNC-QFLGSNLVSWASKRQS 1512
LKGT ++ L++KK E K+ GY D+D+AG +RKST+G + NL+ W +KRQ+
Sbjct: 1228 LKGTIDMKLIFKKNLAFENKIIGYVDSDWAGSEIDRKSTTGYLFKMFDFNLICWNTKRQN 1287
Query: 1513 TIALSTAEAEYISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSR 1571
++A S+ EAEY++ + LW+K L I LE+ I IY DN IS++ NP H R
Sbjct: 1288 SVAASSTEAEYMALFEAVREALWLKFLLTSINIKLENPIKIYEDNQGCISIANNPSCHKR 1347
Query: 1572 AKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRF 1615
AKHI++KYHF R+ VQ V+ L+++ T++Q ADIFTK L RF
Sbjct: 1348 AKHIDIKYHFAREQVQNNVICLEYIPTENQLADIFTKPLPAARF 1391
Score = 182 bits (462), Expect = 7e-45
Identities = 123/411 (29%), Positives = 213/411 (50%), Gaps = 24/411 (5%)
Query: 612 LDSGCSRHMTGEKRMFRE-LKLKPGGEVGFGGNEKGKII-----GTGTICVDSSPCIDNV 665
LDSG S H+ ++ ++ + +++ P ++ ++G+ I G + D +++V
Sbjct: 291 LDSGASDHLINDESLYTDSVEVVPPLKIAVA--KQGEFIYATKRGIVRLRNDHEITLEDV 348
Query: 666 LLVDGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEA 725
L NL+S+ +L + G + F+ KS +S+ V+ NS NN+ I +A
Sbjct: 349 LFCKEAAGNLMSVKRLQEAGMSIEFD-KSGVTISKNGLMVVKNSGMLNNVPVINF---QA 404
Query: 726 QNVKCLLSVNEEQWVWHRRLGHASMRKISQLSKLNLVRG---LPNLKFASDALCEACQKG 782
++ N +WH R GH S K+ ++ + N+ L NL+ + + +CE C G
Sbjct: 405 YSINAKHKNNFR--LWHERFGHISDGKLLEIKRKNMFSDQSLLNNLELSCE-ICEPCLNG 461
Query: 783 KFTKVPFKA-KNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTR 841
K ++PFK K+ RPL ++H D+ GP+ ++ K Y ++ VD ++ + +
Sbjct: 462 KQARLPFKQLKDKTHIKRPLFVVHSDVCGPITPVTLDDKNYFVIFVDQFTHYCVTYLIKY 521
Query: 842 KDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQ 901
K + ++F F+A+ + ++V + D G E+ +++ GI++ + P TPQ
Sbjct: 522 KSDVFSMFQDFVAKSEAHFNLKVVYLYIDNGREYLSNEMRQFCVKKGISYHLTVPHTPQL 581
Query: 902 NGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPIL--NKTPYELWK 959
NGV ER RT+ E ARTM+ + K FW E V TA Y+ NRI R ++ +KTPYE+W
Sbjct: 582 NGVSERMIRTITEKARTMVSGAKLDKSFWGEAVLTATYLINRIPSRALVDSSKTPYEMWH 641
Query: 960 NIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYN 1010
N KP + + FG YV + K++ KFD KS K + +GY GF+ ++
Sbjct: 642 NKKPYLKHLRVFGATVYV-HIKNKQGKFDDKSFKSIFVGY--EPNGFKLWD 689
>YCH4_YEAST (P25600) Transposon Ty5-1 34.5 kDa hypothetical protein
Length = 308
Score = 164 bits (415), Expect = 2e-39
Identities = 97/309 (31%), Positives = 167/309 (53%), Gaps = 17/309 (5%)
Query: 1229 MDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLE 1288
MDV +AFLN + E +YV QPPGF +E+ PD+V++L +YGLKQAP W E +++ L +
Sbjct: 1 MDVDTAFLNSTMDEPIYVKQPPGFVNERNPDYVWELYGGMYGLKQAPLLWNEHINNTLKK 60
Query: 1289 NEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGEL 1348
F R + + L+ ++ D + + +YVDD++ + + + + + + M +G++
Sbjct: 61 IGFCRHEGEHGLYFRSTSDGPIYIGVYVDDLLVAAPSPKIYDRVKQELTKLYSMKDLGKV 120
Query: 1349 KYFLGIQVDQTPEG-------TYIHQSKYTKELLKKFNMLESTVAKT-PMHPTCILEKED 1400
FLG+ + Q+ G YI ++ E + F + ++ + + P+ T +D
Sbjct: 121 DKFLGLNIHQSTNGDITLSLQDYIAKAASESE-INTFKLTQTPLCNSKPLFETTSPHLKD 179
Query: 1401 KSGKVCQKLYRGMIGSLLY-LTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGT 1459
+ Y+ ++G LL+ RPDI + V L +RF +PR HL + +R+LRYL T
Sbjct: 180 ITP------YQSIVGQLLFCANTGRPDISYPVSLLSRFLREPRAIHLESARRVLRYLYTT 233
Query: 1460 TNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKR-QSTIALST 1518
++ L Y+ S+ L+ YCDA + ST G L V+W+SK+ + I + +
Sbjct: 234 RSMCLKYRSGSQVALTVYCDASHGAIHDLPHSTGGYVTLLAGAPVTWSSKKLKGVIPVPS 293
Query: 1519 AEAEYISAA 1527
EAEYI+A+
Sbjct: 294 TEAEYITAS 302
>M810_ARATH (P92519) Hypothetical mitochondrial protein AtMg00810
(ORF240b)
Length = 240
Score = 157 bits (398), Expect = 2e-37
Identities = 76/222 (34%), Positives = 130/222 (58%), Gaps = 1/222 (0%)
Query: 1314 IYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKE 1373
+YVDDI+ ++ +L + + F M +G + YFLGIQ+ P G ++ Q+KY ++
Sbjct: 5 LYVDDILLTGSSNTLLNMLIFQLSSTFSMKDLGPVHYFLGIQIKTHPSGLFLSQTKYAEQ 64
Query: 1374 LLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLTASRPDILFSVHL 1433
+L ML+ TP+ P + + +R ++G+L YLT +RPDI ++V++
Sbjct: 65 ILNNAGMLDCKPMSTPL-PLKLNSSVSTAKYPDPSDFRSIVGALQYLTLTRPDISYAVNI 123
Query: 1434 CARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTS 1493
+ +P +KR+LRY+KGT GL K S+ + +CD+D+AG + R+ST+
Sbjct: 124 VCQRMHEPTLADFDLLKRVLRYVKGTIFHGLYIHKNSKLNVQAFCDSDWAGCTSTRRSTT 183
Query: 1494 GNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQMLW 1535
G C FLG N++SW++KRQ T++ S+ E EY + A+ + ++ W
Sbjct: 184 GFCTFLGCNIISWSAKRQPTVSRSSTETEYRALALTAAELTW 225
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 120 bits (300), Expect = 4e-26
Identities = 113/388 (29%), Positives = 178/388 (45%), Gaps = 36/388 (9%)
Query: 671 LTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKI--------RLSE 722
+ ++LLS+S+LA++ F + + + + DG+VL + + Y + +S+
Sbjct: 518 IAYDLLSLSELANQNITACFTRNT---LERSDGTVLAPIVKHGDFYWLSKKYLIPSHISK 574
Query: 723 LEAQNVKCLLSVNEEQW-VWHRRLGHASMRKISQLSKLNLVRGLPNLKF----ASDALCE 777
L NV SVN+ + + HR LGHA+ R I + K N V L AS C
Sbjct: 575 LTINNVNKSKSVNKYPYPLIHRMLGHANFRSIQKSLKKNAVTYLKESDIEWSNASTYQCP 634
Query: 778 ACQKGKFTK---VPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWT 834
C GK TK V + P + LH D+FGPV Y + D+ +R+
Sbjct: 635 DCLIGKSTKHRHVKGSRLKYQESYEPFQYLHTDIFGPVHHLPKSAPSYFISFTDEKTRFQ 694
Query: 835 WVKFL-TRKDESHA-VFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHD 892
WV L R++ES VF++ +A ++N+ R++ ++ D G E+ N F + GI
Sbjct: 695 WVYPLHDRREESILNVFTSILAFIKNQFNARVLVIQMDRGSEYTNKTLHKFFTNRGITAC 754
Query: 893 FSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNK 952
++ + +GV ER NRTL RT+L +G+ H W V + I+N + V P +K
Sbjct: 755 YTTTADSRAHGVAERLNRTLLNDCRTLLHCSGLPNHLWFSAVEFSTIIRNSL-VSPKNDK 813
Query: 953 TPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGY----SDRSKGFRF 1008
+ + +I+ PFG V N H D+K + GY S S G+
Sbjct: 814 SARQHAGLAGLDITTILPFGQPVIVNN-----HNPDSKIHPRGIPGYALHPSRNSYGYII 868
Query: 1009 YNTD-AKTIEESIHVRFDDKLDSDQSKL 1035
Y KT++ + +V DK QSKL
Sbjct: 869 YLPSLKKTVDTTNYVILQDK----QSKL 892
Score = 112 bits (280), Expect = 8e-24
Identities = 129/566 (22%), Positives = 257/566 (44%), Gaps = 63/566 (11%)
Query: 1095 EPVRTRSAFRPSEETLLSLKGLVSLIEPKSI---DEAL-------QDKEWILAMEEELNQ 1144
EP R++ + ++KG+ S+ ++ DEA+ + ++ A +E++Q
Sbjct: 1220 EPPRSKKRIN----LIAAIKGVKSIKPVRTTLRYDEAITYNKDNKEKDRYVEAYHKEISQ 1275
Query: 1145 FSKNDVWSLVK-----KPESVHVIGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDY 1199
K + W K + VI + ++F N+K D +KAR VA+G Q
Sbjct: 1276 LLKMNTWDTNKYYDRNDIDPKKVINSMFIF----NKKRDGT-HKARFVARGDIQHPDTYD 1330
Query: 1200 TETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPP--GFEDEKK 1257
++ + A+ +S +++++ + Q+D+ SA+L I EE+Y+ PP G D+
Sbjct: 1331 SDMQSNTVHHYALMTSLSIALDNDYYITQLDISSAYLYADIKEELYIRPPPHLGLNDK-- 1388
Query: 1258 PDHVFKLKKSLYGLKQAPRAWYERLSSFLL---ENEFVRGKVDTTLFCKTYKDDILIVQI 1314
+ +L+KSLYGLKQ+ WYE + S+L+ + + VRG + +K+ + + +
Sbjct: 1389 ---LLRLRKSLYGLKQSGANWYETIKSYLINCCDMQEVRG------WSCVFKNSQVTICL 1439
Query: 1315 YVDDIIFGSANQSLCKEFSEMMQAEFEMSMM------GELKY-FLGIQVD-QTPEGTYIH 1366
+VDD+I S + + K+ ++ +++ ++ E++Y LG+++ Q + +
Sbjct: 1440 FVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESDNEIQYDILGLEIKYQRSKYMKLG 1499
Query: 1367 QSKYTKELLKKFNMLESTVAK---TPMHPTCILEKED---KSGKVCQKLY--RGMIGSLL 1418
K E L K N+ + K P P +++++ + +K++ + +IG
Sbjct: 1500 MEKSLTEKLPKLNVPLNPKGKKLRAPGQPGHYIDQDELEIDEDEYKEKVHEMQKLIGLAS 1559
Query: 1419 YLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTS----EYK 1473
Y+ R D+L+ ++ A+ P L +++++ T + L++ K + K
Sbjct: 1560 YVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELIQFMWDTRDKQLIWHKNKPTKPDNK 1619
Query: 1474 LSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASKRQSTIALSTAEAEYISAAICSTQM 1533
L DA Y G++ KS GN L ++ S + S ST EAE + + +
Sbjct: 1620 LVAISDASY-GNQPYYKSQIGNIFLLNGKVIGGKSTKASLTCTSTTEAEIHAVSEAIPLL 1678
Query: 1534 LWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS-RAKHIEVKYHFIRDYVQKGVLL 1592
+ H +++ + D+ + IS+ K+ R + K +RD V L
Sbjct: 1679 NNLSHLVQELNKKPIIKGLLTDSRSTISIIKSTNEEKFRNRFFGTKAMRLRDEVSGNNLY 1738
Query: 1593 LKFVDTDHQWADIFTKSLAEDRFNFI 1618
+ +++T AD+ TK L F +
Sbjct: 1739 VYYIETKKNIADVMTKPLPIKTFKLL 1764
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 117 bits (294), Expect = 2e-25
Identities = 133/531 (25%), Positives = 239/531 (44%), Gaps = 50/531 (9%)
Query: 1121 EPKSIDEALQDKE-WILAMEEELNQFSKNDVWSLV-----KKPESVHVIGTKWVFRNKLN 1174
E + ++ +++KE +I A +E+NQ K W K+ + VI + ++F N
Sbjct: 1236 EAITYNKDIKEKEKYIEAYHKEVNQLLKMKTWDTDEYYDRKEIDPKRVINSMFIF----N 1291
Query: 1175 EKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSA 1234
+K D +KAR VA+G Q + + A+ +S ++++N + Q+D+ SA
Sbjct: 1292 KKRDGT-HKARFVARGDIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSA 1350
Query: 1235 FLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EF 1291
+L I EE+Y+ PP D + +LKKSLYGLKQ+ WYE + S+L++ E
Sbjct: 1351 YLYADIKEELYIRPPPHLGMN---DKLIRLKKSLYGLKQSGANWYETIKSYLIQQCGMEE 1407
Query: 1292 VRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMG----- 1346
VRG + +K+ + + ++VDD++ S N + K E ++ +++ ++
Sbjct: 1408 VRG------WSCVFKNSQVTICLFVDDMVLFSKNLNSNKRIIEKLKMQYDTKIINLGESD 1461
Query: 1347 -ELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCI---- 1395
E++Y LG+++ + G Y + E + K N+ + P P
Sbjct: 1462 EEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQ 1520
Query: 1396 -LEKEDKSGKVCQKLYRGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRIL 1453
LE E+ K+ + +IG Y+ R D+L+ ++ A+ P + L ++
Sbjct: 1521 ELELEEDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELI 1580
Query: 1454 RYLKGTTNLGLMYKKTSEY----KLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASK 1509
+++ T + L++ K+ KL DA Y G++ KS GN L ++ S
Sbjct: 1581 QFIWNTRDKQLIWHKSKPVKPTNKLVVISDASY-GNQPYYKSQIGNIYLLNGKVIGGKST 1639
Query: 1510 RQSTIALSTAEAEY--ISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPI 1567
+ S ST EAE IS ++ L Q D + + + +T +I +S N
Sbjct: 1640 KASLTCTSTTEAEIHAISESVPLLNNLSYLIQELDKKPITKGLLTDSKSTISIIISNNE- 1698
Query: 1568 LHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFI 1618
R + K +RD V L + +++T AD+ TK L F +
Sbjct: 1699 EKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1749
Score = 98.2 bits (243), Expect = 2e-19
Identities = 122/538 (22%), Positives = 208/538 (37%), Gaps = 70/538 (13%)
Query: 612 LDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTICV---DSSPCIDNVLLV 668
LDSG SR + P V I G + D++ VL
Sbjct: 460 LDSGASRTLIRSAHHIHSASSNPDINVVDAQKRNIPINAIGDLQFHFQDNTKTSIKVLHT 519
Query: 669 DGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKI--------RL 720
+ ++LLS+++LA F + + + DG+VL + + Y + +
Sbjct: 520 PNIAYDLLSLNELAAVDITACFTKN---VLERSDGTVLAPIVKYGDFYWVSKKYLLPSNI 576
Query: 721 SELEAQNVKCLLSVNEEQWVW-HRRLGHASMRKISQLSKLNLVRGLP----NLKFASDAL 775
S NV S + + + HR L HA+ + I K N + + A D
Sbjct: 577 SVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDWSSAIDYQ 636
Query: 776 CEACQKGKFTK---VPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSR 832
C C GK TK + ++ P + LH D+FGPV Y + D+ ++
Sbjct: 637 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 696
Query: 833 WTWVKFL--TRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIA 890
+ WV L R+D VF+T +A ++N+ ++ ++ D G E+ N + GI
Sbjct: 697 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 756
Query: 891 HDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRIS----- 945
++ + +GV ER NRTL + RT LQ +G+ H W + + ++N ++
Sbjct: 757 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 816
Query: 946 -------------VRPILNKTPYELWKNIKPNISYFHPFGCVCYVLNTK----------D 982
+ +L + + PN S HP G Y L+
Sbjct: 817 KSARQHAGLAGLDISTLLPFGQPVIVNDHNPN-SKIHPRGIPGYALHPSRNSYGYIIYLP 875
Query: 983 RLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDDKLDSDQSKLVEKFADL 1042
L K ++ +L G R F N DA T +E ++ S +++ DL
Sbjct: 876 SLKKTVDTTNYVILQGKESRLDQF---NYDALTFDEDLNRLTASYQSFIASNEIQQSDDL 932
Query: 1043 SINVSDKGKAPEEVEPEED--------EPEEEAGPS----NSQTLKKSRITAAHPKEL 1088
+I ++ E+ PE+ P + PS +S+ + K+ I A P+E+
Sbjct: 933 NIESDHDFQSDIELHPEQPRNVLSKAVSPTDSTPPSTHTEDSKRVSKTNIRA--PREV 988
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 115 bits (287), Expect = 1e-24
Identities = 128/535 (23%), Positives = 240/535 (43%), Gaps = 58/535 (10%)
Query: 1121 EPKSIDEALQDKE-WILAMEEELNQFSKNDVWSLV-----KKPESVHVIGTKWVFRNKLN 1174
E + ++ +++KE +I A +E+NQ K W K+ + VI + ++F K +
Sbjct: 809 EAITYNKDIKEKEKYIQAYHKEVNQLLKMKTWDTDRYYDRKEIDPKRVINSMFIFNRKRD 868
Query: 1175 EKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLE-----AIRLLISFSVNHNIVLHQM 1229
+KAR VA+G I + +T+ P + A+ +S ++++N + Q+
Sbjct: 869 GT-----HKARFVARG-----DIQHPDTYDPGMQSNTVHHYALMTSLSLALDNNYYITQL 918
Query: 1230 DVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN 1289
D+ SA+L I EE+Y+ PP D + +LKKSLYGLKQ+ WYE + S+L++
Sbjct: 919 DISSAYLYADIKEELYIRPPPHLGMN---DKLIRLKKSLYGLKQSGANWYETIKSYLIKQ 975
Query: 1290 ---EFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMM- 1345
E VRG + +K+ + + ++VDD+I S + + K+ ++ +++ ++
Sbjct: 976 CGMEEVRG------WSCVFKNSQVTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIIN 1029
Query: 1346 -----GELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTC 1394
E++Y LG+++ + G Y + E + K N+ + P P
Sbjct: 1030 LGESDNEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGL 1088
Query: 1395 I-----LEKEDKSGKVCQKLYRGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTA 1448
LE E+ K+ + +IG Y+ R D+L+ ++ A+ P + L
Sbjct: 1089 YIDQQELELEEDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDM 1148
Query: 1449 VKRILRYLKGTTNLGLMYKKTSEY----KLSGYCDADYAGDRTERKSTSGNCQFLGSNLV 1504
+++++ T + L++ K+ KL DA Y G++ KS GN L ++
Sbjct: 1149 TYELIQFIWNTRDKQLIWHKSKPVKPTNKLVVISDASY-GNQPYYKSQIGNIYLLNGKVI 1207
Query: 1505 SWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAIS-LS 1563
S + S ST EAE + + + + H +++ + D+ + IS +
Sbjct: 1208 GGKSTKASLTCTSTTEAEIHAISESVPLLNNLSHLVQELNKKPITKGLLTDSKSTISIII 1267
Query: 1564 KNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFI 1618
N R + K +RD V L + +++T AD+ TK L F +
Sbjct: 1268 SNNEEKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1322
Score = 98.2 bits (243), Expect = 2e-19
Identities = 122/538 (22%), Positives = 208/538 (37%), Gaps = 70/538 (13%)
Query: 612 LDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTICV---DSSPCIDNVLLV 668
LDSG SR + P V I G + D++ VL
Sbjct: 33 LDSGASRTLIRSAHHIHSASSNPDINVVDAQKRNIPINAIGDLQFHFQDNTKTSIKVLHT 92
Query: 669 DGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKI--------RL 720
+ ++LLS+++LA F + + + DG+VL + + Y + +
Sbjct: 93 PNIAYDLLSLNELAAVDITACFTKN---VLERSDGTVLAPIVKYGDFYWVSKKYLLPSNI 149
Query: 721 SELEAQNVKCLLSVNEEQWVW-HRRLGHASMRKISQLSKLNLVRGLP----NLKFASDAL 775
S NV S + + + HR L HA+ + I K N + + A D
Sbjct: 150 SVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDWSSAIDYQ 209
Query: 776 CEACQKGKFTK---VPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSR 832
C C GK TK + ++ P + LH D+FGPV Y + D+ ++
Sbjct: 210 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 269
Query: 833 WTWVKFL--TRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIA 890
+ WV L R+D VF+T +A ++N+ ++ ++ D G E+ N + GI
Sbjct: 270 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 329
Query: 891 HDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRIS----- 945
++ + +GV ER NRTL + RT LQ +G+ H W + + ++N ++
Sbjct: 330 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 389
Query: 946 -------------VRPILNKTPYELWKNIKPNISYFHPFGCVCYVLNTK----------D 982
+ +L + + PN S HP G Y L+
Sbjct: 390 KSARQHAGLAGLDISTLLPFGQPVIVNDHNPN-SKIHPRGIPGYALHPSRNSYGYIIYLP 448
Query: 983 RLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDDKLDSDQSKLVEKFADL 1042
L K ++ +L G R F N DA T +E ++ S +++ DL
Sbjct: 449 SLKKTVDTTNYVILQGKESRLDQF---NYDALTFDEDLNRLTASYQSFIASNEIQQSDDL 505
Query: 1043 SINVSDKGKAPEEVEPEED--------EPEEEAGPS----NSQTLKKSRITAAHPKEL 1088
+I ++ E+ PE+ P + PS +S+ + K+ I A P+E+
Sbjct: 506 NIESDHDFQSDIELHPEQPRNVLSKAVSPTDSTPPSTHTEDSKRVSKTNIRA--PREV 561
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 115 bits (287), Expect = 1e-24
Identities = 128/533 (24%), Positives = 246/533 (46%), Gaps = 54/533 (10%)
Query: 1121 EPKSIDEALQDKE-WILAMEEELNQFSKNDVWSLV-----KKPESVHVIGTKWVFRNKLN 1174
E + ++ +++KE +I A +E+NQ K W K+ + VI + ++F N
Sbjct: 809 EAITYNKDIKEKEKYIEAYHKEVNQLLKMKTWDTDEYYDRKEIDPKRVINSMFIF----N 864
Query: 1175 EKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSA 1234
+K D +KAR VA+G Q + + A+ +S ++++N + Q+D+ SA
Sbjct: 865 KKRDGT-HKARFVARGDIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSA 923
Query: 1235 FLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EF 1291
+L I EE+Y+ PP D + +LKKSLYGLKQ+ WYE + S+L++ E
Sbjct: 924 YLYADIKEELYIRPPPHLGMN---DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEE 980
Query: 1292 VRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMM------ 1345
VRG + +K+ + + ++VDD+I S + + K+ ++ +++ ++
Sbjct: 981 VRG------WSCVFKNSQVTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESD 1034
Query: 1346 GELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCILEKE 1399
E++Y LG+++ + G Y + E + K N+ + P P ++++
Sbjct: 1035 NEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQD 1093
Query: 1400 D---KSGKVCQKLY--RGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRIL 1453
+ + +K++ + +IG Y+ R D+L+ ++ A+ P L ++
Sbjct: 1094 ELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELI 1153
Query: 1454 RYLKGTTNLGLMYKKTS----EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASK 1509
+++ T + L++ K + KL DA Y G++ KS GN L ++ S
Sbjct: 1154 QFMWDTRDKQLIWHKNKPTEPDNKLVAISDASY-GNQPYYKSQIGNIYLLNGKVIGGKST 1212
Query: 1510 RQSTIALSTAEAEY--ISAAI-CSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNP 1566
+ S ST EAE IS ++ + ++ +L I++ + D+ + IS+ K+
Sbjct: 1213 KASLTCTSTTEAEIHAISESVPLLNNLSYLIQELNKKPIIKG---LLTDSRSTISIIKST 1269
Query: 1567 ILHS-RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFI 1618
R + K +RD V L + +++T AD+ TK L F +
Sbjct: 1270 NEEKFRNRFFGTKAMRLRDEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKLL 1322
Score = 98.2 bits (243), Expect = 2e-19
Identities = 122/538 (22%), Positives = 208/538 (37%), Gaps = 70/538 (13%)
Query: 612 LDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTICV---DSSPCIDNVLLV 668
LDSG SR + P V I G + D++ VL
Sbjct: 33 LDSGASRTLIRSAHHIHSASSNPDINVVDAQKRNIPINAIGDLQFHFQDNTKTSIKVLHT 92
Query: 669 DGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKI--------RL 720
+ ++LLS+++LA F + + + DG+VL + + Y + +
Sbjct: 93 PNIAYDLLSLNELAAVDITACFTKN---VLERSDGTVLAPIVKYGDFYWVSKKYLLPSNI 149
Query: 721 SELEAQNVKCLLSVNEEQWVW-HRRLGHASMRKISQLSKLNLVRGLP----NLKFASDAL 775
S NV S + + + HR L HA+ + I K N + + A D
Sbjct: 150 SVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDRSSAIDYQ 209
Query: 776 CEACQKGKFTK---VPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSR 832
C C GK TK + ++ P + LH D+FGPV Y + D+ ++
Sbjct: 210 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 269
Query: 833 WTWVKFL--TRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIA 890
+ WV L R+D VF+T +A ++N+ ++ ++ D G E+ N + GI
Sbjct: 270 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 329
Query: 891 HDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRIS----- 945
++ + +GV ER NRTL + RT LQ +G+ H W + + ++N ++
Sbjct: 330 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 389
Query: 946 -------------VRPILNKTPYELWKNIKPNISYFHPFGCVCYVLNTK----------D 982
+ +L + + PN S HP G Y L+
Sbjct: 390 KSARQHAGLAGLDISTLLPFGQPVIVNDHNPN-SKIHPRGIPGYALHPSRNSYGYIIYLP 448
Query: 983 RLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDDKLDSDQSKLVEKFADL 1042
L K ++ +L G R F N DA T +E ++ S +++ DL
Sbjct: 449 SLKKTVDTTNYVILQGKESRLDQF---NYDALTFDEDLNRLTASYQSFIASNEIQQSDDL 505
Query: 1043 SINVSDKGKAPEEVEPEED--------EPEEEAGPS----NSQTLKKSRITAAHPKEL 1088
+I ++ E+ PE+ P + PS +S+ + K+ I A P+E+
Sbjct: 506 NIESDHDFQSDIELHPEQPRNVLSKAVSPTDSTPPSTHTEDSKRVSKTNIRA--PREV 561
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 114 bits (285), Expect = 2e-24
Identities = 129/531 (24%), Positives = 237/531 (44%), Gaps = 50/531 (9%)
Query: 1121 EPKSIDEALQDKE-WILAMEEELNQFSKNDVWSLVK-----KPESVHVIGTKWVFRNKLN 1174
E + ++ +++KE +I A +E+NQ K W K + + VI + ++F K +
Sbjct: 809 EAITYNKDIKEKEKYIEAYHKEVNQLLKMKTWDTDKYYDRKEIDPKRVINSMFIFNRKRD 868
Query: 1175 EKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSA 1234
+KAR VA+G Q + + A+ +S ++++N + Q+D+ SA
Sbjct: 869 GT-----HKARFVARGDIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSA 923
Query: 1235 FLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EF 1291
+L I EE+Y+ PP D + +LKKSLYGLKQ+ WYE + S+L++ E
Sbjct: 924 YLYADIKEELYIRPPPHLGMN---DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEE 980
Query: 1292 VRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMG----- 1346
VRG + +++ + + ++VDD++ S N + K + ++ +++ ++
Sbjct: 981 VRG------WSCVFENSQVTICLFVDDMVLFSKNLNSNKRIIDKLKMQYDTKIINLGESD 1034
Query: 1347 -ELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCI---- 1395
E++Y LG+++ + G Y + E + K N+ + P P
Sbjct: 1035 EEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQQ 1093
Query: 1396 -LEKEDKSGKVCQKLYRGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRIL 1453
LE E+ K+ + +IG Y+ R D+L+ ++ A+ P + L ++
Sbjct: 1094 ELELEEDDYKMKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSKQVLDMTYELI 1153
Query: 1454 RYLKGTTNLGLMYKKTSEY----KLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASK 1509
+++ T + L++ K+ KL DA Y G++ KS GN L ++ S
Sbjct: 1154 QFIWNTRDKQLIWHKSKPVKPTNKLVVISDASY-GNQPYYKSQIGNIYLLNGKVIGGKST 1212
Query: 1510 RQSTIALSTAEAEY--ISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPI 1567
+ S ST EAE IS ++ L Q D + + + +T +I +S N
Sbjct: 1213 KASLTCTSTTEAEIHAISESVPLLNNLSYLIQELDKKPITKGLLTDSKSTISIIISNNE- 1271
Query: 1568 LHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFI 1618
R + K +RD V L + +++T AD+ TK L F +
Sbjct: 1272 EKFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1322
Score = 97.1 bits (240), Expect = 4e-19
Identities = 122/538 (22%), Positives = 208/538 (37%), Gaps = 70/538 (13%)
Query: 612 LDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTICV---DSSPCIDNVLLV 668
LDSG SR + P V I G + D++ VL
Sbjct: 33 LDSGASRTLIRSAHHIHSASSNPDINVVDAQKRNIPINAIGDLQFHFQDNTKTSIKVLHT 92
Query: 669 DGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKI--------RL 720
+ ++LLS+++LA F + + + DG+VL + + Y + +
Sbjct: 93 PNIAYDLLSLNELAAVDITACFTKN---VLERSDGTVLAPIVKYGDFYWVSKKYLLPSNI 149
Query: 721 SELEAQNVKCLLSVNEEQWVW-HRRLGHASMRKISQLSKLNLVRGLP----NLKFASDAL 775
S NV S + + + HR L HA+ + I K N + + A D
Sbjct: 150 SVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDWSSAIDYQ 209
Query: 776 CEACQKGKFTK---VPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSR 832
C C GK TK + ++ P + LH D+FGPV Y + D+ ++
Sbjct: 210 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 269
Query: 833 WTWVKFL--TRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIA 890
+ WV L R+D VF+T +A ++N+ ++ ++ D G E+ N + GI
Sbjct: 270 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 329
Query: 891 HDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRIS----- 945
++ + +GV ER NRTL + RT LQ +G+ H W + + ++N ++
Sbjct: 330 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 389
Query: 946 -------------VRPILNKTPYELWKNIKPNISYFHPFGCVCYVLNTK----------D 982
+ +L + + PN S HP G Y L+
Sbjct: 390 KSARQHAGLAGLDISTLLPFGQPVIVNDHNPN-SKIHPRGIPGYALHPSRNSYGYIIYLP 448
Query: 983 RLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDDKLDSDQSKLVEKFADL 1042
L K ++ +L G R F N DA T +E ++ S +++ DL
Sbjct: 449 SLKKTVDTTNYVILQGKESRLDQF---NYDALTFDEDLNRLTASYHSFIASNEIQQSNDL 505
Query: 1043 SINVSDKGKAPEEVEPEE--------DEPEEEAGPS----NSQTLKKSRITAAHPKEL 1088
+I ++ E+ PE+ P + PS +S+ + K+ I A P+E+
Sbjct: 506 NIESDHDFQSDIELHPEQLRNVLSKAVSPTDSTPPSTHTEDSKRVSKTNIRA--PREV 561
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 114 bits (285), Expect = 2e-24
Identities = 123/530 (23%), Positives = 240/530 (45%), Gaps = 48/530 (9%)
Query: 1121 EPKSIDEALQDKE-WILAMEEELNQFSKNDVWSLVK-----KPESVHVIGTKWVFRNKLN 1174
E + ++ +++KE +I A +E+NQ K + W K + + VI + ++F K +
Sbjct: 809 EAITYNKDIKEKEKYIEAYHKEVNQLLKMNTWDTDKYYDRKEIDPKRVINSMFIFNRKRD 868
Query: 1175 EKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSA 1234
+KAR VA+G Q + + A+ +S ++++N + Q+D+ SA
Sbjct: 869 GT-----HKARFVARGDIQHPDTYDSGMQSNTVHHYALMTSLSLALDNNYYITQLDISSA 923
Query: 1235 FLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EF 1291
+L I EE+Y+ PP D + +LKKSLYGLKQ+ WYE + S+L++ E
Sbjct: 924 YLYADIKEELYIRPPPHLGMN---DKLIRLKKSLYGLKQSGANWYETIKSYLIKQCGMEE 980
Query: 1292 VRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMM------ 1345
VRG + +K+ + + ++VDD+I S + + K+ ++ +++ ++
Sbjct: 981 VRG------WSCVFKNSQVTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESD 1034
Query: 1346 GELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCILEKE 1399
E++Y LG+++ + G Y + E + K N+ + P P ++++
Sbjct: 1035 NEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQD 1093
Query: 1400 D---KSGKVCQKLY--RGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRIL 1453
+ + +K++ + +IG Y+ R D+L+ ++ A+ P L ++
Sbjct: 1094 ELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELI 1153
Query: 1454 RYLKGTTNLGLMYKKTS----EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASK 1509
+++ T + L++ K + KL DA Y G++ KS GN L ++ S
Sbjct: 1154 QFMWDTRDKQLIWHKNKPTEPDNKLVAISDASY-GNQPYYKSQIGNIYLLNGKVIGGKST 1212
Query: 1510 RQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAIS-LSKNPIL 1568
+ S ST EAE + + + + H +++ + D+ + IS + N
Sbjct: 1213 KASLTCTSTTEAEIHAISESVPLLNNLSHLVQELNKKPITKGLLTDSKSTISIIISNNEE 1272
Query: 1569 HSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFI 1618
R + K +RD V L + +++T AD+ TK L F +
Sbjct: 1273 KFRNRFFGTKAMRLRDEVSGNHLHVCYIETKKNIADVMTKPLPIKTFKLL 1322
Score = 98.2 bits (243), Expect = 2e-19
Identities = 122/538 (22%), Positives = 208/538 (37%), Gaps = 70/538 (13%)
Query: 612 LDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTICV---DSSPCIDNVLLV 668
LDSG SR + P V I G + D++ VL
Sbjct: 33 LDSGASRTLIRSAHHIHSASSNPDINVVDAQKRNIPINAIGDLQFHFQDNTKTSIKVLHT 92
Query: 669 DGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKI--------RL 720
+ ++LLS+++LA F + + + DG+VL + + Y + +
Sbjct: 93 PNIAYDLLSLNELAAVDITACFTKN---VLERSDGTVLAPIVKYGDFYWVSKKYLLPSNI 149
Query: 721 SELEAQNVKCLLSVNEEQWVW-HRRLGHASMRKISQLSKLNLVRGLP----NLKFASDAL 775
S NV S + + + HR L HA+ + I K N + + A D
Sbjct: 150 SVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDWSSAIDYQ 209
Query: 776 CEACQKGKFTK---VPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSR 832
C C GK TK + ++ P + LH D+FGPV Y + D+ ++
Sbjct: 210 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 269
Query: 833 WTWVKFL--TRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIA 890
+ WV L R+D VF+T +A ++N+ ++ ++ D G E+ N + GI
Sbjct: 270 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 329
Query: 891 HDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRIS----- 945
++ + +GV ER NRTL + RT LQ +G+ H W + + ++N ++
Sbjct: 330 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 389
Query: 946 -------------VRPILNKTPYELWKNIKPNISYFHPFGCVCYVLNTK----------D 982
+ +L + + PN S HP G Y L+
Sbjct: 390 KSARQHAGLAGLDISTLLPFGQPVIVNDHNPN-SKIHPRGIPGYALHPSRNSYGYIIYLP 448
Query: 983 RLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDDKLDSDQSKLVEKFADL 1042
L K ++ +L G R F N DA T +E ++ S +++ DL
Sbjct: 449 SLKKTVDTTNYVILQGKESRLDQF---NYDALTFDEDLNRLTASYQSFIASNEIQQSDDL 505
Query: 1043 SINVSDKGKAPEEVEPEED--------EPEEEAGPS----NSQTLKKSRITAAHPKEL 1088
+I ++ E+ PE+ P + PS +S+ + K+ I A P+E+
Sbjct: 506 NIESDHDFQSDIELHPEQPRNVLSKAVSPTDSTPPSTHTEDSKRVSKTNIRA--PREV 561
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 113 bits (282), Expect = 5e-24
Identities = 128/533 (24%), Positives = 245/533 (45%), Gaps = 54/533 (10%)
Query: 1121 EPKSIDEALQDKE-WILAMEEELNQFSKNDVWSLV-----KKPESVHVIGTKWVFRNKLN 1174
E + ++ +++KE +I A +E+NQ K W K+ + VI + ++F N
Sbjct: 1236 EAITYNKDIKEKEKYIEAYHKEVNQLLKMKTWDTDEYYDRKEIDPKRVINSMFIF----N 1291
Query: 1175 EKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNHNIVLHQMDVKSA 1234
+K D +KAR VA+G Q T + A+ +S ++++N + Q+D+ SA
Sbjct: 1292 KKRDGT-HKARFVARGDIQHPDTYDTGMQSNTVHHYALMTSLSLALDNNYYITQLDISSA 1350
Query: 1235 FLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERLSSFLLEN---EF 1291
+L I EE+Y+ PP D + +LKKS YGLKQ+ WYE + S+L++ E
Sbjct: 1351 YLYADIKEELYIRPPPHLGMN---DKLIRLKKSHYGLKQSGANWYETIKSYLIKQCGMEE 1407
Query: 1292 VRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEMSMM------ 1345
VRG + +K+ + + ++VDD+I S + + K+ ++ +++ ++
Sbjct: 1408 VRG------WSCVFKNSQVTICLFVDDMILFSKDLNANKKIITTLKKQYDTKIINLGESD 1461
Query: 1346 GELKY-FLGIQVDQTPEGTY--IHQSKYTKELLKKFNM---LESTVAKTPMHPTCILEKE 1399
E++Y LG+++ + G Y + E + K N+ + P P ++++
Sbjct: 1462 NEIQYDILGLEI-KYQRGKYMKLGMENSLTEKIPKLNVPLNPKGRKLSAPGQPGLYIDQD 1520
Query: 1400 D---KSGKVCQKLY--RGMIGSLLYLTAS-RPDILFSVHLCARFQSDPRETHLTAVKRIL 1453
+ + +K++ + +IG Y+ R D+L+ ++ A+ P L ++
Sbjct: 1521 ELEIDEDEYKEKVHEMQKLIGLASYVGYKFRFDLLYYINTLAQHILFPSRQVLDMTYELI 1580
Query: 1454 RYLKGTTNLGLMYKKTS----EYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWASK 1509
+++ T + L++ K + KL DA Y G++ KS GN L ++ S
Sbjct: 1581 QFMWDTRDKQLIWHKNKPTEPDNKLVAISDASY-GNQPYYKSQIGNIFLLNGKVIGGKST 1639
Query: 1510 RQSTIALSTAEAEY--ISAAI-CSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNP 1566
+ S ST EAE IS ++ + ++ +L I++ + D+ + IS+ K+
Sbjct: 1640 KASLTCTSTTEAEIHAISESVPLLNNLSYLIQELNKKPIIKG---LLTDSRSTISIIKST 1696
Query: 1567 ILHS-RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFI 1618
R + K +RD V L + +++T AD+ TK L F +
Sbjct: 1697 NEEKFRNRFFGTKAMRLRDEVSGNNLYVYYIETKKNIADVMTKPLPIKTFKLL 1749
Score = 97.8 bits (242), Expect = 2e-19
Identities = 122/538 (22%), Positives = 208/538 (37%), Gaps = 70/538 (13%)
Query: 612 LDSGCSRHMTGEKRMFRELKLKPGGEVGFGGNEKGKIIGTGTICV---DSSPCIDNVLLV 668
LDSG SR + P V I G + D++ VL
Sbjct: 460 LDSGASRTLIRSAHHIHSASSNPDINVVDAQKRNIPINAIGDLQFHFQDNTKTSIKVLHT 519
Query: 669 DGLTHNLLSISQLADKGYDVIFNQKSCRAVSQIDGSVLFNSKRKNNIYKI--------RL 720
+ ++LLS+++LA F + + + DG+VL + + Y + +
Sbjct: 520 PNIAYDLLSLNELAAVDITACFTKN---VLERSDGTVLAPIVQYGDFYWVSKRYLLPSNI 576
Query: 721 SELEAQNVKCLLSVNEEQWVW-HRRLGHASMRKISQLSKLNLVRGLP----NLKFASDAL 775
S NV S + + + HR L HA+ + I K N + + A D
Sbjct: 577 SVPTINNVHTSESTRKYPYPFIHRMLAHANAQTIRYSLKNNTITYFNESDVDWSSAIDYQ 636
Query: 776 CEACQKGKFTK---VPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSR 832
C C GK TK + ++ P + LH D+FGPV Y + D+ ++
Sbjct: 637 CPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSAPSYFISFTDETTK 696
Query: 833 WTWVKFL--TRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIA 890
+ WV L R+D VF+T +A ++N+ ++ ++ D G E+ N + GI
Sbjct: 697 FRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTNRTLHKFLEKNGIT 756
Query: 891 HDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRIS----- 945
++ + +GV ER NRTL + RT LQ +G+ H W + + ++N ++
Sbjct: 757 PCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSK 816
Query: 946 -------------VRPILNKTPYELWKNIKPNISYFHPFGCVCYVLNTK----------D 982
+ +L + + PN S HP G Y L+
Sbjct: 817 KSARQHAGLAGLDISTLLPFGQPVIVNDHNPN-SKIHPRGIPGYALHPSRNSYGYIIYLP 875
Query: 983 RLHKFDAKSSKCLLLGYSDRSKGFRFYNTDAKTIEESIHVRFDDKLDSDQSKLVEKFADL 1042
L K ++ +L G R F N DA T +E ++ S +++ DL
Sbjct: 876 SLKKTVDTTNYVILQGKESRLDQF---NYDALTFDEDLNRLTASYQSFIASNEIQESNDL 932
Query: 1043 SINVSDKGKAPEEVEPEED--------EPEEEAGPS----NSQTLKKSRITAAHPKEL 1088
+I ++ E+ PE+ P + PS +S+ + K+ I A P+E+
Sbjct: 933 NIESDHDFQSDIELHPEQPRNVLSKAVSPTDSTPPSTHTEDSKRVSKTNIRA--PREV 988
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 110 bits (276), Expect = 2e-23
Identities = 143/614 (23%), Positives = 267/614 (43%), Gaps = 94/614 (15%)
Query: 1048 DKGKAPEEVEPEEDEPEEEAG-----PSNSQTLKKSRITAAHPKELILGNKDEPVRTRSA 1102
DK + E E D+ + P N +T+ +I A + E I N D ++ +
Sbjct: 1231 DKNNSLTSYELERDKKRSKKNRVKLIPDNMETVSAPKIRAIYYNEAISKNPD--LKEKHE 1288
Query: 1103 FRPSEETLLSLKGLVSLIEPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHV 1162
++ + K L +L + K D ++ +S++++ P+++ +
Sbjct: 1289 YKQAYH-----KELQNLKDMKVFDVDVK--------------YSRSEI------PDNL-I 1322
Query: 1163 IGTKWVFRNKLNEKGDVVRNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLISFSVNH 1222
+ T +F K N KAR+V +G +Q Y+ I++ + + N
Sbjct: 1323 VPTNTIFTKKRNGI-----YKARIVCRGDTQSPDT-YSVITTESLNHNHIKIFLMIANNR 1376
Query: 1223 NIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKKSLYGLKQAPRAWYERL 1282
N+ + +D+ AFL + EE+Y+ P D + V KL K+LYGLKQ+P+ W + L
Sbjct: 1377 NMFMKTLDINHAFLYAKLEEEIYIPHP---HDRRC---VVKLNKALYGLKQSPKEWNDHL 1430
Query: 1283 SSFL-----LENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQSLCKEFSEMMQ 1337
+L +N + G T +D L++ +YVDD + ++N+ EF ++
Sbjct: 1431 RQYLNGIGLKDNSYTPGLYQT-------EDKNLMIAVYVDDCVIAASNEQRLDEFINKLK 1483
Query: 1338 AEFEMSMMGEL------KYFLGIQ---------VDQTPEGTYIHQ--SKYTKEL--LKKF 1378
+ FE+ + G L LG+ +D T + ++I++ KY +EL ++K
Sbjct: 1484 SNFELKITGTLIDDVLDTDILGMDLVYNKRLGTIDLTLK-SFINRMDKKYNEELKKIRKS 1542
Query: 1379 NMLESTVAKTPMHPTCILEKEDKSGKVCQKLYRGMIGSLLYLT-ASRPDILFSVHLCARF 1437
++ + K + E++ + KL + ++G L Y+ R DI F+V AR
Sbjct: 1543 SIPHMSTYKIDPKKDVLQMSEEEFRQGVLKLQQ-LLGELNYVRHKCRYDIEFAVKKVARL 1601
Query: 1438 QSDPRETHLTAVKRILRYLKGTTNLGLMYKK--TSEYKLSGYCDADYAGDRTERKSTSGN 1495
+ P E + +I++YL ++G+ Y + + K+ DA G + +S G
Sbjct: 1602 VNYPHERVFYMIYKIIQYLVRYKDIGIHYDRDCNKDKKVIAITDAS-VGSEYDAQSRIGV 1660
Query: 1496 CQFLGSNLVSWASKRQSTIALSTAEAE----YISAAICSTQMLWMKHQLEDYQILESNIP 1551
+ G N+ + S + + +S+ EAE Y A T + +K E ++I
Sbjct: 1661 ILWYGMNIFNVYSNKSTNRCVSSTEAELHAIYEGYADSETLKVTLKELGEGD---NNDIV 1717
Query: 1552 IYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYV-QKGVLLLKFVDTDHQWADIFTKSL 1610
+ D+ AI + K +K I++ + +K + LLK + AD+ TK +
Sbjct: 1718 MITDSKPAIQGLNRSYQQPKEKFTWIKTEIIKEKIKEKSIKLLKITGKGN-IADLLTKPV 1776
Query: 1611 AED---RFNFILKN 1621
+ RF +LKN
Sbjct: 1777 SASDFKRFIQVLKN 1790
Score = 87.4 bits (215), Expect = 3e-16
Identities = 72/276 (26%), Positives = 122/276 (44%), Gaps = 11/276 (3%)
Query: 742 HRRLGHASMRKISQLSKLN-LVRGLPNLKFASDALCEACQKGKFTK---VPFKAKNVVST 797
H+R+GH +++I K N L +K ++ C+ C+ K TK N +
Sbjct: 562 HKRMGHTGIQQIENSIKHNHYEESLDLIKEPNEFWCQTCKISKATKRNHYTGSMNNHSTD 621
Query: 798 SRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRW--TWVKFLTRKDESHAVFSTFIAQ 855
P +D+FGPV + + KRY +++VD+ +R+ T F + A I
Sbjct: 622 HEPGSSWCMDIFGPVSSSNADTKRYMLIMVDNNTRYCMTSTHFNKNAETILAQVRKNIQY 681
Query: 856 VQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKNRTLQEM 915
V+ + ++ + SD G EF ND+ E F S GI H + + NG ER RT+
Sbjct: 682 VETQFDRKVREINSDRGTEFTNDQIEEYFISKGIHHILTSTQDHAANGRAERYIRTIITD 741
Query: 916 ARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKN--IKPNISYFHPFGC 973
A T+L+++ + FW V +A I+N + + K P + + + F PFG
Sbjct: 742 ATTLLRQSNLRVKFWEYAVTSATNIRNYLEHKS-TGKLPLKAISRQPVTVRLMSFLPFGE 800
Query: 974 VCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFY 1009
+ N + K ++L S G++F+
Sbjct: 801 KGIIWNHNHK--KLKPSGLPSIILCKDPNSYGYKFF 834
>M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820
(ORF170)
Length = 170
Score = 98.6 bits (244), Expect = 1e-19
Identities = 46/97 (47%), Positives = 67/97 (68%)
Query: 1121 EPKSIDEALQDKEWILAMEEELNQFSKNDVWSLVKKPESVHVIGTKWVFRNKLNEKGDVV 1180
EPKS+ AL+D W AM+EEL+ S+N W LV P + +++G KWVF+ KL+ G +
Sbjct: 27 EPKSVIFALKDPGWCQAMQEELDALSRNKTWILVPPPVNQNILGCKWVFKTKLHSDGTLD 86
Query: 1181 RNKARLVAQGYSQQEGIDYTETFAPVARLEAIRLLIS 1217
R KARLVA+G+ Q+EGI + ET++PV R IR +++
Sbjct: 87 RLKARLVAKGFHQEEGIYFVETYSPVVRTATIRTILN 123
>M240_ARATH (P93290) Hypothetical mitochondrial protein AtMg00240
(ORF111a)
Length = 111
Score = 69.3 bits (168), Expect = 8e-11
Identities = 31/82 (37%), Positives = 50/82 (60%)
Query: 1418 LYLTASRPDILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTSEYKLSGY 1477
+YLT +RPD+ F+V+ ++F S R + AV ++L Y+KGT GL Y TS+ +L +
Sbjct: 1 MYLTITRPDLTFAVNRLSQFSSASRTAQMQAVYKVLHYVKGTVGQGLFYSATSDLQLKAF 60
Query: 1478 CDADYAGDRTERKSTSGNCQFL 1499
D+D+A R+S +G C +
Sbjct: 61 ADSDWASCPDTRRSVTGFCSLV 82
>POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1189
Score = 60.5 bits (145), Expect = 4e-08
Identities = 70/276 (25%), Positives = 115/276 (41%), Gaps = 19/276 (6%)
Query: 691 NQKSCRAVSQIDGSVLFNSKRKNNIYKIRLSELEAQNVKCLLSVNEEQWVWHRRLGHASM 750
+Q+ RA+ + N +++ KI L + EA L++ ++ W LG+ +
Sbjct: 806 DQEEARAIGATENKDTRNWEKEG---KIVLPQKEA------LAMIQQMHAW-THLGNRKL 855
Query: 751 RKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKVPFKAKNVVSTSRPLELLHIDLFG 810
+ + + + + R ++ + A C+ CQ+ A +RP ID F
Sbjct: 856 KLLIEKTDFLIPRASTLIEQVTSA-CKVCQQVNAGATRVPAGKRTRGNRPGVYWEID-FT 913
Query: 811 PVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSD 870
VK G K Y +V VD +S W F TR++ +H V + ++ V + SD
Sbjct: 914 EVKPHYAGYK-YLLVFVDTFSGWVEA-FPTRQETAHIVAKKILEEIFPRFGLPKV-IGSD 970
Query: 871 LGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFW 930
G F + + L GI C PQ +G VER NRT++E + ETG+ W
Sbjct: 971 NGPAFVSQVSQGLARILGINWKLHCAYRPQSSGQVERMNRTIKETLTKLTLETGLKD--W 1028
Query: 931 AETVNTACYIQNRISVRPILNKTPYELWKNIKPNIS 966
++ A R TPYE+ P +S
Sbjct: 1029 RRLLSLALLRARNTPNR--FGLTPYEILYGGPPPLS 1062
>POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1196
Score = 56.6 bits (135), Expect = 5e-07
Identities = 63/231 (27%), Positives = 97/231 (41%), Gaps = 14/231 (6%)
Query: 744 RLGHASMRKISQL-----SKLNLVRGLPNLKFASDALCEACQKGKFTKVPFKAKNVVSTS 798
RL H +K+ L S ++ L++ +D+ C C + +K A V
Sbjct: 849 RLTHLGYQKMKALLDRGESPYYMLNRDKTLQYVADS-CTVCAQVNASKAKIGAGVRVRGH 907
Query: 799 RPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDESHAVFSTFIAQVQN 858
RP ID F VK + G +Y +V VD +S W F T+++ + V + ++
Sbjct: 908 RPGSHWEID-FTEVKP-GLYGYKYLLVFVDTFSGWVEA-FPTKRETARVVSKKLLEEIFP 964
Query: 859 EKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKNRTLQEMART 918
V + SD G F + +S+ D GI C PQ +G VER NRT++E
Sbjct: 965 RFGMPQV-LGSDNGPAFTSQVSQSVADLLGIDWKLHCAYRPQSSGQVERMNRTIKETLTK 1023
Query: 919 MLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKPNISYFH 969
+ G W + A Y + R + P TPYE+ P + FH
Sbjct: 1024 LTLAAGTRD--WVLLLPLALY-RARNTPGP-HGLTPYEILYGAPPPLVNFH 1070
>M710_ARATH (P92512) Hypothetical mitochondrial protein AtMg00710
(ORF120)
Length = 120
Score = 56.2 bits (134), Expect = 7e-07
Identities = 29/69 (42%), Positives = 37/69 (53%)
Query: 909 NRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKPNISYF 968
NRT+ E R+ML E G+ K F A+ NTA +I N+ I P E+W P SY
Sbjct: 2 NRTIIEKVRSMLCECGLPKTFRADAANTAVHIINKYPSTAINFHVPDEVWFQSVPTYSYL 61
Query: 969 HPFGCVCYV 977
FGCV Y+
Sbjct: 62 RRFGCVAYI 70
>POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 1046
Score = 55.5 bits (132), Expect = 1e-06
Identities = 56/193 (29%), Positives = 83/193 (42%), Gaps = 12/193 (6%)
Query: 776 CEACQK--GKFTKVPFKAKNVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRW 833
C+ CQ+ T+VP + +RP ID F VK G K Y +V VD +S W
Sbjct: 737 CKVCQQVNAGATRVPEGKRT--RGNRPGVYWEID-FTEVKPHYAGYK-YLLVFVDTFSGW 792
Query: 834 TWVKFLTRKDESHAVFSTFIAQVQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDF 893
+ TR++ +H V + ++ V + SD G F + + L + GI
Sbjct: 793 VEA-YPTRQETAHMVAKKILEEIFPRFGLPKV-IGSDNGPAFVSQVSQGLARTLGINWKL 850
Query: 894 SCPRTPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKT 953
C PQ +G VER NRT++E + ETG+ W ++ A R T
Sbjct: 851 HCAYRPQSSGQVERMNRTIKETLTKLTLETGLKD--WRRLLSLALLRARNTPNR--FGLT 906
Query: 954 PYELWKNIKPNIS 966
PYE+ P +S
Sbjct: 907 PYEILYGGPPPLS 919
>POL_MLVMO (P03355) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1199
Score = 53.1 bits (126), Expect = 6e-06
Identities = 63/233 (27%), Positives = 97/233 (41%), Gaps = 20/233 (8%)
Query: 744 RLGHASMRKISQLSK--------LNLVRGLPNLKFASDALCEACQKGKFTKVPFKAKNVV 795
+L H S K+ L + LN R L N+ C+AC + +K K V
Sbjct: 849 QLTHLSFSKMKALLERSHSPYYMLNRDRTLKNIT----ETCKACAQVNASKSAVKQGTRV 904
Query: 796 STSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDESHAVFSTFIAQ 855
RP ID F +K + G +Y +V +D +S W F T+K+ + V + +
Sbjct: 905 RGHRPGTHWEID-FTEIKP-GLYGYKYLLVFIDTFSGWIEA-FPTKKETAKVVTKKLLEE 961
Query: 856 VQNEKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKNRTLQEM 915
+ V + +D G F + +++ D GI C PQ +G VER NRT++E
Sbjct: 962 IFPRFGMPQV-LGTDNGPAFVSKVSQTVADLLGIDWKLHCAYRPQSSGQVERMNRTIKET 1020
Query: 916 ARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKPNISYF 968
+ TG W + A Y + R + P TPYE+ P + F
Sbjct: 1021 LTKLTLATG--SRDWVLLLPLALY-RARNTPGP-HGLTPYEILYGAPPPLVNF 1069
>POL_MLVAK (P03357) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 843
Score = 53.1 bits (126), Expect = 6e-06
Identities = 63/231 (27%), Positives = 97/231 (41%), Gaps = 15/231 (6%)
Query: 744 RLGHASMRKISQL-----SKLNLVRGLPNLKFASDALCEACQKGKFTKVPFKAKNVVSTS 798
RL H +K+ L S ++ L++ +D+ C C + +K A V
Sbjct: 497 RLTHLGYQKMKALLDRGESPYYMLNRDKTLQYVADS-CTVCAQVNASKAKIGAGVRVRGH 555
Query: 799 RPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDESHAVFSTFIAQVQN 858
RP ID F VK + G +Y +V VD +S W F T+++ + V + ++
Sbjct: 556 RPGSHWEID-FTEVKP-GLYGYKYLLVFVDTFSGWVEA-FPTKRETARVVSKKLLEEIFP 612
Query: 859 EKACRIVRVRSDLGGEFENDKFESLFDSYGIAHDFSCPRTPQQNGVVERKNRTLQEMART 918
V + SD G F + +S+ D GI C PQ +G VER NRT++E
Sbjct: 613 RFGMPQV-LGSDNGPAFTSQVSQSVADLLGI-DKLHCAYRPQSSGQVERMNRTIKETLTK 670
Query: 919 MLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNIKPNISYFH 969
+ G W + A Y + R + P TPYE+ P + FH
Sbjct: 671 LTLAAGTRD--WVLLLPLALY-RARNTPGP-HGLTPYEILYGAPPPLVNFH 717
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.332 0.142 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 170,532,879
Number of Sequences: 164201
Number of extensions: 6836399
Number of successful extensions: 22644
Number of sequences better than 10.0: 103
Number of HSP's better than 10.0 without gapping: 54
Number of HSP's successfully gapped in prelim test: 50
Number of HSP's that attempted gapping in prelim test: 22406
Number of HSP's gapped (non-prelim): 181
length of query: 1626
length of database: 59,974,054
effective HSP length: 124
effective length of query: 1502
effective length of database: 39,613,130
effective search space: 59498921260
effective search space used: 59498921260
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 73 (32.7 bits)
Lotus: description of TM0243.12


