

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0239.11
(307 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
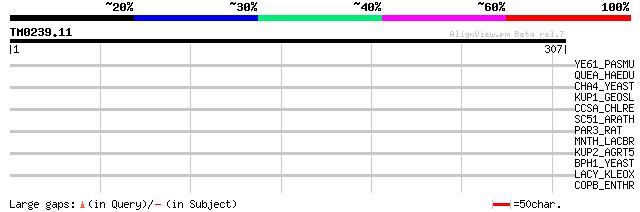
Score E
Sequences producing significant alignments: (bits) Value
YE61_PASMU (Q9CKY9) Hypothetical UPF0324 membrane protein PM1461 33 0.68
QUEA_HAEDU (Q7VLY1) S-adenosylmethionine:tRNA ribosyltransferase... 31 3.4
CHA4_YEAST (P43634) CHA4 activatory protein 31 3.4
KUP1_GEOSL (Q74AK4) Probable potassium transport system protein ... 30 5.8
CCSA_CHLRE (P48269) Cytochrome c biogenesis protein ccsA 30 5.8
SC51_ARATH (Q39208) Delta-7-sterol-C5(6)-desaturase 1 (EC 1.3.3.... 30 7.5
PAR3_RAT (Q920E1) Proteinase activated receptor 3 precursor (PAR... 30 7.5
MNTH_LACBR (Q93V04) Manganese transport protein mntH (Hop-induci... 30 7.5
KUP2_AGRT5 (Q8UJL0) Probable potassium transport system protein ... 30 7.5
BPH1_YEAST (P25356) Beige protein homolog 1 30 7.5
LACY_KLEOX (P18817) Lactose permease (Lactose-proton symport) 30 9.8
COPB_ENTHR (P05425) Probable copper exporting ATPase B (EC 3.6.3.4) 30 9.8
>YE61_PASMU (Q9CKY9) Hypothetical UPF0324 membrane protein PM1461
Length = 336
Score = 33.5 bits (75), Expect = 0.68
Identities = 32/133 (24%), Positives = 57/133 (42%), Gaps = 15/133 (11%)
Query: 16 GGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLF 75
GG +P + ++ +++ F LL S K Q T + P F+FLF
Sbjct: 205 GGNISPAVADTAVISKMIRVMMLAPFLLLVSWLLTKQNDQAKTTNISIPW-----FAFLF 259
Query: 76 TLALQINPCIL---YVLVSIVSIATLIN-------GLMGKITLLNDSSTSPVLQPSLYIA 125
L IN L +++ IV I +L+ GL I+ + + P++ +L +
Sbjct: 260 ILMAVINSFSLIPAHIVAWIVEIDSLLLIAAMTALGLTTHISAIKQAGIKPLILGALVLC 319
Query: 126 WILLCAFQFCVGL 138
W+++ F VG+
Sbjct: 320 WLVIGGFFVNVGI 332
>QUEA_HAEDU (Q7VLY1) S-adenosylmethionine:tRNA
ribosyltransferase-isomerase (EC 5.-.-.-) (Queuosine
biosynthesis protein queA)
Length = 361
Score = 31.2 bits (69), Expect = 3.4
Identities = 20/73 (27%), Positives = 36/73 (48%), Gaps = 1/73 (1%)
Query: 11 FFSNHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQP 70
FFS+ P TF ++ + + + LP S ++ +SA Y T + H+ QQ+ +
Sbjct: 283 FFSDTNIFLYPGKTFKIVDVLITNFHLPESTLIML-VSAFAGYRHTMNAYQHAVQQKYRF 341
Query: 71 FSFLFTLALQINP 83
FS+ + + NP
Sbjct: 342 FSYGDAMLINKNP 354
>CHA4_YEAST (P43634) CHA4 activatory protein
Length = 648
Score = 31.2 bits (69), Expect = 3.4
Identities = 15/42 (35%), Positives = 24/42 (56%)
Query: 148 IFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGLHETTQVWY 189
IF+ ++SS + SSS + A ++LL F+ +G E WY
Sbjct: 291 IFQLEDSSLAESSSSSKLAIIQTLLCLAFYDIGSGENPMAWY 332
>KUP1_GEOSL (Q74AK4) Probable potassium transport system protein
kup1
Length = 631
Score = 30.4 bits (67), Expect = 5.8
Identities = 55/235 (23%), Positives = 93/235 (39%), Gaps = 32/235 (13%)
Query: 35 LLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQINPCILYVLVSIVS 94
L + L++F + +Y L L P + +PF L P LY LV + +
Sbjct: 249 LPIRLAWFCCVLPALLLNYFGQGALLLSDPSEATEPFYHLAP------PWALYPLVLLAT 302
Query: 95 IATLING---LMGKITLLNDS---STSP----VLQPSLYIAWILLCAFQFCVGLGIEASI 144
+AT+I + G +L + SP V S I I + A + + L ++
Sbjct: 303 LATIIASQAVISGVFSLTRQAIQLRLSPRMRIVQTSSEEIGQIYIPAVNWALMLAT-ITL 361
Query: 145 AAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGLHETTQVWYRVVVRPVVD-----D 199
AG SSS ++++G + A +++ + + E W+ + V + D
Sbjct: 362 VAGF----GSSSGLAAAYGVAVATTMVITALLVRFVMLERWH-WHPLAVAGLTVVFLTVD 416
Query: 200 TVFGGAKKEK-----WIERVAVAASLGVLWWWRLREEVESLVVMAEAKKSEQFGE 249
F GA K WI A A V+ WR E+ + ++A+A F E
Sbjct: 417 LAFFGANILKVGAGGWIPLAAGLAVFTVMITWRRGRELVTTHLLAQATPLPSFLE 471
>CCSA_CHLRE (P48269) Cytochrome c biogenesis protein ccsA
Length = 353
Score = 30.4 bits (67), Expect = 5.8
Identities = 29/85 (34%), Positives = 43/85 (50%), Gaps = 7/85 (8%)
Query: 26 HLLTITLLSLLLPLSF-FLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQINPC 84
+LL T +LL+ L+A S + TF+L P++ QQ + ALQ N
Sbjct: 156 YLLYCTSFTLLVEKMLGSLIAPCSLLMNAFATFSL----PKEMQQASPLV--PALQSNWL 209
Query: 85 ILYVLVSIVSIATLINGLMGKITLL 109
+++V V I+S ATLI G + I L
Sbjct: 210 MMHVTVMIISYATLIIGSLLSILFL 234
>SC51_ARATH (Q39208) Delta-7-sterol-C5(6)-desaturase 1 (EC 1.3.3.-)
(Delta-7-C-5 sterol desaturase 1)
(Delta7-sterol-C5-desaturase 1) (STEROL1 protein) (Dwarf
7 protein)
Length = 281
Score = 30.0 bits (66), Expect = 7.5
Identities = 19/68 (27%), Positives = 32/68 (46%), Gaps = 7/68 (10%)
Query: 213 RVAVAASLGVLWWWRLREEV-ESLVVMAEAKKSEQFGELELKDFIGWWLYYVTVTIGMVR 271
R+ + ++ + W+ L V ES++ K GE GW LY+V + I +V
Sbjct: 85 RLQMFVAMKAMPWYTLLPTVSESMIERGWTKCFASIGEF------GWILYFVYIAIYLVF 138
Query: 272 IVKGLMWM 279
+ G+ WM
Sbjct: 139 VEFGIYWM 146
>PAR3_RAT (Q920E1) Proteinase activated receptor 3 precursor (PAR-3)
(Thrombin receptor-like 2) (Coagulation factor II
receptor-like 2)
Length = 368
Score = 30.0 bits (66), Expect = 7.5
Identities = 23/85 (27%), Positives = 39/85 (45%), Gaps = 3/85 (3%)
Query: 21 PKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQ---TFTLFLHSPQQQQQPFSFLFTL 77
PK F LL ++ +++ L LA L H +Q T +H + PF F + +
Sbjct: 203 PKRNFTLLMCGVVWVMVVLYMLPLAILKQEYHLVQPGITTCHDVHDTCESPLPFQFYYFV 262
Query: 78 ALQINPCILYVLVSIVSIATLINGL 102
+L ++ +VS+ TLI+ L
Sbjct: 263 SLAFFGFLIPFVVSVFCYTTLIHKL 287
>MNTH_LACBR (Q93V04) Manganese transport protein mntH (Hop-inducible
cation transporter)
Length = 464
Score = 30.0 bits (66), Expect = 7.5
Identities = 21/77 (27%), Positives = 38/77 (49%), Gaps = 4/77 (5%)
Query: 30 ITLLSLLLPLSF----FLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQINPCI 85
+ +L LL+ L F ++A+L A + T+ +FL PQ Q ++ T + N +
Sbjct: 170 VLILLLLMKLGFRKIEAIVATLVAVILLVFTYEVFLAGPQLDQMFAGYMPTKDIVTNKSM 229
Query: 86 LYVLVSIVSIATLINGL 102
LY+ + IV + + L
Sbjct: 230 LYLALGIVGATVMPHDL 246
>KUP2_AGRT5 (Q8UJL0) Probable potassium transport system protein
kup2
Length = 637
Score = 30.0 bits (66), Expect = 7.5
Identities = 42/219 (19%), Positives = 85/219 (38%), Gaps = 33/219 (15%)
Query: 37 LPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQINPCILYVLVSIVSIA 96
+ +++F+L + A +YL L L +P + PF L+ IL + ++++
Sbjct: 251 ISVAWFILVFPALALNYLGQGALVLSNPAAIENPFYLLYPEWALFPMIILATMATVIASQ 310
Query: 97 TLINGL---------MGKITLLNDSSTSPVLQPSLYIAWILLCAFQFCVGLGIEASIAAG 147
+I G +G + L TS +Y+ + + F +
Sbjct: 311 AVITGAFSLARQAVHLGFLPKLMIRFTSETNTGQIYVPAVNMVLF---------IGVLVL 361
Query: 148 IFEFDNSSSSSSSSFGESAAERSLLSKVF---FLLGLHETTQVWYRVVVRPVVD-DTVFG 203
IF F S S ++++G S +++ + FL + + +++ P+ + VF
Sbjct: 362 IFSF-GDSESLATAYGISVTGAMVVTTLMAFQFLRSIRGKSAFTAAILLAPLFSIEAVFL 420
Query: 204 GAKKEK-----WIERVAVAASLGVLWWWR-----LREEV 232
A K W+ + V+W W LRE++
Sbjct: 421 AANLLKVHDGGWVPLALAGVIILVMWTWTKGSRYLREKI 459
>BPH1_YEAST (P25356) Beige protein homolog 1
Length = 2167
Score = 30.0 bits (66), Expect = 7.5
Identities = 13/44 (29%), Positives = 25/44 (56%)
Query: 184 TTQVWYRVVVRPVVDDTVFGGAKKEKWIERVAVAASLGVLWWWR 227
TT+VW +V ++P+ + T+F +K + ++ V S +WR
Sbjct: 1848 TTEVWRKVPMKPIFEKTIFNLNEKNRSVDYVIHDPSYFDSLYWR 1891
>LACY_KLEOX (P18817) Lactose permease (Lactose-proton symport)
Length = 416
Score = 29.6 bits (65), Expect = 9.8
Identities = 30/145 (20%), Positives = 65/145 (44%), Gaps = 17/145 (11%)
Query: 51 KHYLQTFTLFLHSPQQQQQPFSFLFTLALQINPCILYVLVSIVS-----IATLINGLMGK 105
+ + F F SPQ+ + F F+ T +N I++ +I++ A LI GL+
Sbjct: 246 QQFANFFKGFFSSPQRGTEVFGFVTTGGELLNALIMFCAPAIINRIGAKNALLIAGLIMS 305
Query: 106 ITLLNDSSTSPVLQPSLY-------IAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSS 158
+ +L S + ++ + I ++L+ F++ + + ++A +F + S
Sbjct: 306 VRILGSSFATSAVEVIILKMLHMFEIPFLLVGTFKY-ISSAFKGKLSATLFLIGFNLSKQ 364
Query: 159 SSSFGESAAERSLLSKVFFLLGLHE 183
SS SA + +++ +G H+
Sbjct: 365 LSSVVLSA----WVGRMYDTVGFHQ 385
>COPB_ENTHR (P05425) Probable copper exporting ATPase B (EC 3.6.3.4)
Length = 745
Score = 29.6 bits (65), Expect = 9.8
Identities = 26/95 (27%), Positives = 45/95 (47%), Gaps = 16/95 (16%)
Query: 193 VRPVVDDTVFGGAKKEKWIERVAVAAS--LGVLWWWRLREEVESLVVMAEAKKSEQFGEL 250
V+ V D+V GG+ + V + G L +V +V A+ +KS+ L
Sbjct: 300 VKKQVGDSVIGGSINGDGTIEITVTGTGENGYL------AKVMEMVRKAQGEKSK----L 349
Query: 251 E-LKDFIGWWLYYVTVTIGMVRIVKGLMWMFMVAL 284
E L D + WL+YV + +G++ + W+F+ L
Sbjct: 350 EFLSDKVAKWLFYVALVVGIIAFI---AWLFLANL 381
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,376,414
Number of Sequences: 164201
Number of extensions: 1241263
Number of successful extensions: 4695
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 10
Number of HSP's that attempted gapping in prelim test: 4693
Number of HSP's gapped (non-prelim): 12
length of query: 307
length of database: 59,974,054
effective HSP length: 110
effective length of query: 197
effective length of database: 41,911,944
effective search space: 8256652968
effective search space used: 8256652968
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0239.11


