

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0234.12
(607 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
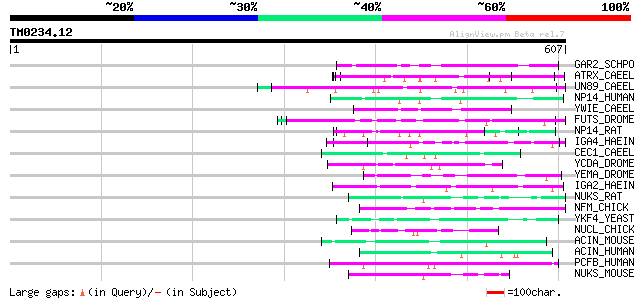
Score E
Sequences producing significant alignments: (bits) Value
GAR2_SCHPO (P41891) Protein gar2 60 2e-08
ATRX_CAEEL (Q9U7E0) Transcriptional regulator ATRX homolog (X-li... 59 5e-08
UN89_CAEEL (O01761) Muscle M-line assembly protein unc-89 (Uncoo... 55 5e-07
NP14_HUMAN (Q14978) Nucleolar phosphoprotein p130 (Nucleolar 130... 53 2e-06
YWIE_CAEEL (Q23525) Hypothetical protein ZK546.14 in chromosome II 52 3e-06
FUTS_DROME (Q9W596) Microtubule-associated protein futsch 52 3e-06
NP14_RAT (P41777) Nucleolar phosphoprotein p130 (Nucleolar 130 k... 51 8e-06
IGA4_HAEIN (P45386) Immunoglobulin A1 protease precursor (EC 3.4... 51 8e-06
CEC1_CAEEL (P34618) Chromo domain protein cec-1 51 8e-06
YCDA_DROME (Q9VGW1) Putative cadherin CG4509 precursor 50 2e-05
YEMA_DROME (P25992) Yemanuclein-alpha 49 3e-05
IGA2_HAEIN (P45384) Immunoglobulin A1 protease precursor (EC 3.4... 49 4e-05
NUKS_RAT (Q9EPJ0) Nuclear ubiquitous casein and cyclin-dependent... 49 5e-05
NFM_CHICK (P16053) Neurofilament triplet M protein (160 kDa neur... 48 6e-05
YKF4_YEAST (P35732) Hypothetical 84.0 kDa protein in NUP120-CSE4... 47 1e-04
NUCL_CHICK (P15771) Nucleolin (Protein C23) 46 3e-04
ACIN_MOUSE (Q9JIX8) Apoptotic chromatin condensation inducer in ... 46 3e-04
ACIN_HUMAN (Q9UKV3) Apoptotic chromatin condensation inducer in ... 46 3e-04
PCFB_HUMAN (O94913) Pre-mRNA cleavage complex II protein Pcf11 (... 45 4e-04
NUKS_MOUSE (Q80XU3) Nuclear ubiquitous casein and cyclin-depende... 45 4e-04
>GAR2_SCHPO (P41891) Protein gar2
Length = 500
Score = 59.7 bits (143), Expect = 2e-08
Identities = 54/243 (22%), Positives = 106/243 (43%), Gaps = 36/243 (14%)
Query: 358 PKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQM 417
PK + ++++RA+S +PS K + K++ +KKK S + E S +S +
Sbjct: 51 PKKSKKEAKRASSPEPSK---KSVKKQKKSKKKEESSSESESESSSSESESSSSESESSS 107
Query: 418 SDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAE 477
S+SESS +SSS E+E+E +I++ EE ++ E +S +E
Sbjct: 108 SESESSS---SESSSSESEEE---------VIVKTEEKKESSSE----------SSSSSE 145
Query: 478 ESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRT 537
+ +A+ ++ +K + ++ ++ +SE + E ++ + S +
Sbjct: 146 SEEEEEAVVKIEEKKESSSDSSSESSSSESESESSSSESEEEEEVVEKTEEKKEGSSESS 205
Query: 538 SPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKS 597
S S S +ES +SDS+ S S S+ + + + +P ++ P
Sbjct: 206 SDSESSSDSSSESGDSDSS-----------SDSESESSSEDEKKRKAEPASEERPAKITK 254
Query: 598 PIQ 600
P Q
Sbjct: 255 PSQ 257
Score = 43.1 bits (100), Expect = 0.002
Identities = 41/194 (21%), Positives = 86/194 (44%), Gaps = 33/194 (17%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
ES + ++ +S ++S S E + ++ + ++ E E + +S+ +
Sbjct: 86 ESESESSSSESESSSSESESSSSESESSSSESSSSESEEEVIVKTEEKKESSSESSSSSE 145
Query: 415 QQMSDS---------ESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQN 465
+ + ESS D+ +SSS E+E E E E+E++ E +
Sbjct: 146 SEEEEEAVVKIEEKKESSSDSSSESSSSESESESSS---------SESEEEEEVVEKTEE 196
Query: 466 VIQGIRE-ASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKR 524
+G E +SD+E S+DS + +++++ SE +SE +++R+++
Sbjct: 197 KKEGSSESSSDSESSSDSSS-------------ESGDSDSSSDSESESSSEDEKKRKAEP 243
Query: 525 TSDPARAASLTRTS 538
S+ R A +T+ S
Sbjct: 244 ASE-ERPAKITKPS 256
>ATRX_CAEEL (Q9U7E0) Transcriptional regulator ATRX homolog
(X-linked nuclear protein-1)
Length = 1359
Score = 58.5 bits (140), Expect = 5e-08
Identities = 55/272 (20%), Positives = 122/272 (44%), Gaps = 36/272 (13%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
ESP K + + +RA S SD + EE KK + K V ++ + + TS+ +
Sbjct: 73 ESPRKSSKKSRKRAKSESESDESDE----EEDRKKSKSKKKVDQKKKEKSKKKRTTSSSE 128
Query: 415 QQMSDSESSEDTWKDS----------SSEETEDEDMPLAKRR------KIILEEEEDEQD 458
+ SD E + + K S SSEE+E+E ++ K E E+ +
Sbjct: 129 DEDSDEEREQKSKKKSKKTKKQTSSESSEESEEERKVKKSKKNKEKSVKKRAETSEESDE 188
Query: 459 DHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQR 518
D + + +G+++ + +E ++S+ VK+ K + + +KE+ +E + + ++
Sbjct: 189 DEKPSKKSKKGLKKKAKSESESESEDEKEVKKSKKKSK-KVVKKESESEDEAPEKKKTEK 247
Query: 519 ERRSKRTSDPARAASLT--------------RTSPIRSLSKVITESINSDSALVVIPEQP 564
+RSK +S+ + + + + P+ ++ K+ ++ + +S + V+P++
Sbjct: 248 RKRSKTSSEESSESEKSDEEEEEKESSPKPKKKKPL-AVKKLSSDEESEESDVEVLPQKK 306
Query: 565 MPISTSLPTSTQTEPIQPETQPQTQAEPQAPK 596
+ +L + ++ E Q + E + K
Sbjct: 307 KRGAVTLISDSEDEKDQKSESEASDVEEKVSK 338
Score = 54.7 bits (130), Expect = 7e-07
Identities = 56/259 (21%), Positives = 107/259 (40%), Gaps = 19/259 (7%)
Query: 363 RKSRRATSSKPS-DPKGKGILKEEPAKKKAASKTVIIREPSPERPLK-----RTSAQQQQ 416
RK +RA K + +GK K+ PAKK+ AS + + E P K R A+ +
Sbjct: 31 RKEKRAQKLKEKREREGKPPPKKRPAKKRKASSSEEDDDDEEESPRKSSKKSRKRAKSES 90
Query: 417 MSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQ---NVIQGIREA 473
SD E+ K S S++ D+ ++K EDE D E Q + ++
Sbjct: 91 ESDESDEEEDRKKSKSKKKVDQKKKEKSKKKRTTSSSEDEDSDEEREQKSKKKSKKTKKQ 150
Query: 474 SDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAAS 533
+ +E S +S+ VK+ K ++++ +E++ E ++ ++ + + + S
Sbjct: 151 TSSESSEESEEERKVKKSKKNKEKSVKKRAETSEESDEDEKPSKKSKKGLKKKAKSESES 210
Query: 534 LT------RTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQ 587
+ + S +S V ES + D A PE+ ++ E + E +
Sbjct: 211 ESEDEKEVKKSKKKSKKVVKKESESEDEA----PEKKKTEKRKRSKTSSEESSESEKSDE 266
Query: 588 TQAEPQAPKSPIQTATTAI 606
+ E ++ P + A+
Sbjct: 267 EEEEKESSPKPKKKKPLAV 285
Score = 52.0 bits (123), Expect = 4e-06
Identities = 46/212 (21%), Positives = 95/212 (44%), Gaps = 24/212 (11%)
Query: 357 PPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQ 416
PPK K R+A+SS+ D + ++ K + +K+ + S E ++ S +++
Sbjct: 50 PPKKRPAKKRKASSSEEDDDDEEESPRKSSKKSRKRAKSESESDESDEEEDRKKSKSKKK 109
Query: 417 MSDSESSEDTWKDSSSEETEDEDMPLAKRRKI--------------ILEEEEDEQDDHEI 462
+ D + E + K ++ +EDED + +K EE E+E+ +
Sbjct: 110 V-DQKKKEKSKKKRTTSSSEDEDSDEEREQKSKKKSKKTKKQTSSESSEESEEERKVKKS 168
Query: 463 IQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRS 522
+N + +++ ++ E +D D P K +K G ++ ++ +E SE EV++ ++
Sbjct: 169 KKNKEKSVKKRAETSEESDEDEKPSKKSKK----GLKKKAKSESESESEDEKEVKKSKKK 224
Query: 523 -----KRTSDPARAASLTRTSPIRSLSKVITE 549
K+ S+ A + + R SK +E
Sbjct: 225 SKKVVKKESESEDEAPEKKKTEKRKRSKTSSE 256
Score = 46.6 bits (109), Expect = 2e-04
Identities = 40/171 (23%), Positives = 73/171 (42%), Gaps = 23/171 (13%)
Query: 354 PESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQ 413
P K +K ++ S S+ + K + K + KK K + +PE+ K+T +
Sbjct: 192 PSKKSKKGLKKKAKSESESESEDE-KEVKKSKKKSKKVVKKESESEDEAPEK--KKTEKR 248
Query: 414 QQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREA 473
++ + SE S ++ K S EE E E P K++K + +++
Sbjct: 249 KRSKTSSEESSESEK-SDEEEEEKESSPKPKKKKPL-------------------AVKKL 288
Query: 474 SDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKR 524
S EES +SD + +++K + E +Q SE + E+ SK+
Sbjct: 289 SSDEESEESDVEVLPQKKKRGAVTLISDSEDEKDQKSESEASDVEEKVSKK 339
>UN89_CAEEL (O01761) Muscle M-line assembly protein unc-89
(Uncoordinated protein 89)
Length = 6632
Score = 55.1 bits (131), Expect = 5e-07
Identities = 71/344 (20%), Positives = 144/344 (41%), Gaps = 31/344 (9%)
Query: 287 LKKMQIIEKIQVAPKEDTSDEVRQRDYPIDDF--PIWSKK------DNPACILEYVRMLR 338
+KK + EK+ PK T + ++ P+ +K + PA + +
Sbjct: 1526 VKKEKTPEKVDEKPKSPTKKDKSPEKSITEEIKSPVKKEKSPEKVEEKPASPTKKEKSPE 1585
Query: 339 RQGDPITLTEFVNTLP----ESPPKLATR--KSRRATSSKPSDPKGKGILKEEPAKKKAA 392
+ P +E P +SP K KS + S + +D K K K+E + +K+A
Sbjct: 1586 KPASPTKKSENEVKSPTKKEKSPEKSVVEELKSPKEKSPEKADDKPKSPTKKEKSPEKSA 1645
Query: 393 SKTV---IIREPSPER-PLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKI 448
++ V +E SPE+ K TS +++ S ++ ++D K T+ E P K
Sbjct: 1646 TEDVKSPTKKEKSPEKVEEKPTSPTKKESSPTKKTDDEVKSP----TKKEKSPQTVEEKP 1701
Query: 449 ILEEEEDEQDDHEIIQNVIQGIREASD-AEESTDSDAIPVVKRRKLPLRGPLQQKE--TA 505
++++ + +++ V ++ + AEE S K+ K P + ++ + T
Sbjct: 1702 ASPTKKEKSPEKSVVEEVKSPKEKSPEKAEEKPKSPT----KKEKSPEKSAAEEVKSPTK 1757
Query: 506 AEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLS--KVITESINSDSALVVIPEQ 563
E++ E ++E + + +K+ S P + A SP + + + E S + PE+
Sbjct: 1758 KEKSPEKSAEEKPKSPTKKESSPVKMADDEVKSPTKKEKSPEKVEEKPASPTKKEKTPEK 1817
Query: 564 PMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQTATTAIP 607
PT + P P + +++ ++P+ P + + P
Sbjct: 1818 SAAEELKSPTKKEKSPSSPTKKTGDESKEKSPEKPEEKPKSPTP 1861
Score = 51.2 bits (121), Expect = 8e-06
Identities = 68/364 (18%), Positives = 138/364 (37%), Gaps = 42/364 (11%)
Query: 272 EDLAMTIGDALNGIKLKKMQIIEKIQVAPKEDTSDEVRQRDYPIDDFPIWSKKDNPACIL 331
E IG G+ + + KI+V K D+ +VR+ +++++ P ++
Sbjct: 1311 ESATTVIGGGSGGVTEGSISV-SKIEVVSKTDSQTDVREGTPKRR--VSFAEEELPKEVI 1367
Query: 332 EYVRMLRRQGDPITLTEFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKA 391
+ R ++ P + E P + S K P ++ PA ++
Sbjct: 1368 DSDRKKKKSPSPDKKEKSPEKTEEKPASPTKKTGEEVKSPKEKSPASPTKKEKSPAAEEV 1427
Query: 392 ASKTVIIREP--------SPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLA 443
S T + P SP P K+T + ++ S +S T K+ S E+ ED P+
Sbjct: 1428 KSPTKKEKSPSSPTKKEKSPSSPTKKTGDEVKEKSPPKS--PTKKEKSPEKPEDVKSPVK 1485
Query: 444 KRRK----IILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIP---------VVKR 490
K + I+E + + + E+ ++ S + P K+
Sbjct: 1486 KEKSPDATNIVEVSSETTIEKTETTMTTEMTHESEESRTSVKKEKTPEKVDEKPKSPTKK 1545
Query: 491 RKLP-------LRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPI--- 540
K P ++ P++++++ + +P S ++E+ ++ + P + + SP
Sbjct: 1546 DKSPEKSITEEIKSPVKKEKSPEKVEEKPASPTKKEKSPEKPASPTKKSENEVKSPTKKE 1605
Query: 541 RSLSKVITESINSDSALVVIPEQPMPISTS----LPTSTQTEPIQPETQPQTQAE--PQA 594
+S K + E + S P S + P + TE ++ T+ + E +
Sbjct: 1606 KSPEKSVVEELKSPKEKSPEKADDKPKSPTKKEKSPEKSATEDVKSPTKKEKSPEKVEEK 1665
Query: 595 PKSP 598
P SP
Sbjct: 1666 PTSP 1669
Score = 40.4 bits (93), Expect = 0.014
Identities = 51/261 (19%), Positives = 94/261 (35%), Gaps = 45/261 (17%)
Query: 348 EFVNTLPESPPKLA---TRKSRRATSSKPSDPKGKGILKEEPA----KKKAASKTVI--I 398
E V P SP K T+K+ S K ++E+PA K+K+ K+V+ +
Sbjct: 1660 EKVEEKPTSPTKKESSPTKKTDDEVKSPTKKEKSPQTVEEKPASPTKKEKSPEKSVVEEV 1719
Query: 399 REPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQD 458
+ P + P K + +S E + + T+ E P + + E
Sbjct: 1720 KSPKEKSPEKAEEKPKSPTKKEKSPEKSAAEEVKSPTKKEKSPEKSAEEKPKSPTKKESS 1779
Query: 459 DHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPT-SEVQ 517
++ + ++ + + E + K+ K P +++AAE+ PT E
Sbjct: 1780 PVKMADDEVKSPTKKEKSPEKVEEKPASPTKKEKTP-------EKSAAEELKSPTKKEKS 1832
Query: 518 RERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQT 577
+K+T D ++ S PE+P S PT ++
Sbjct: 1833 PSSPTKKTGDESKEKS---------------------------PEKPEEKPKS-PTPKKS 1864
Query: 578 EPIQPETQPQTQAEPQAPKSP 598
P P+ + E + P +P
Sbjct: 1865 PPGSPKKKKSKSPEAEKPPAP 1885
>NP14_HUMAN (Q14978) Nucleolar phosphoprotein p130 (Nucleolar 130
kDa protein) (140 kDa nucleolar phosphoprotein)
(Nopp140) (Nucleolar and coiled-body phosphoprotein 1)
Length = 699
Score = 53.1 bits (126), Expect = 2e-06
Identities = 65/282 (23%), Positives = 103/282 (36%), Gaps = 58/282 (20%)
Query: 352 TLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTS 411
+L PPK A K + SS+ S + +EE KK +K V+ + + P K+ +
Sbjct: 303 SLGTQPPKKAVEKQQPVESSEDSSDESDSSSEEE---KKPPTKAVVSKATTKPPPAKKAA 359
Query: 412 AQQQQMSDSESSED---------TWKDSSSEETEDEDMPLAKRR----------KIILEE 452
SDS+SSED T K+SS++ P K + +L
Sbjct: 360 ESSSDSSDSDSSEDDEAPSKPAGTTKNSSNKPAVTTKSPAVKPAAAPKQPVGGGQKLLTR 419
Query: 453 EEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRR-------KLPLRGPLQQKETA 505
+ D E E+S +EE + K + LP + Q +
Sbjct: 420 KADSSSSEE----------ESSSSEEEKTKKMVATTKPKATAKAALSLPAKQAPQGSRDS 469
Query: 506 AEQASEPTSEVQRERRSKRT--SDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQ 563
+ + +SE + E+ SK P + A S S K ES NS S
Sbjct: 470 SSDSDSSSSEEEEEKTSKSAVKKKPQKVAGGAAPSKPASAKKGKAESSNSSS-------- 521
Query: 564 PMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQTATTA 605
+ +E + + + + PQAPK+ +A TA
Sbjct: 522 ---------SDDSSEEEEEKLKGKGSPRPQAPKANGTSALTA 554
Score = 42.0 bits (97), Expect = 0.005
Identities = 55/281 (19%), Positives = 111/281 (38%), Gaps = 31/281 (11%)
Query: 343 PITLTEFVNTLPE---SPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIR 399
P+T+ P+ + PK+A K+ ++SS S EE KTV +
Sbjct: 189 PVTVKAQTKAPPKPARAAPKIANGKAASSSSSSSSSSSSDD--SEEEKAAATPKKTVPKK 246
Query: 400 EPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIIL--------- 450
+ + P+K + ++ S SE DSSS+E E++ P+ +
Sbjct: 247 QVVAKAPVKAATTPTRKSSSSE-------DSSSDEEEEQKKPMKNKPGPYSYAPPPSAPP 299
Query: 451 -EEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQA 509
++ Q + ++ Q + + D+ + +DS + K + K A++A
Sbjct: 300 PKKSLGTQPPKKAVEKQ-QPVESSEDSSDESDSSSEEEKKPPTKAVVSKATTKPPPAKKA 358
Query: 510 SEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPM---- 565
+E +S+ S+ P++ A T+ S + + ++ +A P+QP+
Sbjct: 359 AESSSDSSDSDSSEDDEAPSKPAGTTKNSSNKPAVTTKSPAVKPAAA----PKQPVGGGQ 414
Query: 566 PISTSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQTATTAI 606
+ T S+ +E ++ + + A P TA A+
Sbjct: 415 KLLTRKADSSSSEEESSSSEEEKTKKMVATTKPKATAKAAL 455
>YWIE_CAEEL (Q23525) Hypothetical protein ZK546.14 in chromosome II
Length = 472
Score = 52.4 bits (124), Expect = 3e-06
Identities = 43/173 (24%), Positives = 77/173 (43%), Gaps = 14/173 (8%)
Query: 377 KGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETE 436
K K K +P K A K + + E+ +K+ +Q + +DS+S ED S EE E
Sbjct: 89 KSKQHSKVQPQKVVAPVKRPADQNKNKEKVVKKDQKKQDKKADSDSEED--DSSDDEEKE 146
Query: 437 DEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLR 496
+ D P+AK++K EE D+ +D E + +G A +AE+S +D
Sbjct: 147 ETDEPVAKKQK--KEESSDDDEDSEDGEEP-EGNNGAVEAEDSDSTDE---------EEE 194
Query: 497 GPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITE 549
P + +T A+ + ++ +E + + + + R + L K + E
Sbjct: 195 TPSKPNKTVAQSTLKSNGKIDKEIQKLEDDEDNESPEIRRQIALLRLQKKLKE 247
>FUTS_DROME (Q9W596) Microtubule-associated protein futsch
Length = 5412
Score = 52.4 bits (124), Expect = 3e-06
Identities = 73/326 (22%), Positives = 129/326 (39%), Gaps = 51/326 (15%)
Query: 294 EKIQVAPKEDT--SDEVRQRDYPIDDFPIWSKK-DNPACILEYVRMLRRQGDPITLTEFV 350
EK +A K++ S E +R+ + FP+ SK+ PA + E V+ +
Sbjct: 3294 EKSPLASKDEAEKSKEESRRESVAEQFPLVSKEVSRPASVAESVK------------DEA 3341
Query: 351 NTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASK--TVIIREPSPERPLK 408
E P ++ SR A+ + G +K+E K K S+ +V + P P +
Sbjct: 3342 EKSKEESPLMSKEASRPASVA--------GSVKDEAEKSKEESRRESVAEKSPLPSKEAS 3393
Query: 409 RTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQ 468
R ++ + + D D K+ S E+ E PLA + E E I++ +
Sbjct: 3394 RPASVAESVKD---EADKSKEESRRESGAEKSPLASK------EASRPASVAESIKDEAE 3444
Query: 469 GIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRER-------R 521
+E S E + +P + + P E+ ++A + E +R+
Sbjct: 3445 KSKEESRRESVAEKSPLPSKEASR-----PTSVAESVKDEAEKSKEESRRDSVAEKSPLA 3499
Query: 522 SKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQ 581
SK S PA A + +S + ES+ S L E P S + + E +
Sbjct: 3500 SKEASRPASVAESVQDEAEKSKEESRRESVAEKSPL-ASKEASRPASVAESIKDEAEKSK 3558
Query: 582 PETQPQTQAEPQAPKSPIQTATTAIP 607
E++ ++ AE KSP+ + + P
Sbjct: 3559 EESRRESVAE----KSPLASKEASRP 3580
Score = 51.6 bits (122), Expect = 6e-06
Identities = 66/304 (21%), Positives = 120/304 (38%), Gaps = 16/304 (5%)
Query: 298 VAPKEDTSDEVRQRDYPIDDFPIWSKK-DNPACILEYVRMLRRQGDPITLTEFVNTLPES 356
+ + + S E +R+ + P+ SK+ PA + E ++ + T E V
Sbjct: 2181 IKDEAEKSKEESRRESVAEKSPLPSKEASRPASVAESIKDEAEKSKEETRRESVAEKSPL 2240
Query: 357 PPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQ 416
P K A+R + A S K K KEE ++ AA K+ P P + R ++ +
Sbjct: 2241 PSKEASRPASVAESIKDEAEKS----KEESRRESAAEKS-----PLPSKEASRPASVAES 2291
Query: 417 MSD--SESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREAS 474
+ D +S E++ ++S +E + + + K + L+E + E +++ +E S
Sbjct: 2292 VKDEADKSKEESRRESMAESGKAQSI---KGDQSPLKEVSRPESVAESVKDDPVKSKEPS 2348
Query: 475 DAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASL 534
E S + + PL + + + + +E Q RR +T +
Sbjct: 2349 RRESVAGSVTADSARDDQSPLESKGASRPESVVDSVKDEAEKQESRRESKTESVIPPKAK 2408
Query: 535 TRTSPIRSLSKV-ITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQ 593
SP L V +TE+I D+ + P Q S+ S + + E P E
Sbjct: 2409 DDKSPKEVLQPVSMTETIREDADQPMKPSQAESRRESIAESIKASSPRDEKSPLASKEAS 2468
Query: 594 APKS 597
P S
Sbjct: 2469 RPGS 2472
Score = 49.7 bits (117), Expect = 2e-05
Identities = 64/310 (20%), Positives = 130/310 (41%), Gaps = 30/310 (9%)
Query: 303 DTSDEVRQRDYPIDDFPIWSKK-DNPACILEYVRMLRRQGDPITLTEFVNTLPESPPKLA 361
+ S E +R+ + P+ SK+ PA + E V+ + + E V P K A
Sbjct: 3666 EKSKEESRRESVAEKSPLASKEASRPASVAESVKDEAEKSKEESRRESVAEKSPLPSKEA 3725
Query: 362 TRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSE 421
+R + A S K K KEE ++ A K+ + + ++ + S + ++E
Sbjct: 3726 SRPTSVAESVKDEAEKS----KEESRRESVAEKSSL----ASKKASRPASVAESVKDEAE 3777
Query: 422 SSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTD 481
S K+ S E+ E PLA + E E +++ + +E S E +
Sbjct: 3778 KS----KEESRRESVAEKSPLASK------EASRPASVAESVKDEAEKSKEESRRESVAE 3827
Query: 482 SDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIR 541
+P + + P E+ ++A + E +RE ++++ P + +R + +
Sbjct: 3828 KSPLPSKEASR-----PTSVAESVKDEADKSKEESRRESGAEKS--PLASMEASRPTSVA 3880
Query: 542 SLSKVITESINSDSALVVIPEQ-PMPISTSLPTSTQTEPIQPE---TQPQTQAEPQAPKS 597
K TE +S + E+ P+P + ++ E ++ E ++ +++ E A KS
Sbjct: 3881 ESVKDETEKSKEESRRESVTEKSPLPSKEASRPTSVAESVKDEAEKSKEESRRESVAEKS 3940
Query: 598 PIQTATTAIP 607
P+ + ++ P
Sbjct: 3941 PLASKESSRP 3950
Score = 44.3 bits (103), Expect = 0.001
Identities = 57/264 (21%), Positives = 114/264 (42%), Gaps = 24/264 (9%)
Query: 359 KLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMS 418
K +TR+ A S K S P K + E A +++ +EPS +K +AQ ++ S
Sbjct: 1846 KGSTRRESVAESDKSSQPF-KETSRPESAVGSMKDESMS-KEPSRRESVKDGAAQSRETS 1903
Query: 419 DSESSEDTWKDSSSE---------ETEDEDMPLAKRRKIILEEEEDEQDDH--EIIQNVI 467
S ++ KD + + T+ ++ K K L EE + E +++
Sbjct: 1904 RPASVAESAKDGADDLKELSRPESTTQSKEAGSIKDEKSPLASEEASRPASVAESVKDEA 1963
Query: 468 QGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSD 527
+ +E S E + +P + + P E+ ++A + E +RE ++++
Sbjct: 1964 EKSKEESRRESVAEKSPLPSKEASR-----PASVAESIKDEAEKSKEESRRESVAEKSPL 2018
Query: 528 PARAASLTRTSPIRSLSKVITESINSDSALVVIPEQ-PMPISTSLPTSTQTEPIQPE--- 583
P++ AS R + + K E +S + E+ P+P + ++ E I+ E
Sbjct: 2019 PSKEAS--RPASVAESIKDEAEKSKEESRRESVAEKSPLPSKEASRPASVAESIKDEAEK 2076
Query: 584 TQPQTQAEPQAPKSPIQTATTAIP 607
++ +++ E A KSP+ + + P
Sbjct: 2077 SKEESRRESVAEKSPLPSKEASRP 2100
Score = 43.5 bits (101), Expect = 0.002
Identities = 53/260 (20%), Positives = 100/260 (38%), Gaps = 21/260 (8%)
Query: 353 LPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSA 412
+P + T KS A+ G +KE+ + K+ + P P RP
Sbjct: 1416 VPAPEEAIKTEKSPLASKETSRPESATGSVKEDTEQTKSK------KSPVPSRPESEAKD 1469
Query: 413 QQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQ--DDHEIIQNVIQGI 470
++ + E+S +S +E +DE RR+ I + +DE D + ++ + +
Sbjct: 1470 KKSPFASGEASRP---ESVAESVKDEAGKAESRRESIAKTHKDESSLDKAKEQESRRESL 1526
Query: 471 REASDAEESTDSDAIPVVKRRKLP------LRGPLQQKETAAEQASEPTSEVQRERRSKR 524
E+ E D + K P + P +++ A +E T + + SK
Sbjct: 1527 AESIKPESGIDEKSALASKEASRPESVTDKSKEPSRRESIAESLKAESTKDEKSAPPSKE 1586
Query: 525 TSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPET 584
S P + +S ESI ++SA I + + S+ + E +PE+
Sbjct: 1587 ASRPGSVVESVKDETEKSKEPSRRESI-AESAKPPIEFREVSRPESVIDGIKDESAKPES 1645
Query: 585 Q---PQTQAEPQAPKSPIQT 601
+ P E P+S +++
Sbjct: 1646 RRDSPLASKEASRPESVLES 1665
Score = 39.3 bits (90), Expect = 0.030
Identities = 53/261 (20%), Positives = 107/261 (40%), Gaps = 33/261 (12%)
Query: 359 KLATRKSRRAT-SSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQM 417
K T+K ++ +S+PS +LK++ K+++ +++ + T + M
Sbjct: 3022 KADTKKDGKSQEASRPSSVDE--LLKDDDEKQESRRQSI-----TGSHKAMSTMGDESPM 3074
Query: 418 SDSESSEDTWKDSS------SEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIR 471
++ S++ + S E T+DE+ PL RR + E I ++ +G +
Sbjct: 3075 DKADKSKEPSRPESVAESIKHENTKDEESPLGSRRDSVAE---------SIKSDITKGEK 3125
Query: 472 EASDAEE-STDSDAIPVVKRRKLPLRGPLQQKETAAEQAS-EPTSEVQRERRSKRTSDPA 529
++E S + +K K R +E+ AE E + + SK S P
Sbjct: 3126 SPLPSKEVSRPESVVGSIKDEKAESR-----RESVAESVKPESSKDATSAPPSKEHSRPE 3180
Query: 530 RAASLTRTSPIRSLSK--VITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQ 587
+ ++ S+ + +SI + +L+V E P S + P Q ++P+
Sbjct: 3181 SVLGSLKDEGDKTTSRRVSVADSIKDEKSLLVSQEASRPESEAESLKDAAAPSQETSRPE 3240
Query: 588 TQAEP-QAPKSPIQTATTAIP 607
+ E + KSP+ + + P
Sbjct: 3241 SVTESVKDGKSPVASKEASRP 3261
Score = 36.2 bits (82), Expect = 0.26
Identities = 48/239 (20%), Positives = 95/239 (39%), Gaps = 26/239 (10%)
Query: 359 KLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMS 418
K ++SRR + ++ P+ GI ++ K AS+ + + S E ++++ ++
Sbjct: 1515 KAKEQESRRESLAESIKPES-GIDEKSALASKEASRPESVTDKSKE------PSRRESIA 1567
Query: 419 DSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEE 478
+S +E T + S+ +++ A R ++E +DE + +E S E
Sbjct: 1568 ESLKAESTKDEKSAPPSKE-----ASRPGSVVESVKDETEKS----------KEPSRRES 1612
Query: 479 STDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTS 538
+S P+ R + P + +++++P S SK S P +
Sbjct: 1613 IAESAKPPIEFRE---VSRPESVIDGIKDESAKPESRRDSPLASKEASRPESVLESVKDE 1669
Query: 539 PIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKS 597
PI+S K ES+ ++ P+ TS S ++ AE P+S
Sbjct: 1670 PIKSTEKSRRESVAESFKADSTKDEKSPL-TSKDISRPESAVENVMDAVGSAERSQPES 1727
Score = 35.0 bits (79), Expect = 0.57
Identities = 45/183 (24%), Positives = 72/183 (38%), Gaps = 25/183 (13%)
Query: 404 ERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPL-AKRRKIILEEEEDEQDDHEI 462
E + + Q+ DS+ SE K S+ EE E + A RK LE QD+ E+
Sbjct: 961 ESSMTKEEEIQKHQRDSQESEKKRKKSAEEEIEAAIAKVEAAERKARLEGASARQDESEL 1020
Query: 463 IQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRS 522
E ++ + I + R Q + A E+ S PT E E+ S
Sbjct: 1021 DV-------EPEQSKIKAEVQDIIATAKDIAKSRTEEQLAKPAEEELSSPTPE---EKLS 1070
Query: 523 KRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQP 582
K+TSD T+ I + V+ ++ +PE+ S ++ + T P P
Sbjct: 1071 KKTSD-------TKDDQIGAPVDVLPVNLQES-----LPEE--KFSATIESGATTAPTLP 1116
Query: 583 ETQ 585
E +
Sbjct: 1117 EDE 1119
Score = 35.0 bits (79), Expect = 0.57
Identities = 64/305 (20%), Positives = 118/305 (37%), Gaps = 46/305 (15%)
Query: 332 EYVRMLRRQGDPITLTEFVNTLPESPPKLATRKSRRATSSKPSDPKGK--GILKEE---- 385
+YV+ ++ + I + V TLPE+ P AT P D + G L +
Sbjct: 1135 KYVKEETKEAEAIVVAT-VQTLPEAAPLAIDTILASATKDAPKDANAEALGELPDSGERV 1193
Query: 386 -PAKKKAASKTVIIRE--PSPER----PLKRTSAQQQQMSDSE----SSEDTWKDSSSEE 434
P K ++ ++R+ +P+ P+ + DS+ S ++S+ EE
Sbjct: 1194 LPMKMTFEAQQNLLRDVIKTPDEVADLPVHEEADLGLYEKDSQDAGAKSISHKEESAKEE 1253
Query: 435 TEDEDMPLAKRRKIILEEEEDEQD-DHEIIQNVIQGIRE---------ASDAEESTDSDA 484
E +D K +I L +E ++ D H +++ +Q + E EE ++
Sbjct: 1254 KETDDEKENKVGEIELGDEPNKVDISHVLLKESVQEVAEKVVVIETTVEKKQEEIVEATT 1313
Query: 485 IPVVKRRKLPLRGPLQQKE----------TAAEQASEPTSEVQRERRSKRTSD---PARA 531
+ + + + L ++ KE ++A + S + + S TSD PA+
Sbjct: 1314 V-ITQENQEDLMEQVKDKEEHEQKIESGIITEKEAKKSASTPEEKETSDITSDDELPAQL 1372
Query: 532 ASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAE 591
A T P + + T SI S + E+ + + Q PE +T+
Sbjct: 1373 ADPTTVPPKSAKDREDTGSIESPPTI----EEAIEVEVQAKQEAQKPVPAPEEAIKTEKS 1428
Query: 592 PQAPK 596
P A K
Sbjct: 1429 PLASK 1433
Score = 35.0 bits (79), Expect = 0.57
Identities = 56/281 (19%), Positives = 103/281 (35%), Gaps = 38/281 (13%)
Query: 327 PACILEYVRMLRRQGDPITLTEFVNTLPESP---PKLATRKSRRATSSKPSDPKGKGILK 383
P ++E + +GD ++ PES PK KS+ SS+P
Sbjct: 2833 PPSVVESTKADSTKGD-------ISPSPESVLEGPKDDVEKSKE--SSRPPSVSASITGD 2883
Query: 384 EEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLA 443
+ AS +++ + +R S + + E+ + K SS + +DE LA
Sbjct: 2884 STKDVSRPASVVESVKDEHDKAESRRESIAKVESVIDEAGKSDSKSSSQDSQKDEKSTLA 2943
Query: 444 K----RRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPL---- 495
RR+ ++E +D+ + E ES + PV + K PL
Sbjct: 2944 SKEASRRESVVESSKDDAEKSE-------------SRPESVIASGEPVPRESKSPLDSKD 2990
Query: 496 ---RGPLQQKETAAEQASEPTSEVQRERRSKR--TSDPARAASLTRTSPIRSLSKVITES 550
G + + TA ++ SE S + S + T ++ +R S + L K E
Sbjct: 2991 TSRPGSMVESVTAEDEKSEQQSRRESVAESVKADTKKDGKSQEASRPSSVDELLKDDDEK 3050
Query: 551 INSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAE 591
S + + M + + + ++P++ AE
Sbjct: 3051 QESRRQSITGSHKAMSTMGDESPMDKADKSKEPSRPESVAE 3091
Score = 31.6 bits (70), Expect = 6.3
Identities = 39/204 (19%), Positives = 78/204 (38%), Gaps = 21/204 (10%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQ 414
++ K A + +AT KP + +G+ P+K R P+P P+K+ +
Sbjct: 790 QAEAKTAATGATQATQRKPISRRPRGV---SPSK----------RAPAPGSPVKQAKPKA 836
Query: 415 QQMSDSESSEDTWKDSSSEETEDEDMPLAKRR-------KIILEEEEDEQDDHEIIQNVI 467
+ + + DSS T D A ++ + + E++ E DD + Q V+
Sbjct: 837 ADLKKTRLDKGGTTDSSLVSTPSADEATAAKKLQDLTASQELDAEKQRELDDLKEEQEVV 896
Query: 468 QGIREASDAEESTDSDAIPV-VKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTS 526
+ I +E + + R++P G + E+ E + + S
Sbjct: 897 REIEAVFSRDEMKRQQHQQIKAELREMPAEGTGDGENEPDEEEEYLIIEKEEVEQYTEDS 956
Query: 527 DPARAASLTRTSPIRSLSKVITES 550
+ +S+T+ I+ + ES
Sbjct: 957 IVEQESSMTKEEEIQKHQRDSQES 980
Score = 31.2 bits (69), Expect = 8.2
Identities = 43/227 (18%), Positives = 92/227 (39%), Gaps = 17/227 (7%)
Query: 383 KEEPAKKKAASKTVIIREPSPERPLKRT-SAQQQQMSDSESSEDTWKDSSSEETEDEDMP 441
++ P K AS+ + E + P+K T ++++ +++S ++ T + S ++D P
Sbjct: 1647 RDSPLASKEASRPESVLESVKDEPIKSTEKSRRESVAESFKADSTKDEKSPLTSKDISRP 1706
Query: 442 LAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAE-ESTDSDAIPVVKRRKLPLRGPLQ 500
+ ++ E+ E + R S AE E D+D V +P ++
Sbjct: 1707 ESAVENVMDAVGSAERSQPESVTASRDVSRPESVAESEKDDTDKPESVVESVIPASDVVE 1766
Query: 501 QKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVI 560
++ AA++ ++ + P+ ++R P+ + V ES S +V+
Sbjct: 1767 IEKGAADKEKGVFVSLE----IGKPDSPSEV--ISRPGPV--VESVKPESRRESSTEIVL 1818
Query: 561 P-------EQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQ 600
P E P S ++E ++ T+ ++ AE P +
Sbjct: 1819 PCHAEDSKEPSRPESKVECLKDESEVLKGSTRRESVAESDKSSQPFK 1865
>NP14_RAT (P41777) Nucleolar phosphoprotein p130 (Nucleolar 130 kDa
protein) (140 kDa nucleolar phosphoprotein) (Nopp140)
(Nucleolar and coiled-body phosphoprotein 1)
Length = 704
Score = 51.2 bits (121), Expect = 8e-06
Identities = 48/181 (26%), Positives = 80/181 (43%), Gaps = 21/181 (11%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEE------PAKKKAASKTVIIREPSPERPLK 408
++ P L +++ RA SD + +E+ PAKKKAA V P P P+K
Sbjct: 461 KAAPSLPAKQAPRAGGDSSSDSESSSSEEEKKTPPKPPAKKKAAGAAV----PKP-TPVK 515
Query: 409 RTSAQQQQMSDS--ESSEDTWKDSSSEET------EDEDMPLAKRRKIILEEEEDEQD-- 458
+ +A+ S S +SSE+ K S+ T + +P ++ K E EE+E+D
Sbjct: 516 KAAAESSSSSSSSEDSSEEEKKKPKSKATPKPQAGKANGVPASQNGKAGKESEEEEEDTE 575
Query: 459 DHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQR 518
++ G + E+ D A P K+ KL +++ ++AS P V+
Sbjct: 576 QNKKAAGTKPGSGKKRKHNETADEAATPQSKKVKLQTPNTFPKRKKGEKRASSPFRRVRE 635
Query: 519 E 519
E
Sbjct: 636 E 636
Score = 49.3 bits (116), Expect = 3e-05
Identities = 50/215 (23%), Positives = 83/215 (38%), Gaps = 22/215 (10%)
Query: 358 PKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQM 417
PK A +++ A SS S + +EE KK +KTV+ + P+ P+K+ +
Sbjct: 319 PKKAAAQTQPADSSADSSEESDSSSEEE---KKTPAKTVVSKTPAKPAPVKKKAESSSDS 375
Query: 418 SDSESSED--------TWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQG 469
SDS+SSED K S+ P AK + Q +
Sbjct: 376 SDSDSSEDEAPAKPVSATKSPLSKPAVTPKPPAAKAVATPKQPAGSGQKPQSRKADSSSS 435
Query: 470 IREASDAEESTDSDAIPVVKRRKLPLRGP----LQQKETAAEQASEPTSEVQRERRSKRT 525
E+S +EE ++ K R P Q + +S+ S E +
Sbjct: 436 EEESSSSEEEATKKSVTTPKARVTAKAAPSLPAKQAPRAGGDSSSDSESSSSEEEKKTPP 495
Query: 526 SDPAR----AASLTRTSPIRSLSKVITESINSDSA 556
PA+ A++ + +P++ K ES +S S+
Sbjct: 496 KPPAKKKAAGAAVPKPTPVK---KAAAESSSSSSS 527
Score = 48.5 bits (114), Expect = 5e-05
Identities = 61/255 (23%), Positives = 100/255 (38%), Gaps = 30/255 (11%)
Query: 355 ESPPKLATRK---SRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTS 411
++ PK A K S ++SS SD + P KK A K V+ + P +K T+
Sbjct: 214 KAQPKAANGKAGSSSSSSSSSSSDDSEEEKKAAAPLKKTAPKKQVVAKAP-----VKVTA 268
Query: 412 AQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRR---------KIILEEEEDEQDDHEI 462
A Q+ S SE DSSSEE E++ P+ K+ + L ++ +
Sbjct: 269 APTQKSSSSE-------DSSSEEEEEQKKPMKKKAGPYSSVPPPSVSLSKKSVGAQSPKK 321
Query: 463 IQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRS 522
Q ++D+ E +DS + K + K ++ +E +S+ + S
Sbjct: 322 AAAQTQPADSSADSSEESDSSSEEEKKTPAKTVVSKTPAKPAPVKKKAESSSD-SSDSDS 380
Query: 523 KRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQP 582
PA+ S T++ LSK + V P+QP S P S + +
Sbjct: 381 SEDEAPAKPVSATKS----PLSKPAVTPKPPAAKAVATPKQPAG-SGQKPQSRKADSSSS 435
Query: 583 ETQPQTQAEPQAPKS 597
E + + E KS
Sbjct: 436 EEESSSSEEEATKKS 450
Score = 42.7 bits (99), Expect = 0.003
Identities = 57/266 (21%), Positives = 107/266 (39%), Gaps = 30/266 (11%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTS--- 411
++P K+ +++++SS+ S + + K+ P KKKA + + P P L + S
Sbjct: 261 KAPVKVTAAPTQKSSSSEDSSSEEEEEQKK-PMKKKAGPYSSV---PPPSVSLSKKSVGA 316
Query: 412 ------AQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQN 465
A Q Q +DS + DSSSEE + K ++ + + ++
Sbjct: 317 QSPKKAAAQTQPADSSADSSEESDSSSEEEKKTP------AKTVVSKTPAKPAP---VKK 367
Query: 466 VIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQA----SEPTSEVQRERR 521
+ ++SD++ S D V K PL P + A +A +P Q+ +
Sbjct: 368 KAESSSDSSDSDSSEDEAPAKPVSATKSPLSKPAVTPKPPAAKAVATPKQPAGSGQKPQS 427
Query: 522 SKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQ 581
K S + S + S ++ + A +P + P + +S+ +E
Sbjct: 428 RKADSSSSEEESSSSEEEATKKSVTTPKARVTAKAAPSLPAKQAPRAGG-DSSSDSESSS 486
Query: 582 PETQPQTQAEPQAPKSPIQTATTAIP 607
E + +T +P A K + A A+P
Sbjct: 487 SEEEKKTPPKPPAKK---KAAGAAVP 509
Score = 35.0 bits (79), Expect = 0.57
Identities = 51/266 (19%), Positives = 108/266 (40%), Gaps = 30/266 (11%)
Query: 352 TLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTS 411
+LP+ K A + S ++S + S+ + + K++P ++KA P P++ +
Sbjct: 113 SLPQHAGKAAAKASESSSSEESSEEEEEKDKKKKPVQQKAVKPQAKAVRPPPKK-----A 167
Query: 412 AQQQQMSDSESSEDTWKDSSSEETED-EDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGI 470
+ SDS S ++ + + P + K + + +
Sbjct: 168 ESSESESDSSSEDEAPQTQKPKAAATAAKAPTKAQTKAPAKPGPPAKAQPKAANGKAGSS 227
Query: 471 REASDAEESTDSD-----AIPVVK---RRKLPLRGPLQ------QKETAAEQASEPTSEV 516
+S + S DS+ A P+ K ++++ + P++ QK +++E +S E
Sbjct: 228 SSSSSSSSSDDSEEEKKAAAPLKKTAPKKQVVAKAPVKVTAAPTQKSSSSEDSSSEEEEE 287
Query: 517 QRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQ 576
Q++ K+ + P SLSK +S+ + S Q P +S +S +
Sbjct: 288 QKKPMKKKAGPYSSV-----PPPSVSLSK---KSVGAQSPKKA-AAQTQPADSSADSSEE 338
Query: 577 TEPIQPETQPQTQAEPQAPKSPIQTA 602
++ E + +T A+ K+P + A
Sbjct: 339 SDS-SSEEEKKTPAKTVVSKTPAKPA 363
>IGA4_HAEIN (P45386) Immunoglobulin A1 protease precursor (EC
3.4.21.72) (IGA1 protease)
Length = 1849
Score = 51.2 bits (121), Expect = 8e-06
Identities = 55/257 (21%), Positives = 101/257 (38%), Gaps = 14/257 (5%)
Query: 355 ESPPK--LATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSA 412
E+ PK AT+ + S PS+ K + A EP+P+
Sbjct: 1075 ETAPKSDTATQTENPNSESVPSETTEKVAENPPQENETVAKNEQEATEPTPQNGEVAKED 1134
Query: 413 QQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIRE 472
Q ++++++E T + +EET+ + + + E + E+ + ++ + +E
Sbjct: 1135 QPTVEANTQTNEATQSEGKTEETQTAETKSEPTESVTVSENQPEKTVSQSTEDKVVVEKE 1194
Query: 473 ASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAA 532
E+ ++ P V ++ P +Q E A E+ PT E ++ + + P A
Sbjct: 1195 EKAKVETEETQKAPQVTSKE-----PPKQAEPAPEEV--PTDTNAEEAQALQQTQPTTVA 1247
Query: 533 SLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEP 592
+ TSP SK E+ P P+ + T+TE + +TQ P
Sbjct: 1248 AAETTSP---NSKPAEETQQPSEKTNAEPVTPVVSENTATQPTETEETAKVEKEKTQEVP 1304
Query: 593 Q--APKSPIQTATTAIP 607
Q + +SP Q A P
Sbjct: 1305 QVASQESPKQEQPAAKP 1321
Score = 46.2 bits (108), Expect = 2e-04
Identities = 56/278 (20%), Positives = 115/278 (41%), Gaps = 31/278 (11%)
Query: 347 TEFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERP 406
+E + E+PP+ ++ + P+ + KE+ +A ++T + +
Sbjct: 1096 SETTEKVAENPPQENETVAKNEQEATEPTPQNGEVAKEDQPTVEANTQTNEATQSEGKTE 1155
Query: 407 LKRTSAQQQQMSDSES-SEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQN 465
+T+ + + ++S + SE+ + + S+ TED K+++E+EE + + E Q
Sbjct: 1156 ETQTAETKSEPTESVTVSENQPEKTVSQSTED---------KVVVEKEEKAKVETEETQK 1206
Query: 466 VIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRT 525
Q + + + +P + LQQ + A+E TS + ++ T
Sbjct: 1207 APQVTSKEPPKQAEPAPEEVPTDTNAE--EAQALQQTQPTTVAAAETTS--PNSKPAEET 1262
Query: 526 SDPARAASLTRTSPIRSLSKVITESINSDSALVVIPE--QPMPISTSLPTSTQTEP-IQP 582
P+ + +P+ S T+ ++ V E Q +P S + Q +P +P
Sbjct: 1263 QQPSEKTNAEPVTPVVS-ENTATQPTETEETAKVEKEKTQEVPQVASQESPKQEQPAAKP 1321
Query: 583 ETQPQTQAEP-------------QAPKSPIQTATTAIP 607
+ Q + QAEP P++ QT +TA+P
Sbjct: 1322 QAQTKPQAEPARENVLTTKNVGEPQPQAQPQTQSTAVP 1359
Score = 45.8 bits (107), Expect = 3e-04
Identities = 42/213 (19%), Positives = 90/213 (41%), Gaps = 28/213 (13%)
Query: 392 ASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETED---EDMPLAKRRKI 448
A+++ I E RP + ++ + + S+E K ++ +TE+ E +P K+
Sbjct: 1043 ATESAIASEQPETRPAETAQPAMEETNTANSTETAPKSDTATQTENPNSESVPSETTEKV 1102
Query: 449 ILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQ 508
E Q++ + +N +A E T + V K + P + T +
Sbjct: 1103 ---AENPPQENETVAKN-------EQEATEPTPQNG-EVAKEDQ-----PTVEANTQTNE 1146
Query: 509 ASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPIS 568
A++ + + + ++ S+P + +++ P +++S+ +++ +VV E+ +
Sbjct: 1147 ATQSEGKTEETQTAETKSEPTESVTVSENQPEKTVSQ------STEDKVVVEKEEKAKVE 1200
Query: 569 TSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQT 601
T TQ P +P QAEP + P T
Sbjct: 1201 TE---ETQKAPQVTSKEPPKQAEPAPEEVPTDT 1230
Score = 44.3 bits (103), Expect = 0.001
Identities = 57/306 (18%), Positives = 117/306 (37%), Gaps = 53/306 (17%)
Query: 294 EKIQVAPKEDTSDEVRQRDYPIDDFPIWSKKDNPACILEYVRMLRRQGDPITLTEFVNTL 353
E+ Q AP+ + + +Q + ++ P + + + +Q P T+ T
Sbjct: 1202 EETQKAPQVTSKEPPKQAEPAPEEVPTDTNAEEAQAL--------QQTQPTTVAAAETTS 1253
Query: 354 PESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQ 413
P S P T++ T+++P P ++ E + +P T +
Sbjct: 1254 PNSKPAEETQQPSEKTNAEPVTP--------------------VVSENTATQP---TETE 1290
Query: 414 QQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREA 473
+ + E +++ +S+E+ ++ P AK + + E +++ +NV + +A
Sbjct: 1291 ETAKVEKEKTQEV-PQVASQESPKQEQPAAKPQAQTKPQAEPARENVLTTKNVGEPQPQA 1349
Query: 474 SDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAAS 533
+ST P P Q + E A E S V + +TS A+
Sbjct: 1350 QPQTQSTAVPTTGETAANSKPAAKPQAQAKPQTEPARENVSTVNTKEPQSQTS-----AT 1404
Query: 534 LTRTSPIRSLSKVITE-----SINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQT 588
++ P + S + + SIN+ SA +T T+ +++ Q ET +
Sbjct: 1405 VSTEQPAKETSSNVEQPAPENSINTGSA-----------TTMTETAEKSDKPQMETVTEN 1453
Query: 589 QAEPQA 594
+P+A
Sbjct: 1454 DRQPEA 1459
>CEC1_CAEEL (P34618) Chromo domain protein cec-1
Length = 304
Score = 51.2 bits (121), Expect = 8e-06
Identities = 54/263 (20%), Positives = 97/263 (36%), Gaps = 52/263 (19%)
Query: 342 DPITLTEFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREP 401
DP + F ++ +K + A K + K + + A A + P
Sbjct: 47 DPTLIEAFFTREAARKAEIKAKKDKMAAGKKGASSKASASVSKASASTPARGAKAAPKPP 106
Query: 402 SPERPLKRTSAQQQQMSDSESSEDTWKDSSS----------EETEDEDMPLAKRRKIILE 451
+ P KR Q+ D DT ++ SS EE ED++ P+ K++K + E
Sbjct: 107 PKKSPPKR---QRLAGGDIRPDSDTDEEHSSADKKSKAEDEEEVEDDEEPVPKKKKEVQE 163
Query: 452 E---------------------------EEDEQDDHEIIQ---------NVIQGIREASD 475
E EEDE+++ E +Q + + E +
Sbjct: 164 EPEEEESVEGEDEEESQEVEDLKEDEKMEEDEKEEEEDVQLESEKNEKEEEEEKVEEKKE 223
Query: 476 AEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLT 535
EE + + I +V K + + + A + SEP+S E+ + AA +
Sbjct: 224 EEEEEEEEEIQLVIVEKTVIETTIVEPAVATPEPSEPSSS---EKAVVENGSSSAAAGNS 280
Query: 536 RTSPIRSLSKVITESINSDSALV 558
+ P S +V+T + D A++
Sbjct: 281 ASKPEVSAVEVVTVEDDDDIAII 303
>YCDA_DROME (Q9VGW1) Putative cadherin CG4509 precursor
Length = 1483
Score = 49.7 bits (117), Expect = 2e-05
Identities = 55/215 (25%), Positives = 93/215 (42%), Gaps = 32/215 (14%)
Query: 348 EFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEE-------PAKKKAASKTVIIRE 400
E+V P + P+ RR T +P P+ + ++ EE P K++ + R
Sbjct: 950 EYVRFNPPNSPEGVYWIKRRRTKKRPRQPRKRIVMVEEVKRKIRTPIKEEEEVQERKKRV 1009
Query: 401 PSPERPLK--RTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQD 458
P P++PL+ +TS ++Q+SD ES +D ++ S+ + + E D D
Sbjct: 1010 P-PKKPLRETKTSILRKQLSD-ESRKDQSRNGESQTGNRHRSESDSHNRDMFMEITDSMD 1067
Query: 459 D------HEIIQNVIQ-------GIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETA 505
+ H I + ++ G + D E S DSD +V R P R ++
Sbjct: 1068 ELASPGSHSIRKIQVEKYYKHSDGDFDEDDTEYSIDSDGDEIVIRTNYPSRAQENERYRR 1127
Query: 506 AEQA-SEPTSEVQRERRSKRTSDPARAASLTRTSP 539
E+ +EP + V R+R PAR +S T + P
Sbjct: 1128 QERTYAEPENPVDRKR-------PARKSSPTDSQP 1155
>YEMA_DROME (P25992) Yemanuclein-alpha
Length = 1002
Score = 49.3 bits (116), Expect = 3e-05
Identities = 54/223 (24%), Positives = 96/223 (42%), Gaps = 51/223 (22%)
Query: 388 KKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRK 447
KK ++T I + PER KR + + S S S +D D + E +DE
Sbjct: 184 KKSYTTRTDAIIK-MPERSRKRMVSSSSESSSSSSGDDDENDDGNNEEDDES-------- 234
Query: 448 IILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAE 507
+ E+D +++ E SD+E+ ++S+++ + A+
Sbjct: 235 ---DSEDDSEENDE------------SDSEDDSESESLED------------EDSAATAK 267
Query: 508 QASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPI 567
+S+ Q +R + ++ +S + TS + +K IT S +S+S P P
Sbjct: 268 SSSKYKDNHQAKRAKVIVTGKSKPSSSSLTSGKKPPTKPITTSSSSNS--------PRPS 319
Query: 568 STSL-PTSTQTEPI--QPETQ----PQTQAEPQAPKSPIQTAT 603
+ + T +PI QP +Q PQ+QA+ QA K ++T T
Sbjct: 320 TVEISDTEDGQDPIQTQPSSQLQSLPQSQAQAQALKKVVKTTT 362
>IGA2_HAEIN (P45384) Immunoglobulin A1 protease precursor (EC
3.4.21.72) (IGA1 protease)
Length = 1702
Score = 48.9 bits (115), Expect = 4e-05
Identities = 57/269 (21%), Positives = 114/269 (42%), Gaps = 31/269 (11%)
Query: 354 PESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQ 413
P PP AT T ++ S + K + K E + ++ + E + +P + + Q
Sbjct: 1029 PVPPPAPATPSETTETVAENSKQESKTVEKNEQDATETTAQNGEVAEEA--KPSVKANTQ 1086
Query: 414 QQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREA 473
+++ S S + + + +ET + K K +E+EE + + + IQ Q E
Sbjct: 1087 TNEVAQSGSETEETQTTEIKETAKVE----KEEKAKVEKEEKAKVEKDEIQEAPQMASET 1142
Query: 474 SDA-------EESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERR-SKRT 525
S E STD+ V+ ++ + Q AA +A+ P S+ E + S++T
Sbjct: 1143 SPKQAKPAPKEVSTDTK----VEETQVQAQPQTQSTTVAAAEATSPNSKPAEETQPSEKT 1198
Query: 526 S----DPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQ 581
+ P + + T + + + T + ++ E P S + P Q+E +Q
Sbjct: 1199 NAEPVTPVVSKNQTENTTDQPTEREKTAKVETEKT----QEPPQVASQASPKQEQSETVQ 1254
Query: 582 PETQPQTQAEP-----QAPKSPIQTATTA 605
P+ +++ P + ++ +QT T+A
Sbjct: 1255 PQAVLESENVPTVNNAEEVQAQLQTQTSA 1283
Score = 35.0 bits (79), Expect = 0.57
Identities = 47/228 (20%), Positives = 82/228 (35%), Gaps = 63/228 (27%)
Query: 355 ESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKA----------ASKTVIIREPSPE 404
+ PP++A++ S + S+ P+ + P A S TV ++P+PE
Sbjct: 1234 QEPPQVASQASPKQEQSETVQPQAVLESENVPTVNNAEEVQAQLQTQTSATVSTKQPAPE 1293
Query: 405 RPLKRTSA----------QQQQMSDSESSED-------TWKDSSSEETEDEDMPLAKRRK 447
+ SA + Q + S+ED T D+S + P ++RR+
Sbjct: 1294 NSINTGSATAITETAEKSDKPQTETAASTEDASQHKANTVADNSVANNSESSEPKSRRRR 1353
Query: 448 IILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAE 507
I + +E + AEE+T + ET
Sbjct: 1354 SISQPQE-------------------TSAEETTAAST-----------------DETTIA 1377
Query: 508 QASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDS 555
S+ + +R RRS R+ S T +R L+ T ++ SD+
Sbjct: 1378 DNSKRSKPNRRSRRSVRSEPTVTNGSDRSTVALRDLTSTNTNAVISDA 1425
>NUKS_RAT (Q9EPJ0) Nuclear ubiquitous casein and cyclin-dependent
kinases substrate
Length = 243
Score = 48.5 bits (114), Expect = 5e-05
Identities = 57/241 (23%), Positives = 94/241 (38%), Gaps = 50/241 (20%)
Query: 371 SKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDS 430
S P K + +E K+++ + E S E+ +K DSE +D K S
Sbjct: 30 SGPPAKKIRSSPREAKNKRRSGKNSQEDSEDSEEKDVKTKKDDSHSAEDSEDEKDDHK-S 88
Query: 431 SSEETEDEDMPLAKRRKIILE----EEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIP 486
++ + +K+R+++LE EEE E+DD Q G E E+ DSD
Sbjct: 89 VRQQRQAASKAASKQREMLLEDVGSEEEPEEDDEAPFQEKDSGSDEDFLMEDDDDSD--- 145
Query: 487 VVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKV 546
G ++K + S+P ER+ K+ P A++T SP++ KV
Sbjct: 146 ---------YGSSKKKNKKMVKKSKP------ERKEKKMPKPRLKATVT-PSPVKGKGKV 189
Query: 547 ITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQTATTAI 606
PT+++T E P + E + +SP + T+A
Sbjct: 190 -----------------------GRPTASKT---SKEKTPSPKEEDEEAESPPEKKTSAS 223
Query: 607 P 607
P
Sbjct: 224 P 224
>NFM_CHICK (P16053) Neurofilament triplet M protein (160 kDa
neurofilament protein) (Neurofilament medium
polypeptide) (NF-M)
Length = 857
Score = 48.1 bits (113), Expect = 6e-05
Identities = 46/230 (20%), Positives = 103/230 (44%), Gaps = 24/230 (10%)
Query: 383 KEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPL 442
+EE +++ A + + E E+ ++ + ++++ + E+ K ++EE
Sbjct: 481 QEEEQEEEKAEEEAVEEEAVSEKAAEQAAEEEEKEEEEAEEEEAAKSDAAEEG------- 533
Query: 443 AKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAE-ESTDSDAIPVVKRRKLPLRGPLQQ 501
+++ I E+EE E+ + E + +G E + A+ E S P K P + P+ +
Sbjct: 534 GSKKEEIEEKEEGEEAEEE--EAEAKGKAEEAGAKVEKVKSP--PAKSPPKSPPKSPVTE 589
Query: 502 KETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIP 561
+ A ++A+ +EV +++++++ ++ A + S K T + S
Sbjct: 590 QAKAVQKAA---AEVGKDQKAEKAAEKA-----AKEEKAASPEKPATPKVTSPEKPATPE 641
Query: 562 EQPMPISTSLPTSTQT--EPIQPE--TQPQTQAEPQAPKSPIQTATTAIP 607
+ P P P ++ +P PE P+ A P+ P++P + A+ P
Sbjct: 642 KPPTPEKAITPEKVRSPEKPTTPEKVVSPEKPASPEKPRTPEKPASPEKP 691
Score = 41.2 bits (95), Expect = 0.008
Identities = 51/233 (21%), Positives = 85/233 (35%), Gaps = 29/233 (12%)
Query: 383 KEEPAKKKAASKTV--IIREPSPERPLKRT----SAQQQQMSDSESSEDTWKDSSSEETE 436
K EP K K K V II E E SA ++M+ E+ ++ + EE
Sbjct: 436 KIEPPKLKVQHKFVEEIIEETKVEDEKSEMEDALSAIAEEMAAKAQEEEQEEEKAEEEAV 495
Query: 437 DEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLR 496
+E+ K + EEEE E+++ E EE+ SDA +K +
Sbjct: 496 EEEAVSEKAAEQAAEEEEKEEEEAE--------------EEEAAKSDAAEEGGSKKEEIE 541
Query: 497 GPLQQKETAAEQ------ASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITES 550
+ +E E+ A E ++V++ + S P T +++ K E
Sbjct: 542 EKEEGEEAEEEEAEAKGKAEEAGAKVEKVKSPPAKSPPKSPPKSPVTEQAKAVQKAAAEV 601
Query: 551 INSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQTAT 603
A + + P T + T P+ A P+ P +P + T
Sbjct: 602 GKDQKAEKAAEKAAKEEKAASPEKPATPKV---TSPEKPATPEKPPTPEKAIT 651
>YKF4_YEAST (P35732) Hypothetical 84.0 kDa protein in NUP120-CSE4
intergenic region
Length = 738
Score = 47.4 bits (111), Expect = 1e-04
Identities = 56/243 (23%), Positives = 95/243 (39%), Gaps = 46/243 (18%)
Query: 358 PKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQM 417
P L+++K+ TS+ S PK K KA KT E PL+ + ++++
Sbjct: 223 PSLSSKKTTSRTSA--SQPKKMSWAAIATPKPKAVKKT--------ESPLENVAELKKEI 272
Query: 418 SDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAE 477
SD + +D K +SEE +E A+ ++ E +E +D + E E
Sbjct: 273 SDIK--KDDQKSEASEEKVNEQETSAQEQEEETAEPSEENEDR---------VPEVDGEE 321
Query: 478 ESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRT 537
+++ VK + Q+++ A E T ++ S S+P + T
Sbjct: 322 VQEEAEKKEQVKEEEQTAEELEQEQDNVAAPEEEVTVVEEKVEISAVISEPPEDQANTVP 381
Query: 538 SPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKS 597
P + + P+QP +P QP+ QPQ Q +PQ P+
Sbjct: 382 QPQQQSQQ---------------PQQPQQ---------PQQPQQPQ-QPQQQQQPQQPQQ 416
Query: 598 PIQ 600
P Q
Sbjct: 417 PQQ 419
>NUCL_CHICK (P15771) Nucleolin (Protein C23)
Length = 694
Score = 45.8 bits (107), Expect = 3e-04
Identities = 47/175 (26%), Positives = 78/175 (43%), Gaps = 41/175 (23%)
Query: 374 SDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDSSSE 433
SD + + + +PA KK+A+ V ++P+ P K+ S ++++ D E E+ +
Sbjct: 136 SDEEEEPAVPVKPAAKKSAA-AVPAKKPAVV-PAKQESEEEEEEDDEEEDEE------DD 187
Query: 434 ETEDEDM-----PLAK---------RRKIILEEEEDEQDDHEIIQNVIQGIREASDAEES 479
E+EDE M P+ K + K E+EEDE+D+ E ++ E D EES
Sbjct: 188 ESEDEAMDTTPAPVKKPTPAKATPAKAKAESEDEEDEEDEDEDEEDEDD---EEEDEEES 244
Query: 480 TDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASL 534
D + KE ++ E ++ E + K+T PA A SL
Sbjct: 245 EDEKPV----------------KEAPGKRKKEMANKSAPEAKKKKTETPASAFSL 283
Score = 32.3 bits (72), Expect = 3.7
Identities = 35/212 (16%), Positives = 85/212 (39%), Gaps = 16/212 (7%)
Query: 356 SPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQ 415
+PP +S SS + G+ ++ P K++ A+ T + +P + K + ++
Sbjct: 18 APPPKKVEESEEEESSDLEESSGEEVMVP-PKKQQKAAVTPAKKAATPAK--KAATPAKK 74
Query: 416 QMSDSESSEDTWKDSSSEETEDEDMPL---AKRRKIILEEEEDEQDDHEIIQNVIQGIRE 472
++ ++ + T + + + + AK K +EE +E+D+ + + E
Sbjct: 75 AVTPAKKAVATPAKKAVAPSPKKAAVVGKGAKNGKNAKKEESEEEDEDDEDDEEDEDEEE 134
Query: 473 ASDAEE----------STDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRS 522
SD EE + A+P K +P + +++E ++ + + +
Sbjct: 135 ESDEEEEPAVPVKPAAKKSAAAVPAKKPAVVPAKQESEEEEEEDDEEEDEEDDESEDEAM 194
Query: 523 KRTSDPARAASLTRTSPIRSLSKVITESINSD 554
T P + + + +P ++ ++ E D
Sbjct: 195 DTTPAPVKKPTPAKATPAKAKAESEDEEDEED 226
Score = 32.0 bits (71), Expect = 4.8
Identities = 32/117 (27%), Positives = 49/117 (41%), Gaps = 23/117 (19%)
Query: 354 PESPPKLATRKSRRATSSKP---------SDPKGKGILKEEPAKKKAASKTVIIREPSPE 404
P P K A +KS A +K S+ + + +EE + + + P+P
Sbjct: 142 PAVPVKPAAKKSAAAVPAKKPAVVPAKQESEEEEEEDDEEEDEEDDESEDEAMDTTPAPV 201
Query: 405 R---PLKRTSAQQQQMS-DSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQ 457
+ P K T A+ + S D E ED +D E+ E+ED EE EDE+
Sbjct: 202 KKPTPAKATPAKAKAESEDEEDEEDEDEDEEDEDDEEEDE----------EESEDEK 248
>ACIN_MOUSE (Q9JIX8) Apoptotic chromatin condensation inducer in the
nucleus (Acinus)
Length = 1338
Score = 45.8 bits (107), Expect = 3e-04
Identities = 44/249 (17%), Positives = 94/249 (37%), Gaps = 25/249 (10%)
Query: 342 DPITLTEFVNTLPESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREP 401
D +T E V +L PP + +R P+ EE + +
Sbjct: 171 DEMTHPEGVASL--LPPDFQSSLNRPELELSTHSPRKSSSFSEEKGESD---------DE 219
Query: 402 SPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHE 461
P + +R+S +Q S T ++ +ET ++ + R + +EEEE+E+++
Sbjct: 220 KPRKGERRSSRVRQAKSKLPEYSQTAEEEEDQETPSRNLRVRADRNLKIEEEEEEEEE-- 277
Query: 462 IIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRER- 520
E D EE + D + + P + +E ++P V +
Sbjct: 278 ---------EEDDDDEEEEEVDEAQKSREAEAPTLKQFEDEEGEERTRAKPEKVVDEKPL 328
Query: 521 --RSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTE 578
RS+ + + +TR+ S + + ++ +V +P++ + +S +++
Sbjct: 329 NIRSQEKGELEKGGRVTRSQEEARRSHLARQQQEKETQIVSLPQEENEVKSSQSLEEKSQ 388
Query: 579 PIQPETQPQ 587
P P+
Sbjct: 389 SPSPPPLPE 397
>ACIN_HUMAN (Q9UKV3) Apoptotic chromatin condensation inducer in the
nucleus (Acinus)
Length = 1341
Score = 45.8 bits (107), Expect = 3e-04
Identities = 51/244 (20%), Positives = 96/244 (38%), Gaps = 39/244 (15%)
Query: 383 KEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTW--KDSSSEETEDEDM 440
+ P K + S+ + R +R S++ +Q ++ SE + ++ +ET ++
Sbjct: 200 RHSPRKSSSISEEKGDSDDEKPRKGERRSSRVRQARAAKLSEGSQPAEEEEDQETPSRNL 259
Query: 441 PLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRK-----LPL 495
+ R + EEEE+E+++ E + E +E S P++K K +P
Sbjct: 260 RVRADRNLKTEEEEEEEEEEE------EDDEEEEGDDEGQKSREAPILKEFKEEGEEIPR 313
Query: 496 RGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTR---------TSPIRSLSKV 546
P + + + S+ ++R R R+ + AR + L R TSP+ +
Sbjct: 314 VKPEEMMDERPKTRSQEQEVLERGGRFTRSQEEARKSHLARQQQEKEMKTTSPLEEEERE 373
Query: 547 ITES--------------INSD---SALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQ 589
I S + D ++LV +PEQ + P E P PQ
Sbjct: 374 IKSSQGLKEKSKSPSPPRLTEDRKKASLVALPEQTASEEETPPPLLTKEASSPPPHPQLH 433
Query: 590 AEPQ 593
+E +
Sbjct: 434 SEEE 437
Score = 35.8 bits (81), Expect = 0.33
Identities = 55/274 (20%), Positives = 104/274 (37%), Gaps = 28/274 (10%)
Query: 353 LPESPPKLATRKSRRATSSKPS---DPKGKGILKEEPAKKKAASKTVIIREPSPERPLKR 409
L +S ++R S ++SS S P G P + K + + + P R
Sbjct: 570 LKQSADSSSSRSSSSSSSSSRSRSRSPDSSGSRSHSPLRSKQRD---VAQARTHANPRGR 626
Query: 410 TSAQQQQMSDSESSEDTWKDSSSEETEDEDMP-LAKRRKIILEEEEDEQDDHEIIQNVIQ 468
+ S+S S + S+S + P +++ E +D E+ +
Sbjct: 627 PKMGSRSTSESRSRSRSRSRSASSNSRKSLSPGVSRDSSTSYTETKDPSSGQEVATPPVP 686
Query: 469 GIREASDAEE-STDSDAIPV------------VKRRKLPLRGPLQQKETAAEQASEPTSE 515
++ E ST S ++ V +R P RG K+ AE+A EP +
Sbjct: 687 QLQVCEPKERTSTSSSSVQARRLSQPESAEKHVTQRLQPERG--SPKKCEAEEA-EPPAA 743
Query: 516 VQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTST 575
Q + +TS + + T + + +T + + +PE PMPI+ +
Sbjct: 744 TQPQTSETQTSHLPESERIHHTVEEK---EEVTMDTSENRPENDVPEPPMPIADQVSNDD 800
Query: 576 QTEPIQPETQPQTQAEPQAPKSPIQ--TATTAIP 607
+ E + + + + P++ K I +AT +P
Sbjct: 801 RPEGSVEDEEKKESSLPKSFKRKISVVSATKGVP 834
>PCFB_HUMAN (O94913) Pre-mRNA cleavage complex II protein Pcf11
(Fragment)
Length = 1654
Score = 45.4 bits (106), Expect = 4e-04
Identities = 55/269 (20%), Positives = 110/269 (40%), Gaps = 19/269 (7%)
Query: 350 VNTLPESPPKLA-TRKSRRATSSKPSDPK-GKGILKEE------PAKKKAASKTVIIREP 401
+NTL +S K + T S + SSK K G+ I K+E +K K+ S + + +
Sbjct: 418 MNTLNQSDTKTSKTIPSEKLNSSKQEKSKSGEKITKKELDQLDSKSKSKSKSPSPLKNKL 477
Query: 402 SPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDE---QD 458
S + LK ++ ++SD + K ++T+ +D + ++RK ++++DE
Sbjct: 478 SHTKDLKNQESESMRLSDMNKRDPRLKKHLQDKTDGKDDDVKEKRKTAEKKDKDEHMKSS 537
Query: 459 DHEII---QNVIQGIREASD--AEESTDSDAIP--VVKRRKLPLRGPLQQKETAAEQASE 511
+H + +I GI + D EES P R++ R P +
Sbjct: 538 EHRLAGSRNKIINGIVQKQDTITEESEKQGTKPGRSSTRKRSRSRSPKSRSPIIHSPKRR 597
Query: 512 PTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSL 571
+R +RS + +A + ++ +S + T D +Q + +
Sbjct: 598 DRRSPKRRQRSMSPTSTPKAGKIRQSGAKQSHMEEFTPPSREDRNAKRSTKQDIRDPRRM 657
Query: 572 PTSTQTEPIQPETQPQTQAEPQAPKSPIQ 600
+ + P + Q T++ + PK ++
Sbjct: 658 KKTEEERPQETTNQHSTKSGTE-PKENVE 685
>NUKS_MOUSE (Q80XU3) Nuclear ubiquitous casein and cyclin-dependent
kinases substrate (JC7)
Length = 234
Score = 45.4 bits (106), Expect = 4e-04
Identities = 45/180 (25%), Positives = 76/180 (42%), Gaps = 24/180 (13%)
Query: 371 SKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWKDS 430
S P K + +E K+++ + E S E+ +K DSE +D K+
Sbjct: 30 SGPPAKKIRSSPREAKNKRRSGKNSQEDSEDSEEKDVKTKKDDSHSAEDSEDEKDDHKNV 89
Query: 431 SSEETEDEDMPLAKRRKIILE----EEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIP 486
++ + +K+R+++LE EEE E+DD Q G E E+ DSD
Sbjct: 90 R-QQRQAASKAASKQREMLLEDVGSEEEPEEDDEAPFQEKDSGSDEDFLMEDDDDSD--- 145
Query: 487 VVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKV 546
G ++K + S+P ER+ K+ P A++T SP++ +KV
Sbjct: 146 ---------YGSSKKKNKKMVKKSKP------ERKEKKMPKPRLKATVT-PSPVKGKAKV 189
Score = 32.3 bits (72), Expect = 3.7
Identities = 26/81 (32%), Positives = 38/81 (46%), Gaps = 8/81 (9%)
Query: 354 PESPPKLATRKSRRATSSKPSDPKGKGILKEEPAKKKAASKTVIIREPSPERPLKRTSAQ 413
PE K + +AT + PS KGK + A KK+ KT PSP+ + +
Sbjct: 162 PERKEKKMPKPRLKATVT-PSPVKGKAKVGRPTASKKSKEKT-----PSPKEEDEEAESP 215
Query: 414 QQQMSDSESSEDTWKDSSSEE 434
++ S E SED + SS E+
Sbjct: 216 PEKKSGDEGSED--EASSGED 234
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.129 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 68,848,697
Number of Sequences: 164201
Number of extensions: 2972529
Number of successful extensions: 16753
Number of sequences better than 10.0: 690
Number of HSP's better than 10.0 without gapping: 84
Number of HSP's successfully gapped in prelim test: 636
Number of HSP's that attempted gapping in prelim test: 13591
Number of HSP's gapped (non-prelim): 2179
length of query: 607
length of database: 59,974,054
effective HSP length: 116
effective length of query: 491
effective length of database: 40,926,738
effective search space: 20095028358
effective search space used: 20095028358
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0234.12


