

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0228a.2
(510 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
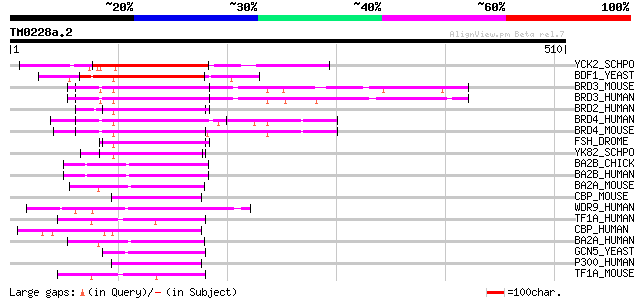
Score E
Sequences producing significant alignments: (bits) Value
YCK2_SCHPO (Q9Y7N0) Hypothetical bromodomain protein C1450.02 106 1e-22
BDF1_YEAST (P35817) BDF1 protein 99 3e-20
BRD3_MOUSE (Q8K2F0) Bromodomain-containing protein 3 (Bromodomai... 99 3e-20
BRD3_HUMAN (Q15059) Bromodomain-containing protein 3 (RING3-like... 95 4e-19
BRD2_HUMAN (P25440) Bromodomain-containing protein 2 (RING3 prot... 94 8e-19
BRD4_HUMAN (O60885) Bromodomain-containing protein 4 (HUNK1 prot... 94 1e-18
BRD4_MOUSE (Q9ESU6) Bromodomain-containing protein 4 (Mitotic ch... 93 1e-18
FSH_DROME (P13709) Female sterile homeotic protein (Fragile-chor... 91 9e-18
YK82_SCHPO (Q9HGP4) Hypothetical bromodomain protein C631.02 84 7e-16
BA2B_CHICK (Q9DE13) Bromodomain adjacent to zinc finger domain 2... 76 2e-13
BA2B_HUMAN (Q9UIF8) Bromodomain adjacent to zinc finger domain 2... 74 9e-13
BA2A_MOUSE (Q91YE5) Bromodomain adjacent to zinc finger domain 2... 71 8e-12
CBP_MOUSE (P45481) CREB-binding protein (EC 2.3.1.48) 70 2e-11
WDR9_HUMAN (Q9NSI6) WD-repeat protein 9 69 2e-11
TF1A_HUMAN (O15164) Transcription intermediary factor 1-alpha (T... 69 4e-11
CBP_HUMAN (Q92793) CREB-binding protein (EC 2.3.1.48) 68 5e-11
BA2A_HUMAN (Q9UIF9) Bromodomain adjacent to zinc finger domain 2... 68 5e-11
GCN5_YEAST (Q03330) Histone acetyltransferase GCN5 (EC 2.3.1.48) 67 8e-11
P300_HUMAN (Q09472) E1A-associated protein p300 (EC 2.3.1.48) 67 1e-10
TF1A_MOUSE (Q64127) Transcription intermediary factor 1-alpha (T... 67 1e-10
>YCK2_SCHPO (Q9Y7N0) Hypothetical bromodomain protein C1450.02
Length = 578
Score = 106 bits (265), Expect = 1e-22
Identities = 53/112 (47%), Positives = 72/112 (63%), Gaps = 6/112 (5%)
Query: 77 RKGSTQ---CAAILKCLMSYPY---SWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKL 130
RK ++Q C+ +LK L Y ++ F +PVDPVA + PDYF +I +PMDL TI SKL
Sbjct: 251 RKNNSQMRFCSTVLKELYKRQYESFAFPFYQPVDPVACDCPDYFDVIKEPMDLSTIQSKL 310
Query: 131 KKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDK 182
KN YS +EEF +D+ L F+N TYNPP VH+M ++L +F KW+ K
Sbjct: 311 NKNEYSTLEEFESDILLMFNNCFTYNPPGTPVHVMGRQLENVFKEKWEARPK 362
Score = 85.9 bits (211), Expect = 2e-16
Identities = 83/303 (27%), Positives = 124/303 (40%), Gaps = 40/303 (13%)
Query: 10 TRKLKIKFSTKRIEVDPGQKCEFGQRGLQTDENGRCNSAEKLF-KPDSIKRGPPEGFEGQ 68
+R+ ++K TK + G G +Q+ + + E L K DS G G Q
Sbjct: 5 SRENEVKAETKDEIANDGSPQLNGDNNIQSSDGHNDENEESLSRKRDS--SGATVGDLKQ 62
Query: 69 KEKR-----------QKIDRKG-----STQCAAILKCLMSYPYSWVFNKPVDPVALNIPD 112
+EK +KI G C AI++ L S F PVDP+ NIPD
Sbjct: 63 EEKESMPKKEPEPTVKKIRGSGMPPPQQKYCLAIVRQLKRTKNSAPFKVPVDPIKQNIPD 122
Query: 113 YFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQI 172
Y I+ PMDLGTI KL YS +EF D+ L FSN YN + V M K L ++
Sbjct: 123 YPTIVKNPMDLGTIEKKLTSYEYSVPQEFIDDMNLMFSNCFLYNGTESPVGSMGKALQEV 182
Query: 173 FDSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQKDTFPKKSQVSEHKRVLKI 232
F+ + K + + + +K++ +KS + PR +R +
Sbjct: 183 FERQLKQL-------PDAEQPAAAPVKKSKQKSASTAPPRT-------------RRNSSV 222
Query: 233 SSLAARDAKVEVPKLSQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTS-PLERK 291
SS +A A PK + K + + + R + KRQ +S + P +
Sbjct: 223 SSTSASVAASTAPKAASPAVLPEGKPRRRKNNSQMRFCSTVLKELYKRQYESFAFPFYQP 282
Query: 292 SDP 294
DP
Sbjct: 283 VDP 285
>BDF1_YEAST (P35817) BDF1 protein
Length = 686
Score = 99.0 bits (245), Expect = 3e-20
Identities = 49/118 (41%), Positives = 75/118 (63%), Gaps = 4/118 (3%)
Query: 65 FEGQKEKRQKIDRKGSTQCAAILKCLMSYP---YSWVFNKPVDPVALNIPDYFAIIAKPM 121
+E +K K +++ ++ C ++LK LM+ Y++ F +PVDPV++N+P YF + +PM
Sbjct: 304 YESKKPKSKRL-QQAMKFCQSVLKELMAKKHASYNYPFLEPVDPVSMNLPTYFDYVKEPM 362
Query: 122 DLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKD 179
DLGTI+ KL Y +E+F DVRL F N T+NP V+MM L ++F+SKW D
Sbjct: 363 DLGTIAKKLNDWQYQTMEDFERDVRLVFKNCYTFNPDGTIVNMMGHRLEEVFNSKWAD 420
Score = 75.1 bits (183), Expect = 4e-13
Identities = 57/225 (25%), Positives = 92/225 (40%), Gaps = 23/225 (10%)
Query: 27 GQKCEFGQRGLQTDENGRCNSAEKLFK----------PDSIKRGPPEGFEGQKEKRQKID 76
G E Q+GL+ +E G+ E L + P PP + + I
Sbjct: 91 GSGAEDEQQGLKKEEGGQGTKQEDLDENSKQELPMEVPKEPAPAPPPEPDMNNLPQNPIP 150
Query: 77 RKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYS 136
+ +K + + F +PVDPV L+IP YF I +PMDL TI KL Y
Sbjct: 151 KHQQKHALLAIKAVKRLKDARPFLQPVDPVKLDIPFYFNYIKRPMDLSTIERKLNVGAYE 210
Query: 137 CVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTG 196
E+ D L +N++ +N P+ + MA+ + F+ +M K + K
Sbjct: 211 VPEQITEDFNLMVNNSIKFNGPNAGISQMARNIQASFEKHMLNMPAK-DAPPVIAKGRRS 269
Query: 197 TIKETA------------RKSCNVMHPRQKDTFPKKSQVSEHKRV 229
+ +E A R + P+ KD +P +S+ + KR+
Sbjct: 270 SAQEDAPIVIRRAQTHNGRPKRTIHPPKSKDIYPYESKKPKSKRL 314
>BRD3_MOUSE (Q8K2F0) Bromodomain-containing protein 3
(Bromodomain-containing FSH-like protein FSRG2)
Length = 726
Score = 98.6 bits (244), Expect = 3e-20
Identities = 99/401 (24%), Positives = 168/401 (41%), Gaps = 57/401 (14%)
Query: 61 PPEGFEGQKEKRQKIDRKGSTQ-----CAAILKCLMSYP---YSWVFNKPVDPVALNIPD 112
PP+ E Q +KG C +IL+ ++S Y+W F KPVD AL + D
Sbjct: 287 PPKKDLEDGEVPQHAGKKGKLSEHLRHCDSILREMLSKKHAAYAWPFYKPVDAEALELHD 346
Query: 113 YFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQI 172
Y II PMDL T+ K+ Y + FAAD+RL FSN YNPP ++V MA++L +
Sbjct: 347 YHDIIKHPMDLSTVKRKMDSREYPDAQGFAADIRLMFSNCYKYNPPDHEVVAMARKLQDV 406
Query: 173 FDSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQKDTFPKKSQVSEHKRVLKI 232
F+ ++ M + L + + A S ++ + S SE +R ++
Sbjct: 407 FEMRFAKMPDEPMEAPALPAPTAPIVSKGAESS----RSSEESSSDSGSSDSEEERATRL 462
Query: 233 SSL------------AARDAKVEVPKLSQV----------HCKSIEKDLIKGSEERDRIA 270
+ L A A V PK + K EK+ K E ++ A
Sbjct: 463 AELQEQLKAVHEQLAALSQAPVNKPKKKKEKKEKEKKKKDKDKDKEKEKHKAKSEEEKKA 522
Query: 271 PGAVAFKQKRQIKSTSPLERKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTP 330
A A KQ +Q K P ++ S + G + A+ + D+ E
Sbjct: 523 KAAPAAKQAQQ---------KKAPTKKANSTTTASRQLKKGGKQASASYDSEEEE----E 569
Query: 331 DFPVQLSPKKAL-----RAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQ-LEKERLER 384
P+ K+ L R K I+++++ +L + + + L+ L
Sbjct: 570 GLPMSYDEKRQLSLDINRLPGEKLGRVVHIIQSREPSLRDSNPDEIEIDFETLKPTTLRE 629
Query: 385 IQREERARIEAQ----IKTAEAAARARAEEELRQQREKERE 421
++R ++ ++ + + T+ A+++EEL Q+++KE E
Sbjct: 630 LERYVKSCLQKKQRKPLSTSGKKQAAKSKEELAQEKKKELE 670
Score = 83.2 bits (204), Expect = 1e-15
Identities = 44/130 (33%), Positives = 66/130 (49%), Gaps = 2/130 (1%)
Query: 54 PDSIKRGPPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDY 113
P + PPE K R+ + ++K L + ++W F +PVD + LN+PDY
Sbjct: 15 PGPVNPPPPEVSNPSKPGRKTNQLQYMQN--VVVKTLWKHQFAWPFYQPVDAIKLNLPDY 72
Query: 114 FAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIF 173
II PMD+GTI +L+ N Y E D F+N YN P +D+ +MA+ L +IF
Sbjct: 73 HKIIKNPMDMGTIKKRLENNYYWSASECMQDFNTMFTNCYIYNKPTDDIVLMAQALEKIF 132
Query: 174 DSKWKDMDKK 183
K M ++
Sbjct: 133 LQKVAQMPQE 142
>BRD3_HUMAN (Q15059) Bromodomain-containing protein 3 (RING3-like
protein)
Length = 726
Score = 95.1 bits (235), Expect = 4e-19
Identities = 102/396 (25%), Positives = 163/396 (40%), Gaps = 49/396 (12%)
Query: 61 PPEGFEGQKEKRQKIDRKGSTQ-----CAAILKCLMSYP---YSWVFNKPVDPVALNIPD 112
PP+ E Q +KG C +IL+ ++S Y+W F KPVD AL + D
Sbjct: 288 PPKKDLEDGEVPQHAGKKGKLSEHLRYCDSILREMLSKKHAAYAWPFYKPVDAEALELHD 347
Query: 113 YFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQI 172
Y II PMDL T+ K+ Y + FAADVRL FSN YNPP ++V MA++L +
Sbjct: 348 YHDIIKHPMDLSTVKRKMDGREYPDAQGFAADVRLMFSNCYKYNPPDHEVVAMARKLQDV 407
Query: 173 FDSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQKDTFPKKSQVSEHKRVLKI 232
F+ ++ M + L + + A S ++ + S SE +R ++
Sbjct: 408 FEMRFAKMPDEPVEAPALPAPAAPMVSKGAESS----RSSEESSSDSGSSDSEEERATRL 463
Query: 233 SSL------------AARDAKVEVPKLSQVHC----------KSIEKDLIKGSEERD-RI 269
+ L A A V PK + K EK +K EE+ ++
Sbjct: 464 AELQEQLKAVHEQLAALSQAPVNKPKKKKEKKEKEKKKKDKEKEKEKHKVKAEEEKKAKV 523
Query: 270 APGAVAFKQKR----QIKSTSPLERKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGE 325
AP A +QK+ + ST+ R+ A +S DSE G + S +
Sbjct: 524 APPAKQAQQKKAPAKKANSTTTAGRQLKKGGKQASASYDSEEEEEGLPMSYDEKRQLSLD 583
Query: 326 VWSTPDFPVQLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERI 385
+ P K +++SR + ++ +LE+ +
Sbjct: 584 INRLP------GEKLGRVVHIIQSREPSLRDSNPDEIEIDFETLKPTTLRELERYVKSCL 637
Query: 386 QREERARIEAQIKTAEAAARARAEEELRQQREKERE 421
Q+++R A K A +++EEL Q+++KE E
Sbjct: 638 QKKQRKPFSASGKKQAA----KSKEELAQEKKKELE 669
Score = 83.2 bits (204), Expect = 1e-15
Identities = 44/130 (33%), Positives = 66/130 (49%), Gaps = 2/130 (1%)
Query: 54 PDSIKRGPPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDY 113
P + PPE K R+ + ++K L + ++W F +PVD + LN+PDY
Sbjct: 16 PGPVNPPPPEVSNPSKPGRKTNQLQYMQN--VVVKTLWKHQFAWPFYQPVDAIKLNLPDY 73
Query: 114 FAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIF 173
II PMD+GTI +L+ N Y E D F+N YN P +D+ +MA+ L +IF
Sbjct: 74 HKIIKNPMDMGTIKKRLENNYYWSASECMQDFNTMFTNCYIYNKPTDDIVLMAQALEKIF 133
Query: 174 DSKWKDMDKK 183
K M ++
Sbjct: 134 LQKVAQMPQE 143
>BRD2_HUMAN (P25440) Bromodomain-containing protein 2 (RING3
protein) (O27.1.1)
Length = 801
Score = 94.0 bits (232), Expect = 8e-19
Identities = 52/123 (42%), Positives = 70/123 (56%), Gaps = 4/123 (3%)
Query: 61 PPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYP---YSWVFNKPVDPVALNIPDYFAII 117
P + Q K+ K+ + C ILK L+S Y+W F KPVD AL + DY II
Sbjct: 332 PDSQQQHQSSKKGKLSEQ-LKHCNGILKELLSKKHAAYAWPFYKPVDASALGLHDYHDII 390
Query: 118 AKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKW 177
PMDL T+ K++ Y +EFAADVRL FSN YNPP +DV MA++L +F+ ++
Sbjct: 391 KHPMDLSTVKRKMENRDYRDAQEFAADVRLMFSNCYKYNPPDHDVVAMARKLQDVFEFRY 450
Query: 178 KDM 180
M
Sbjct: 451 AKM 453
Score = 82.8 bits (203), Expect = 2e-15
Identities = 38/98 (38%), Positives = 57/98 (57%)
Query: 86 ILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADV 145
++K L + ++W F +PVD V L +PDY II +PMD+GTI +L+ N Y E D
Sbjct: 86 VMKALWKHQFAWPFRQPVDAVKLGLPDYHKIIKQPMDMGTIKRRLENNYYWAASECMQDF 145
Query: 146 RLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
F+N YN P +D+ +MA+ L +IF K M ++
Sbjct: 146 NTMFTNCYIYNKPTDDIVLMAQTLEKIFLQKVASMPQE 183
>BRD4_HUMAN (O60885) Bromodomain-containing protein 4 (HUNK1
protein)
Length = 1362
Score = 93.6 bits (231), Expect = 1e-18
Identities = 82/280 (29%), Positives = 120/280 (42%), Gaps = 31/280 (11%)
Query: 38 QTDENGRCNSAEKLFKPDSIKRGPPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYP--- 94
Q E+ R K PDS + PE K K+ + C+ ILK + +
Sbjct: 320 QRRESSRPVKPPKKDVPDSQQHPAPE-------KSSKVSEQLKC-CSGILKEMFAKKHAA 371
Query: 95 YSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMT 154
Y+W F KPVD AL + DY II PMD+ TI SKL+ Y +EF ADVRL FSN
Sbjct: 372 YAWPFYKPVDVEALGLHDYCDIIKHPMDMSTIKSKLEAREYRDAQEFGADVRLMFSNCYK 431
Query: 155 YNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQK 214
YNPP ++V MA++L +F+ ++ K DE + V V+ P
Sbjct: 432 YNPPDHEVVAMARKLQDVFEMRFA------KMPDEPEEPVVAVSSPAVPPPTKVVAPPSS 485
Query: 215 DTFPKKSQV--------SEHKRVLKISSL-----AARDAKVEVPKLSQVHCKSIEKDLIK 261
S SE +R +++ L A + + + Q K EKD +
Sbjct: 486 SDSSSDSSSDSDSSTDDSEEERAQRLAELQEQLKAVHEQLAALSQPQQNKPKKKEKDKKE 545
Query: 262 GSEERDRIAPGAVAFKQKRQIKSTSPLERKSDPDSDGAVS 301
+E+ + V +K + K P + K + S+ VS
Sbjct: 546 KKKEKHK-RKEEVEENKKSKAKEPPPKKTKKNNSSNSNVS 584
Score = 89.0 bits (219), Expect = 3e-17
Identities = 52/149 (34%), Positives = 77/149 (50%), Gaps = 15/149 (10%)
Query: 61 PPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKP 120
PPE K KRQ + + +LK L + ++W F +PVD V LN+PDY+ II P
Sbjct: 47 PPETSNPNKPKRQTNQLQYLLR--VVLKTLWKHQFAWPFQQPVDAVKLNLPDYYKIIKTP 104
Query: 121 MDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDM 180
MD+GTI +L+ N Y +E D F+N YN P +D+ +MA+ L ++F K ++
Sbjct: 105 MDMGTIKKRLENNYYWNAQECIQDFNTMFTNCYIYNKPGDDIVLMAEALEKLFLQKINEL 164
Query: 181 DKKWKCEDEL----------GKSVTGTIK 199
+ E E+ G+ TGT K
Sbjct: 165 PTE---ETEIMIVQAKGRGRGRKETGTAK 190
Score = 35.8 bits (81), Expect = 0.27
Identities = 45/190 (23%), Positives = 88/190 (45%), Gaps = 29/190 (15%)
Query: 240 AKVEVPKLSQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTSPLERKSDPD--SD 297
A V +P+ ++ + + +I+ E+ P K K++ + +P+ K D +
Sbjct: 1142 APVHLPQRPEMKPVDVGRPVIRPPEQN--APPPGAPDKDKQKQEPKTPVAPKKDLKIKNM 1199
Query: 298 GAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKALRAAMLKSRFADTILK 357
G+ +SL +H TP++ A S S+ F + R A + + LK
Sbjct: 1200 GSWASLVQKHP------TTPSSTAKS----SSDSF-------EQFRRAAREKEEREKALK 1242
Query: 358 AQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELRQQRE 417
AQ EH +K K +L +ER+ R + +E A +A+ EA R +++ RQ+++
Sbjct: 1243 AQA----EHAEKE---KERLRQERM-RSREDEDALEQARRAHEEARRRQEQQQQQRQEQQ 1294
Query: 418 KEREAARAAI 427
++++ AA+
Sbjct: 1295 QQQQQQAAAV 1304
>BRD4_MOUSE (Q9ESU6) Bromodomain-containing protein 4 (Mitotic
chromosome-associated protein) (MCAP)
Length = 1400
Score = 93.2 bits (230), Expect = 1e-18
Identities = 80/271 (29%), Positives = 120/271 (43%), Gaps = 19/271 (7%)
Query: 41 ENGRCNSAEKLFKPDSIKRGPPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYP---YSW 97
E+ R K PDS + PE K KI + C+ ILK + + Y+W
Sbjct: 324 ESSRPVKPPKKDVPDSQQHPGPE-------KSSKISEQLKC-CSGILKEMFAKKHAAYAW 375
Query: 98 VFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNP 157
F KPVD AL + DY II PMD+ TI SKL+ Y +EF ADVRL FSN YNP
Sbjct: 376 PFYKPVDVEALGLHDYCDIIKHPMDMSTIKSKLESREYRDAQEFGADVRLMFSNCYKYNP 435
Query: 158 PHNDVHMMAKELNQIFDSKWKDM--DKKWKCEDELGKSVTGTIKETARKSCNVMHPRQKD 215
P ++V MA++L +F+ ++ M + + +V K A S +
Sbjct: 436 PDHEVVAMARKLQDVFEMRFAKMPDEPEEPVVTVSSPAVPPPTKVVAPPSSSDSSSDSSS 495
Query: 216 TFPKKSQVSEHKRVLKISSL-----AARDAKVEVPKLSQVHCKSIEKDLIKGSEERDRIA 270
+ SE +R +++ L A + + + Q K EKD + +E+ +
Sbjct: 496 DSDSSTDDSEEERAQRLAELQEQLKAVHEQLAALSQPQQNKPKKKEKDKKEKKKEKHK-K 554
Query: 271 PGAVAFKQKRQIKSTSPLERKSDPDSDGAVS 301
V +K + K P + K + S+ VS
Sbjct: 555 KEEVEENKKSKTKELPPKKTKKNNSSNSNVS 585
Score = 88.6 bits (218), Expect = 4e-17
Identities = 45/120 (37%), Positives = 67/120 (55%), Gaps = 2/120 (1%)
Query: 61 PPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKP 120
PPE K KRQ + + +LK L + ++W F +PVD V LN+PDY+ II P
Sbjct: 47 PPETSNPNKPKRQTNQLQYLLR--VVLKTLWKHQFAWPFQQPVDAVKLNLPDYYKIIKTP 104
Query: 121 MDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDM 180
MD+GTI +L+ N Y +E D F+N YN P +D+ +MA+ L ++F K ++
Sbjct: 105 MDMGTIKKRLENNYYWNAQECIQDFNTMFTNCYIYNKPGDDIVLMAEALEKLFLQKINEL 164
Score = 35.8 bits (81), Expect = 0.27
Identities = 48/195 (24%), Positives = 84/195 (42%), Gaps = 37/195 (18%)
Query: 240 AKVEVPKLSQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTSPLERKSDPD--SD 297
A V +P+ ++ I + +I+ E+ P K K++ + +P+ K D +
Sbjct: 1178 APVHLPQRPEMKPVDIGRPVIRPPEQS--APPPGAPDKDKQKQEPKTPVAPKKDLKIKNM 1235
Query: 298 GAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKALRAAMLKSRFADTILK 357
G+ +SL +H TP++ A S S+ F + R A + + LK
Sbjct: 1236 GSWASLVQKHP------TTPSSTAKS----SSDSF-------EHFRRAAREKEEREKALK 1278
Query: 358 AQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELRQQRE 417
AQ EH +K EKERL R+ER R E A RA E RQ+++
Sbjct: 1279 AQA----EHAEK--------EKERL----RQERMRSREDEDALEQARRAHEEARRRQEQQ 1322
Query: 418 KEREAARAAIEEMKR 432
++++ R ++ ++
Sbjct: 1323 QQQQQQRQEQQQQQQ 1337
>FSH_DROME (P13709) Female sterile homeotic protein (Fragile-chorion
membrane protein)
Length = 2038
Score = 90.5 bits (223), Expect = 9e-18
Identities = 46/101 (45%), Positives = 60/101 (58%), Gaps = 3/101 (2%)
Query: 83 CAAILKCLMSYP---YSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVE 139
C ILK L S Y+W F KPVD L + DY II KPMDLGT+ K+ Y
Sbjct: 484 CNEILKELFSKKHSGYAWPFYKPVDAEMLGLHDYHDIIKKPMDLGTVKRKMDNREYKSAP 543
Query: 140 EFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDM 180
EFAADVRL F+N YNPP +DV M ++L +F+ ++ ++
Sbjct: 544 EFAADVRLIFTNCYKYNPPDHDVVAMGRKLQDVFEMRYANI 584
Score = 84.7 bits (208), Expect = 5e-16
Identities = 39/98 (39%), Positives = 58/98 (58%)
Query: 86 ILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADV 145
++K + + +SW F +PVD LN+PDY II +PMD+GTI +L+ N Y +E D
Sbjct: 46 VMKVIWKHHFSWPFQQPVDAKKLNLPDYHKIIKQPMDMGTIKKRLENNYYWSAKETIQDF 105
Query: 146 RLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKK 183
F+N YN P DV +MA+ L ++F K + M K+
Sbjct: 106 NTMFNNCYVYNKPGEDVVVMAQTLEKVFLQKIESMPKE 143
>YK82_SCHPO (Q9HGP4) Hypothetical bromodomain protein C631.02
Length = 727
Score = 84.3 bits (207), Expect = 7e-16
Identities = 41/98 (41%), Positives = 57/98 (57%), Gaps = 3/98 (3%)
Query: 83 CAAILKCLMSYP---YSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVE 139
C ++LK L+ Y++ F KPV+P A PDYF +I PMDLGT+ +KL N Y+ ++
Sbjct: 397 CQSVLKELLKKQHEAYAYPFYKPVNPTACGCPDYFKVIKHPMDLGTMQNKLNHNEYASMK 456
Query: 140 EFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKW 177
F AD+ L F N +N VH+M K+L IF W
Sbjct: 457 AFEADMVLMFKNCYKFNSAGTPVHLMGKKLESIFQKLW 494
Score = 75.9 bits (185), Expect = 2e-13
Identities = 42/115 (36%), Positives = 58/115 (49%)
Query: 66 EGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGT 125
E K++ + ++ A+L+ L S F PVDPV NIPDY II P+DLGT
Sbjct: 221 ENDKDQYPPMTKEQHKYIHAMLRQLRRGRDSIPFRAPVDPVKQNIPDYPTIIKNPIDLGT 280
Query: 126 ISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDM 180
+ K VYS + F D+ L FSN YN + V +M K L F+ + K +
Sbjct: 281 MQKKFSSGVYSSAQHFIDDMNLMFSNCFLYNGTESPVGVMGKNLQATFERQLKQL 335
>BA2B_CHICK (Q9DE13) Bromodomain adjacent to zinc finger domain 2B
(Extracellular matrix protein F22)
Length = 2130
Score = 75.9 bits (185), Expect = 2e-13
Identities = 45/133 (33%), Positives = 67/133 (49%), Gaps = 3/133 (2%)
Query: 50 KLFKPDSIKRGPPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALN 109
K+ + S+ +G E F K K ++ D K C+ IL L ++ +W F PV+
Sbjct: 1999 KMDESVSVSQGKQENFTAIK-KPKRDDSKDLAICSMILSELETHEDAWPFLLPVNLKL-- 2055
Query: 110 IPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKEL 169
+P Y +I KPMD TI KL Y VE F+ DVRL F N T+N +D+ +
Sbjct: 2056 VPGYKKVIKKPMDFSTIRDKLTSGQYPNVEAFSLDVRLVFDNCETFNEDDSDIGRAGHNM 2115
Query: 170 NQIFDSKWKDMDK 182
+ F+ KW ++ K
Sbjct: 2116 RKYFEKKWTEIFK 2128
Score = 42.0 bits (97), Expect = 0.004
Identities = 35/123 (28%), Positives = 63/123 (50%), Gaps = 14/123 (11%)
Query: 339 KKALRAAMLKS--RFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQ 396
++A A +L++ R + ++ QQ LL+H +LE+ RL+ ++RE R +
Sbjct: 887 EEAANAKLLEAEKRIKEKEMRRQQAVLLKH--------QELERHRLD-MERERRRQHMML 937
Query: 397 IKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSY 456
+K EA +A +E L+Q++ E+ + E +R +E+E E+ K E + C
Sbjct: 938 MKAMEARKKAEEKERLKQEKRDEKRLNKERKLEQRR-LELEMAKELKKPNEDM--CLADQ 994
Query: 457 KAL 459
KAL
Sbjct: 995 KAL 997
Score = 33.9 bits (76), Expect = 1.0
Identities = 41/166 (24%), Positives = 72/166 (42%), Gaps = 25/166 (15%)
Query: 290 RKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKALRAAMLKS 349
RK P + G+ LDS TDA + Q + K LR +
Sbjct: 779 RKGRPPNVGSTEFLDS-------------TDAKLLRKLQAQEIARQAAQIKLLRKLQKQE 825
Query: 350 RFADTILKAQQKTLLEHGDKG---DPLKMQLEKERLERIQ-----REERAR--IEAQIKT 399
+ +Q+ ++ +K + +K+ ++E+++RIQ +E RA+ +EA+ K
Sbjct: 826 QARAAKEAKKQQAIMAAEEKRKQKEQIKIMKQQEKIKRIQQIRMEKELRAQQILEAKKKK 885
Query: 400 AEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKE 445
E AA A+ E ++R KE+E R +K H L++ +E
Sbjct: 886 KEEAANAKLLEA--EKRIKEKEMRRQQAVLLKHQELERHRLDMERE 929
>BA2B_HUMAN (Q9UIF8) Bromodomain adjacent to zinc finger domain 2B
(hWALp4)
Length = 1972
Score = 73.9 bits (180), Expect = 9e-13
Identities = 45/133 (33%), Positives = 64/133 (47%), Gaps = 3/133 (2%)
Query: 50 KLFKPDSIKRGPPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALN 109
K+ + SI E F K K ++ D K C+ IL + ++ +W F PV+
Sbjct: 1841 KMEENTSINLSKQESFTSVK-KPKRDDSKDLALCSMILTEMETHEDAWPFLLPVNLKL-- 1897
Query: 110 IPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKEL 169
+P Y +I KPMD TI KL Y +E FA DVRL F N T+N +D+ +
Sbjct: 1898 VPGYKKVIKKPMDFSTIREKLSSGQYPNLETFALDVRLVFDNCETFNEDDSDIGRAGHNM 1957
Query: 170 NQIFDSKWKDMDK 182
+ F+ KW D K
Sbjct: 1958 RKYFEKKWTDTFK 1970
Score = 33.5 bits (75), Expect = 1.3
Identities = 30/114 (26%), Positives = 51/114 (44%), Gaps = 6/114 (5%)
Query: 335 QLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIE 394
Q+ +K LRA + +A LLE + +M+ ++ L + Q ER R
Sbjct: 756 QIRMEKELRAQQILEAKKKKKEEAANAKLLEAEKRIKEKEMRRQQAVLLKHQERERRRQH 815
Query: 395 AQI-KTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELE 447
+ K EA +A +E L+Q++ E+ + E +R LE+ KEL+
Sbjct: 816 MMLMKAMEARKKAEEKERLKQEKRDEKRLNKERKLEQRRL-----ELEMAKELK 864
>BA2A_MOUSE (Q91YE5) Bromodomain adjacent to zinc finger domain 2A
(Transcription termination factor-I interacting protein
5) (TTF-I interacting protein 5) (Tip5)
Length = 1850
Score = 70.9 bits (172), Expect = 8e-12
Identities = 39/126 (30%), Positives = 65/126 (50%), Gaps = 4/126 (3%)
Query: 56 SIKRGPPEGFEGQKEKRQKIDRKGS--TQCAAILKCLMSYPYSWVFNKPVDPVALNIPDY 113
++ R P +G K +R + S T C IL + S+ +W F +PV+P ++ Y
Sbjct: 1718 AVPRYPEDGLSPPKRRRHSMRSHHSDLTFCEIILMEMESHDAAWPFLEPVNPRLVS--GY 1775
Query: 114 FAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIF 173
+I PMD T+ +L + Y+ EEFAAD L F N T+N ++V + + F
Sbjct: 1776 RRVIKNPMDFSTMRERLLRGGYTSSEEFAADALLVFDNCQTFNEDDSEVGKAGHVMRRFF 1835
Query: 174 DSKWKD 179
+S+W++
Sbjct: 1836 ESRWEE 1841
Score = 38.5 bits (88), Expect = 0.042
Identities = 28/95 (29%), Positives = 57/95 (59%), Gaps = 5/95 (5%)
Query: 369 KGDPLKMQLEKERL-ERIQREERARIEAQIKTA-EAAARARAEEELR-QQREKEREAARA 425
K + KM+ ++E+L E+++RE++ +++A+ K A RA++ L Q+R +E++ +A
Sbjct: 708 KAEKEKMKTKQEKLKEKVKREKKEKVKAKGKEGPRARPSCRADKTLATQKRLEEQQRQQA 767
Query: 426 AIEEMKRTVE--IEHNLEIVKELETLSGCTLSYKA 458
+EEMK+ E + + + + + G TLS +A
Sbjct: 768 ILEEMKKPTEGMCLSDHQPLPDFTRIPGLTLSSRA 802
>CBP_MOUSE (P45481) CREB-binding protein (EC 2.3.1.48)
Length = 2441
Score = 69.7 bits (169), Expect = 2e-11
Identities = 35/83 (42%), Positives = 47/83 (56%)
Query: 94 PYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAM 153
P S F +PVDP L IPDYF I+ PMDL TI KL Y ++ DVRL F+NA
Sbjct: 1107 PESLPFRQPVDPQLLGIPDYFDIVKNPMDLSTIKRKLDTGQYQEPWQYVDDVRLMFNNAW 1166
Query: 154 TYNPPHNDVHMMAKELNQIFDSK 176
YN + V+ +L ++F+ +
Sbjct: 1167 LYNRKTSRVYKFCSKLAEVFEQE 1189
>WDR9_HUMAN (Q9NSI6) WD-repeat protein 9
Length = 2269
Score = 69.3 bits (168), Expect = 2e-11
Identities = 59/211 (27%), Positives = 90/211 (41%), Gaps = 14/211 (6%)
Query: 16 KFSTKRIEVDPGQKCEFGQRGLQTDENGRCNSAEKLFKPDSIKR--GPPEGFEGQKEKRQ 73
K + + ++ Q C T EN N AE L D K G +G+K R
Sbjct: 1252 KITDQLLKFIKNQHCTNISELSNTSENDEQN-AEDLDDSDLPKTSSGRRRVHDGKKSIRA 1310
Query: 74 K--IDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLK 131
++ QC ++ + S F +PVD V PDY II PMD GT+ L
Sbjct: 1311 TNYVESNWKKQCKELVNLIFQCEDSEPFRQPVDLV--EYPDYRDIIDTPMDFGTVRETLD 1368
Query: 132 KNVYSCVEEFAADVRLTFSNAMTYNP-PHNDVHMMAKELNQIFDSKWKDMDKKWKCEDEL 190
Y EF D+RL FSNA Y P + ++ M L+ +F+ K K + +K +
Sbjct: 1369 AGNYDSPLEFCKDIRLIFSNAKAYTPNKRSKIYSMTLRLSALFEEKMKKISSDFKIGQKF 1428
Query: 191 GKSVTGTIKETARKSCNVMHPRQKDTFPKKS 221
+ + + + R++C + D+ P KS
Sbjct: 1429 NEKLRRSQRFKQRQNC------KGDSQPNKS 1453
Score = 43.5 bits (101), Expect = 0.001
Identities = 32/109 (29%), Positives = 49/109 (44%), Gaps = 7/109 (6%)
Query: 61 PPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKP 120
P G GQK + ++ DR S + L++ + F PVD P Y ++A P
Sbjct: 1152 PQAGEWGQKSRDEECDRIISG-----IDQLLNLDIAAAFAGPVD--LCTYPKYCTVVAYP 1204
Query: 121 MDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKEL 169
DL TI +L Y + +VR NA T+N P + + AK++
Sbjct: 1205 TDLYTIRMRLVNRFYRRLSALVWEVRYIEHNARTFNEPESVIARSAKKI 1253
>TF1A_HUMAN (O15164) Transcription intermediary factor 1-alpha
(TIF1-alpha) (Tripartite motif-containing protein 24)
Length = 1050
Score = 68.6 bits (166), Expect = 4e-11
Identities = 44/141 (31%), Positives = 67/141 (47%), Gaps = 8/141 (5%)
Query: 45 CNSAEKLFKPDSIKRGPPEGFEGQKEKRQ---KIDRKGSTQCAAILKCLMSYPYSWVFNK 101
C L KP+ +K+K + K+ +C +L L + S F
Sbjct: 867 CTFCRDLSKPEVEYDCDAPSHNSEKKKTEGLVKLTPIDKRKCERLLLFLYCHEMSLAFQ- 925
Query: 102 PVDPVALNIPDYFAIIAKPMDLGTISSKLKK--NVYSCVEEFAADVRLTFSNAMTYNPPH 159
DPV L +PDY+ II PMDL TI +L++ ++YS E+F AD RL F N +N P
Sbjct: 926 --DPVPLTVPDYYKIIKNPMDLSTIKKRLQEDYSMYSKPEDFVADFRLIFQNCAEFNEPD 983
Query: 160 NDVHMMAKELNQIFDSKWKDM 180
++V +L F+ K++
Sbjct: 984 SEVANAGIKLENYFEELLKNL 1004
>CBP_HUMAN (Q92793) CREB-binding protein (EC 2.3.1.48)
Length = 2442
Score = 68.2 bits (165), Expect = 5e-11
Identities = 54/197 (27%), Positives = 85/197 (42%), Gaps = 28/197 (14%)
Query: 8 VPTRKLKIKFSTKRIEVDPGQ-----KCEFGQRGLQ--TDENGRCNSAEKLFKPDSIKRG 60
VP ++K + + E DPG+ + E + LQ + + AE+ +P +
Sbjct: 992 VPVLEMKTETQAEDTEPDPGESKGEPRSEMMEEDLQGASQVKEETDIAEQKSEPMEVDEK 1051
Query: 61 PPEGFEGQKEKRQKIDRKGSTQCAA-------------ILKCLMSY--------PYSWVF 99
PE KE+ + ++Q + + + LM P S F
Sbjct: 1052 KPEVKVEVKEEEESSSNGTASQSTSPSQPRKKIFKPEELRQALMPTLEALYRQDPESLPF 1111
Query: 100 NKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPH 159
+PVDP L IPDYF I+ PMDL TI KL Y ++ DV L F+NA YN
Sbjct: 1112 RQPVDPQLLGIPDYFDIVKNPMDLSTIKRKLDTGQYQEPWQYVDDVWLMFNNAWLYNRKT 1171
Query: 160 NDVHMMAKELNQIFDSK 176
+ V+ +L ++F+ +
Sbjct: 1172 SRVYKFCSKLAEVFEQE 1188
>BA2A_HUMAN (Q9UIF9) Bromodomain adjacent to zinc finger domain 2A
(Transcription termination factor-I interacting protein
5) (TTF-I interacting protein 5) (Tip5) (hWALp3)
Length = 1878
Score = 68.2 bits (165), Expect = 5e-11
Identities = 41/128 (32%), Positives = 64/128 (49%), Gaps = 4/128 (3%)
Query: 54 PDSIKRGPPEGFEGQKEKRQKIDRKGS--TQCAAILKCLMSYPYSWVFNKPVDPVALNIP 111
P + R EG K +R + S T C IL + S+ +W F +PV+P ++
Sbjct: 1744 PAAGPRYSEEGLSPSKRRRLSMRNHHSDLTFCEIILMEMESHDAAWPFLEPVNPRLVS-- 1801
Query: 112 DYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQ 171
Y II PMD T+ +L + Y+ EEFAAD L F N T+N ++V + +
Sbjct: 1802 GYRRIIKNPMDFSTMRERLLRGGYTSSEEFAADALLVFDNCQTFNEDDSEVGKAGHIMRR 1861
Query: 172 IFDSKWKD 179
F+S+W++
Sbjct: 1862 FFESRWEE 1869
Score = 33.1 bits (74), Expect = 1.7
Identities = 22/81 (27%), Positives = 44/81 (54%), Gaps = 8/81 (9%)
Query: 357 KAQQKTLLEHGD-KGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELRQQ 415
K + K E G K + LK ++++E+ E+++ +E+ + +A +A++ L Q
Sbjct: 724 KKKSKAEKEKGKTKQEKLKEKVKREKKEKVKMKEKEEV------TKAKPACKADKTLATQ 777
Query: 416 RE-KEREAARAAIEEMKRTVE 435
R +ER+ + +EEMK+ E
Sbjct: 778 RRLEERQKQQMILEEMKKPTE 798
>GCN5_YEAST (Q03330) Histone acetyltransferase GCN5 (EC 2.3.1.48)
Length = 439
Score = 67.4 bits (163), Expect = 8e-11
Identities = 34/95 (35%), Positives = 54/95 (56%), Gaps = 2/95 (2%)
Query: 86 ILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADV 145
IL L ++ +W F +PV+ +PDY+ I +PMDL T+ KL+ N Y +E+F D
Sbjct: 339 ILTELQNHAAAWPFLQPVNKE--EVPDYYDFIKEPMDLSTMEIKLESNKYQKMEDFIYDA 396
Query: 146 RLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDM 180
RL F+N YN + + A L + F++K K++
Sbjct: 397 RLVFNNCRMYNGENTSYYKYANRLEKFFNNKVKEI 431
>P300_HUMAN (Q09472) E1A-associated protein p300 (EC 2.3.1.48)
Length = 2414
Score = 67.0 bits (162), Expect = 1e-10
Identities = 33/83 (39%), Positives = 47/83 (55%)
Query: 94 PYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAM 153
P S F +PVDP L IPDYF I+ PMDL TI KL Y ++ D+ L F+NA
Sbjct: 1070 PESLPFRQPVDPQLLGIPDYFDIVKSPMDLSTIKRKLDTGQYQEPWQYVDDIWLMFNNAW 1129
Query: 154 TYNPPHNDVHMMAKELNQIFDSK 176
YN + V+ +L+++F+ +
Sbjct: 1130 LYNRKTSRVYKYCSKLSEVFEQE 1152
>TF1A_MOUSE (Q64127) Transcription intermediary factor 1-alpha
(TIF1-alpha) (Tripartite motif-containing protein 24)
Length = 1051
Score = 66.6 bits (161), Expect = 1e-10
Identities = 43/141 (30%), Positives = 66/141 (46%), Gaps = 8/141 (5%)
Query: 45 CNSAEKLFKPDSIKRGPPEGFEGQKEKRQ---KIDRKGSTQCAAILKCLMSYPYSWVFNK 101
C L KP+ +K K + K+ +C +L L + S F
Sbjct: 868 CTFCRDLSKPEVDYDCDVPSHHSEKRKSEGLTKLTPIDKRKCERLLLFLYCHEMSLAFQ- 926
Query: 102 PVDPVALNIPDYFAIIAKPMDLGTISSKLKKN--VYSCVEEFAADVRLTFSNAMTYNPPH 159
DPV L +PDY+ II PMDL TI +L+++ +Y+ E+F AD RL F N +N P
Sbjct: 927 --DPVPLTVPDYYKIIKNPMDLSTIKKRLQEDYCMYTKPEDFVADFRLIFQNCAEFNEPD 984
Query: 160 NDVHMMAKELNQIFDSKWKDM 180
++V +L F+ K++
Sbjct: 985 SEVANAGIKLESYFEELLKNL 1005
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.130 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 59,711,646
Number of Sequences: 164201
Number of extensions: 2541941
Number of successful extensions: 13054
Number of sequences better than 10.0: 568
Number of HSP's better than 10.0 without gapping: 165
Number of HSP's successfully gapped in prelim test: 420
Number of HSP's that attempted gapping in prelim test: 10286
Number of HSP's gapped (non-prelim): 1981
length of query: 510
length of database: 59,974,054
effective HSP length: 115
effective length of query: 395
effective length of database: 41,090,939
effective search space: 16230920905
effective search space used: 16230920905
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0228a.2


