

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0227.11
(425 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
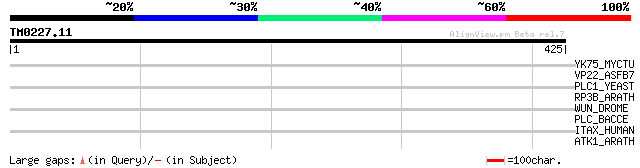
Score E
Sequences producing significant alignments: (bits) Value
YK75_MYCTU (Q10683) Hypothetical protein Rv2075c/MT2135 42 0.002
VP22_ASFB7 (P23169) Protein P22 precursor (Protein K'177) 34 0.82
PLC1_YEAST (P32383) 1-phosphatidylinositol-4,5-bisphosphate phos... 33 1.1
RP3B_ARATH (Q39212) DNA-directed RNA polymerase II 36 kDa polype... 32 2.4
WUN_DROME (Q9V576) Putative phosphatidate phosphatase (EC 3.1.3.... 32 4.0
PLC_BACCE (P14262) 1-phosphatidylinositol phosphodiesterase prec... 30 9.0
ITAX_HUMAN (P20702) Integrin alpha-X precursor (Leukocyte adhesi... 30 9.0
ATK1_ARATH (Q07970) Kinesin 1 (Kinesin-like protein A) 30 9.0
>YK75_MYCTU (Q10683) Hypothetical protein Rv2075c/MT2135
Length = 487
Score = 42.4 bits (98), Expect = 0.002
Identities = 37/126 (29%), Positives = 59/126 (46%), Gaps = 14/126 (11%)
Query: 79 LPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFEN-- 136
+P WL THNSF L S + +V + +NQQ ++ QL+ VR L LDL+
Sbjct: 112 VPLRETQWLGTHNSFNSL---SDSFTVSHADSNQQLSLAQQLDIDVRALELDLHYLPRLE 168
Query: 137 -----DVWLCHSFGGQCYNY-TAFQPAI-NVLKEIQVFLEA--NPSEIVTIIIEDYVTSP 187
V +CH G + N +P + VL +I +L A + E++ + +ED + +
Sbjct: 169 GHGAPGVTVCHGLGPKNANLGCTVEPLLATVLPQIANWLNAPGHTEEVILLYLEDQLKNA 228
Query: 188 KGLTKV 193
V
Sbjct: 229 SAYESV 234
>VP22_ASFB7 (P23169) Protein P22 precursor (Protein K'177)
Length = 177
Score = 33.9 bits (76), Expect = 0.82
Identities = 19/52 (36%), Positives = 29/52 (55%), Gaps = 5/52 (9%)
Query: 7 TLAIATTLFTILVLLDSSLALKFKPLQQEGQICVANKNCNSGLHC--ETCVT 56
TL IA + I++L+ + L +K Q ++C +K+C SG HC TC T
Sbjct: 10 TLLIALIILVIIILV---VFLYYKKQQPPKKVCKVDKDCGSGEHCVRGTCST 58
>PLC1_YEAST (P32383) 1-phosphatidylinositol-4,5-bisphosphate
phosphodiesterase 1 (EC 3.1.4.11) (Phosphoinositide
phospholipase C) (PLC-1) (Phospholipase C-1)
Length = 869
Score = 33.5 bits (75), Expect = 1.1
Identities = 38/168 (22%), Positives = 72/168 (42%), Gaps = 24/168 (14%)
Query: 80 PFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVW 139
P N Y ++HN++ L Q + T SV + L G R + +D++D EN
Sbjct: 385 PLNHYFIASSHNTYLLGKQIAETPSV--------EGYIQVLQQGCRCVEIDIWDGENGPV 436
Query: 140 LCHSFGGQCYNYTAFQPAINVLKEIQ--VFLEANPSEIVTIIIEDYVTSPKGLTKVFDAA 197
+CH F T+ P V++ I+ F+ + I+++ I + K + +
Sbjct: 437 VCHGF------LTSAIPLKTVIRVIKKYAFITSPYPLIISLEINCNKDNQKLASLIMREV 490
Query: 198 GLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEGIA 245
+ +F +R K P ++ K ++ SK + EA+ G++
Sbjct: 491 LAEQLYFVGTRTDK----LPSPRELKHK----ILLKSKKTSEATRGLS 530
>RP3B_ARATH (Q39212) DNA-directed RNA polymerase II 36 kDa
polypeptide B (EC 2.7.7.6) (RNA polymerase II subunit 3)
Length = 319
Score = 32.3 bits (72), Expect = 2.4
Identities = 24/83 (28%), Positives = 37/83 (43%), Gaps = 8/83 (9%)
Query: 1 MMMQKPTLAIATTLFTI--LVLLDSSLA--LKFKPLQQEG----QICVANKNCNSGLHCE 52
M+ + PT+AI + VL D +A L PL E + C ++CN HCE
Sbjct: 41 MISEVPTMAIHLVKIEVNSSVLNDEFIAQRLSLIPLTSERAMSMRFCQDCEDCNGDEHCE 100
Query: 53 TCVTNGNVRPRCTRTQPINPTSK 75
C + +C Q ++ TS+
Sbjct: 101 FCSVEFPLSAKCVTDQTLDVTSR 123
>WUN_DROME (Q9V576) Putative phosphatidate phosphatase (EC 3.1.3.4)
(Phosphatidic acid phosphatase type 2) (Wunen protein)
(Germ cell guidance factor)
Length = 379
Score = 31.6 bits (70), Expect = 4.0
Identities = 24/82 (29%), Positives = 41/82 (49%), Gaps = 9/82 (10%)
Query: 268 NRAESPSMNTTSRSLVLVNFFRDLPDVAQSCKDN-----SAPLLSMVST--CNQAAGKRW 320
N+A+ + N TSR V +N+ +LPD C +LS ++T + G+
Sbjct: 161 NKAKQDNGNATSRRYVFMNY--ELPDWMIECYKKIGIYAFGAVLSQLTTDIAKYSIGRLR 218
Query: 321 PNFIAVDFYKRSDGGGAPEAVD 342
P+FIAV + +DG +A++
Sbjct: 219 PHFIAVCQPQMADGSTCDDAIN 240
>PLC_BACCE (P14262) 1-phosphatidylinositol phosphodiesterase
precursor (EC 4.6.1.13) (Phosphatidylinositol
diacylglycerol-lyase) (Phosphatidylinositol-specific
phospholipase C) (PI-PLC)
Length = 329
Score = 30.4 bits (67), Expect = 9.0
Identities = 23/80 (28%), Positives = 36/80 (44%), Gaps = 10/80 (12%)
Query: 124 VRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDY 183
+RG + D +N + L H G Y Y IN K+ FL+ NPSE + + ++
Sbjct: 99 IRGRLTD----DNTIVLHH---GPLYLYVTLHEFINEAKQ---FLKDNPSETIIMSLKKE 148
Query: 184 VTSPKGLTKVFDAAGLRKYW 203
KG F + +KY+
Sbjct: 149 YEDMKGAEDSFSSTFEKKYF 168
>ITAX_HUMAN (P20702) Integrin alpha-X precursor (Leukocyte adhesion
glycoprotein p150,95 alpha chain) (Leukocyte adhesion
receptor p150,95) (CD11c) (Leu M5)
Length = 1163
Score = 30.4 bits (67), Expect = 9.0
Identities = 23/99 (23%), Positives = 42/99 (42%), Gaps = 12/99 (12%)
Query: 142 HSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIE--------DYVTSPKGLTKV 193
H G Y TA Q ++ L + ++I+ +I + DY K + +
Sbjct: 220 HQLQGFTYTATAIQNVVHRLFHASYGARRDATKILIVITDGKKEGDSLDY----KDVIPM 275
Query: 194 FDAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVF 232
DAAG+ +Y V +N W +++D+ K + +F
Sbjct: 276 ADAAGIIRYAIGVGLAFQNRNSWKELNDIASKPSQEHIF 314
>ATK1_ARATH (Q07970) Kinesin 1 (Kinesin-like protein A)
Length = 793
Score = 30.4 bits (67), Expect = 9.0
Identities = 20/61 (32%), Positives = 31/61 (50%), Gaps = 1/61 (1%)
Query: 222 MVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSMNTTSRS 281
M +NQ + A KE GI+++ R VE Q G G +A S N+ + +MN+ S
Sbjct: 1 MASRNQNRPPRSPNAKKEGLGGISFDKRRKVETQGGTGRRQAFSAVNK-QDVTMNSDVGS 59
Query: 282 L 282
+
Sbjct: 60 I 60
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,293,062
Number of Sequences: 164201
Number of extensions: 2128312
Number of successful extensions: 4222
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 4220
Number of HSP's gapped (non-prelim): 8
length of query: 425
length of database: 59,974,054
effective HSP length: 113
effective length of query: 312
effective length of database: 41,419,341
effective search space: 12922834392
effective search space used: 12922834392
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0227.11


