

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.9
(223 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
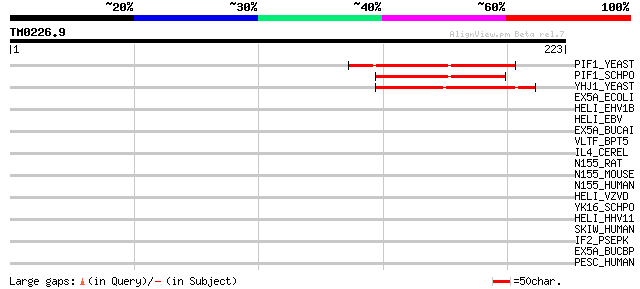
PIF1_YEAST (P07271) DNA repair and recombination protein PIF1, m... 52 1e-06
PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1, m... 49 7e-06
YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergeni... 47 5e-05
EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 36 0.063
HELI_EHV1B (P28934) Probable helicase 35 0.14
HELI_EBV (P03214) Probable helicase 33 0.41
EX5A_BUCAI (P57530) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 33 0.54
VLTF_BPT5 (P13390) L-shaped tail fiber protein (LTF protein) 32 0.91
IL4_CEREL (P51744) Interleukin-4 precursor (IL-4) (B-cell stimul... 32 0.91
N155_RAT (P37199) Nuclear pore complex protein Nup155 (Nucleopor... 32 1.6
N155_MOUSE (Q99P88) Nuclear pore complex protein Nup155 (Nucleop... 32 1.6
N155_HUMAN (O75694) Nuclear pore complex protein Nup155 (Nucleop... 31 2.0
HELI_VZVD (P09303) Probable helicase 31 2.0
YK16_SCHPO (Q9HDY4) Putative ATP-dependent RNA helicase PB1A10.0... 30 4.5
HELI_HHV11 (P10189) Probable helicase 30 4.5
SKIW_HUMAN (Q15477) Helicase SKI2W (Helicase-like protein) (HLP) 30 5.9
IF2_PSEPK (Q88DV7) Translation initiation factor IF-2 30 5.9
EX5A_BUCBP (Q89AB2) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 30 5.9
PESC_HUMAN (O00541) Pescadillo homolog 1 29 7.7
>PIF1_YEAST (P07271) DNA repair and recombination protein PIF1,
mitochondrial precursor
Length = 857
Score = 52.0 bits (123), Expect = 1e-06
Identities = 30/67 (44%), Positives = 47/67 (69%), Gaps = 2/67 (2%)
Query: 137 DSKFPIKFRIRQFPIAICFAMTINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRVRSRKG 196
+++ P+ R+ Q P+ + ++++I+KSQGQTL +V + L R VF GQ YVALSR SR+G
Sbjct: 683 ENEKPLVSRV-QLPLMLAWSLSIHKSQGQTLPKVKVDLRR-VFEKGQAYVALSRAVSREG 740
Query: 197 LKLCILD 203
L++ D
Sbjct: 741 LQVLNFD 747
>PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1,
mitochondrial precursor
Length = 805
Score = 49.3 bits (116), Expect = 7e-06
Identities = 25/52 (48%), Positives = 39/52 (74%), Gaps = 1/52 (1%)
Query: 148 QFPIAICFAMTINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRVRSRKGLKL 199
Q P+ + +A++I+K+QGQTL +V + L R VF GQ YVALSR +++GL++
Sbjct: 710 QIPLILAYAISIHKAQGQTLDRVKVDLGR-VFEKGQAYVALSRATTQEGLQV 760
>YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergenic
region
Length = 723
Score = 46.6 bits (109), Expect = 5e-05
Identities = 26/64 (40%), Positives = 44/64 (68%), Gaps = 2/64 (3%)
Query: 148 QFPIAICFAMTINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRVRSRKGLKLCILDEAGK 207
Q P+ +C+A++I+K+QGQT+ ++ + L R +F GQ+YVALSR + L++ D GK
Sbjct: 648 QIPLMLCWALSIHKAQGQTIQRLKVDL-RRIFEAGQVYVALSRAVTMDTLQVLNFD-PGK 705
Query: 208 QQTS 211
+T+
Sbjct: 706 IRTN 709
>EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5) (Exodeoxyribonuclease V 67 kDa polypeptide)
Length = 608
Score = 36.2 bits (82), Expect = 0.063
Identities = 36/120 (30%), Positives = 48/120 (40%), Gaps = 22/120 (18%)
Query: 81 GTPIMLLRNIDQAASLCNGTRLIVDELGERFIGATVITGTNVNDKVHILRMDLVLSDSKF 140
G P+M+ RN D A L NG I + G+ GT R+ + D
Sbjct: 479 GRPVMIARN-DSALGLFNGDIGIALDRGQ---------GT---------RVWFAMPDGNI 519
Query: 141 PIKFRIRQFPIAICFAMTINKSQGQTLSQVGLFLP---RPVFTHGQLYVALSRVRSRKGL 197
R +AMT++KSQG L LP PV T +Y A++R R R L
Sbjct: 520 KSVQPSRLPEHETTWAMTVHKSQGSEFDHAALILPSQRTPVVTRELVYTAVTRARRRLSL 579
>HELI_EHV1B (P28934) Probable helicase
Length = 881
Score = 35.0 bits (79), Expect = 0.14
Identities = 20/58 (34%), Positives = 32/58 (54%)
Query: 156 AMTINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRVRSRKGLKLCILDEAGKQQTSTV 213
AMTI +SQG +L +V + PR +YVA+SR S + L++ + + + TV
Sbjct: 806 AMTIARSQGLSLDKVAICFPRNNLRINSVYVAMSRTVSSRFLRMNLNPLRERHERDTV 863
>HELI_EBV (P03214) Probable helicase
Length = 809
Score = 33.5 bits (75), Expect = 0.41
Identities = 21/48 (43%), Positives = 29/48 (59%), Gaps = 1/48 (2%)
Query: 146 IRQFPIAICFAMTINKSQGQTLSQVGL-FLPRPVFTHGQLYVALSRVR 192
IR + I+ AMTI K+QG +L++V + F G +YVALSR R
Sbjct: 722 IRDYGISSKLAMTIAKAQGLSLNKVAICFGSHRNIKPGHVYVALSRAR 769
>EX5A_BUCAI (P57530) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5)
Length = 602
Score = 33.1 bits (74), Expect = 0.54
Identities = 31/116 (26%), Positives = 53/116 (44%), Gaps = 21/116 (18%)
Query: 80 VGTPIMLLRNIDQAASLCNGTRLIVDELGERFIGATVITGTNVNDKVHILRMDLVLSDSK 139
+G PIM++ N ++A ++ NG IG IT N N + + + + +
Sbjct: 480 IGKPIMIINN-NRALNVSNGN-----------IG---ITNINKNGILQVSFLKENNTINN 524
Query: 140 FPIKFRIRQFPIAICFAMTINKSQGQTLSQVGLFLPR---PVFTHGQLYVALSRVR 192
P+K +R + A +A+T++KSQG L LP + LY ++R R
Sbjct: 525 IPVKI-LRNYKTA--WAITVHKSQGSEFMNTALILPNFNSHILNKDTLYTGITRSR 577
>VLTF_BPT5 (P13390) L-shaped tail fiber protein (LTF protein)
Length = 1396
Score = 32.3 bits (72), Expect = 0.91
Identities = 18/49 (36%), Positives = 26/49 (52%)
Query: 48 DEDSEIKVEWFTTEFSNDTKSSGMPNHQLLLKVGTPIMLLRNIDQAASL 96
D K+ T EFSND ++ N ++ K TPIML+R+ D S+
Sbjct: 586 DAKISYKISSRTAEFSNDDTNTAATNLRVSGKQHTPIMLVRDSDSNVSV 634
>IL4_CEREL (P51744) Interleukin-4 precursor (IL-4) (B-cell
stimulatory factor 1) (BSF-1) (Lymphocyte stimulatory
factor 1)
Length = 135
Score = 32.3 bits (72), Expect = 0.91
Identities = 26/69 (37%), Positives = 39/69 (55%), Gaps = 4/69 (5%)
Query: 148 QFPIAICFAMTINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRV-RSRKGL--KLCILDE 204
+ P+A FA N ++ +TL + G+ L R +H L LSR+ R+ GL K C ++E
Sbjct: 50 ELPVADVFAAPKNTTEKETLCRAGIELRRIYRSHTCLNRFLSRLDRNLSGLASKTCSVNE 109
Query: 205 AGKQQTSTV 213
A K TST+
Sbjct: 110 A-KTSTSTL 117
>N155_RAT (P37199) Nuclear pore complex protein Nup155 (Nucleoporin
Nup155) (155 kDa nucleoporin) (P140)
Length = 1390
Score = 31.6 bits (70), Expect = 1.6
Identities = 27/93 (29%), Positives = 48/93 (51%), Gaps = 17/93 (18%)
Query: 123 NDKVHILRMDLVL-------SDSKFPIKFRIRQFPIAICFAMTINKSQG---QTLSQVGL 172
+D++H L + LVL + FP+ F ++ +C T+N G QT++++G+
Sbjct: 1237 SDRMHALSLKLVLLGKIYAGTPRFFPLDFIVQFLEQQVC---TLNWDVGFVIQTMNEIGV 1293
Query: 173 FLPRPVFTHGQLYVA----LSRVRSRKGLKLCI 201
LPR + + QL+ + +RV+S L CI
Sbjct: 1294 PLPRLLEVYDQLFKSRDPFWNRVKSPLHLLDCI 1326
>N155_MOUSE (Q99P88) Nuclear pore complex protein Nup155 (Nucleoporin
Nup155) (155 kDa nucleoporin)
Length = 1391
Score = 31.6 bits (70), Expect = 1.6
Identities = 27/93 (29%), Positives = 48/93 (51%), Gaps = 17/93 (18%)
Query: 123 NDKVHILRMDLVL-------SDSKFPIKFRIRQFPIAICFAMTINKSQG---QTLSQVGL 172
+D++H L + LVL + FP+ F ++ +C T+N G QT++++G+
Sbjct: 1238 SDRMHALSLKLVLLGKIYAGTPRFFPLDFIVQFLEQQVC---TLNWDVGFVIQTMNEIGV 1294
Query: 173 FLPRPVFTHGQLYVA----LSRVRSRKGLKLCI 201
LPR + + QL+ + +RV+S L CI
Sbjct: 1295 PLPRLLEVYDQLFKSRDPFWNRVKSPLHLLDCI 1327
>N155_HUMAN (O75694) Nuclear pore complex protein Nup155 (Nucleoporin
Nup155) (155 kDa nucleoporin)
Length = 1391
Score = 31.2 bits (69), Expect = 2.0
Identities = 24/91 (26%), Positives = 46/91 (50%), Gaps = 13/91 (14%)
Query: 123 NDKVHILRMDLVL-------SDSKFPIKFRIRQFPIAICFAMTINKSQG---QTLSQVGL 172
+D++H L + +VL + FP+ F ++ +C T+N G QT++++G+
Sbjct: 1238 SDRMHALSLKIVLLGKIYAGTPRFFPLDFIVQFLEQQVC---TLNWDVGFVIQTMNEIGV 1294
Query: 173 FLPRPVFTHGQLYVALSRVRSRKGLKLCILD 203
LPR + + QL+ + +R L +LD
Sbjct: 1295 PLPRLLEVYDQLFKSRDPFWNRMKKPLHLLD 1325
>HELI_VZVD (P09303) Probable helicase
Length = 881
Score = 31.2 bits (69), Expect = 2.0
Identities = 17/44 (38%), Positives = 25/44 (56%)
Query: 156 AMTINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRVRSRKGLKL 199
AMTI +SQG +L +V + +YVA+SR S + LK+
Sbjct: 806 AMTIARSQGLSLEKVAICFTADKLRLNSVYVAMSRTVSSRFLKM 849
>YK16_SCHPO (Q9HDY4) Putative ATP-dependent RNA helicase PB1A10.06c
(EC 3.6.1.-)
Length = 1183
Score = 30.0 bits (66), Expect = 4.5
Identities = 29/119 (24%), Positives = 55/119 (45%), Gaps = 6/119 (5%)
Query: 29 GMIAAPEHEYLSSDSALRSDEDSEIKVEWFTTEFSNDTKSSGMPNHQLLLKVGTPIMLLR 88
GMIA + +++ S + + ++ F+++ S + N +K T +LLR
Sbjct: 449 GMIAITQPRRVAAVSIAKRVSE---ELTGFSSKVSYQIRFDSTINPDTAIKFMTDGILLR 505
Query: 89 NIDQAASLCNGTRLIVDELGERFIGATVITGTNVNDKVHILRMDLVLSDSKF-PIKFRI 146
+ L + +IVDE ER + ++ G + ++ LR ++ SD K P+K I
Sbjct: 506 ELSSDFLLTAYSAVIVDEAHERSVNTDILLG--LLSRIVRLRREMSKSDQKVKPLKLII 562
>HELI_HHV11 (P10189) Probable helicase
Length = 882
Score = 30.0 bits (66), Expect = 4.5
Identities = 21/56 (37%), Positives = 31/56 (54%), Gaps = 10/56 (17%)
Query: 149 FPIAICFAMTINKSQGQTLSQVGLFLPRPVFTHGQL-----YVALSRVRSRKGLKL 199
+ I+ AMTI +SQG +L +V + FT G L YVA+SR S + L++
Sbjct: 800 YGISSKLAMTITRSQGLSLDKVAI-----CFTPGNLRLNSAYVAMSRTTSSEFLRM 850
>SKIW_HUMAN (Q15477) Helicase SKI2W (Helicase-like protein) (HLP)
Length = 1246
Score = 29.6 bits (65), Expect = 5.9
Identities = 22/72 (30%), Positives = 35/72 (48%), Gaps = 7/72 (9%)
Query: 70 GMPNHQLLLKVGTPI-MLLRNIDQAASLCNGTRLIVDELGERFIGATVITGTNVNDKVHI 128
GMP +L GTP +++R I + A +C R +GE +GA + T +
Sbjct: 1179 GMPFSELAGLSGTPEGLVVRCIQRLAEMCRSLRGAARLVGEPVLGAKMETAAT------L 1232
Query: 129 LRMDLVLSDSKF 140
LR D+V + S +
Sbjct: 1233 LRRDIVFAASLY 1244
>IF2_PSEPK (Q88DV7) Translation initiation factor IF-2
Length = 846
Score = 29.6 bits (65), Expect = 5.9
Identities = 17/67 (25%), Positives = 35/67 (51%), Gaps = 8/67 (11%)
Query: 60 TEFSNDTKSSGMPNHQLLLKVGTPIMLLRNIDQAASLCNGTRLIVDELGERFIGATVITG 119
+E +N G + + K+GTP+ + + +DQ + +LI +ELG + T+++
Sbjct: 276 SELANQMSVKGAEVVKFMFKMGTPVTINQVLDQETA-----QLIAEELGHK---VTLVSD 327
Query: 120 TNVNDKV 126
T + D +
Sbjct: 328 TALEDSL 334
>EX5A_BUCBP (Q89AB2) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5)
Length = 618
Score = 29.6 bits (65), Expect = 5.9
Identities = 15/43 (34%), Positives = 25/43 (57%), Gaps = 3/43 (6%)
Query: 155 FAMTINKSQGQTLSQVGLFLP---RPVFTHGQLYVALSRVRSR 194
+ MT++KSQG S+V L LP + T +Y A++R + +
Sbjct: 544 WTMTVHKSQGSEFSEVVLILPTIMTSILTKELIYTAVTRSKKK 586
>PESC_HUMAN (O00541) Pescadillo homolog 1
Length = 588
Score = 29.3 bits (64), Expect = 7.7
Identities = 16/44 (36%), Positives = 23/44 (51%)
Query: 54 KVEWFTTEFSNDTKSSGMPNHQLLLKVGTPIMLLRNIDQAASLC 97
K EW T E D K + +H + + T I LR++D A S+C
Sbjct: 98 KSEWNTVERLKDNKPNYKLDHIIKERYPTFIDALRDLDDALSMC 141
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.137 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,457,813
Number of Sequences: 164201
Number of extensions: 902664
Number of successful extensions: 1897
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 1884
Number of HSP's gapped (non-prelim): 22
length of query: 223
length of database: 59,974,054
effective HSP length: 106
effective length of query: 117
effective length of database: 42,568,748
effective search space: 4980543516
effective search space used: 4980543516
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0226.9


