

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.8
(518 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
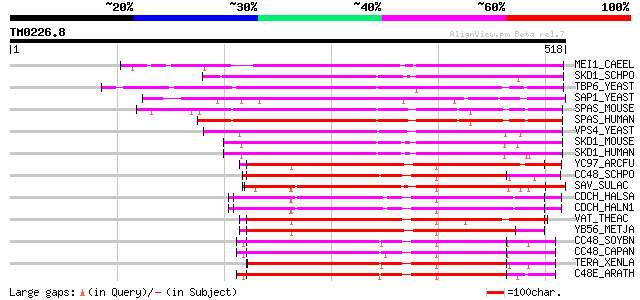
Score E
Sequences producing significant alignments: (bits) Value
MEI1_CAEEL (P34808) Meiotic spindle formation protein mei-1 290 6e-78
SKD1_SCHPO (Q09803) Suppressor protein of bem1/bed5 double mutants 268 3e-71
TBP6_YEAST (P40328) Probable 26S protease subunit YTA6 (TAT-bind... 263 8e-70
SAP1_YEAST (P39955) SAP1 protein 259 2e-68
SPAS_MOUSE (Q9QYY8) Spastin 252 2e-66
SPAS_HUMAN (Q9UBP0) Spastin 251 3e-66
VPS4_YEAST (P52917) Vacuolar protein sorting-associated protein ... 248 3e-65
SKD1_MOUSE (P46467) SKD1 protein (Vacuolar sorting protein 4b) 248 3e-65
SKD1_HUMAN (O75351) SKD1 protein (Vacuolar sorting protein 4b) 248 4e-65
YC97_ARCFU (O28972) Cell division cycle protein 48 homolog AF1297 238 3e-62
CC48_SCHPO (Q9P3A7) Cell division cycle protein 48 homolog 211 5e-54
SAV_SULAC (Q07590) SAV protein 210 8e-54
CDCH_HALSA (P46464) CdcH protein 202 2e-51
CDCH_HALN1 (Q9HPF0) CdcH protein 202 2e-51
VAT_THEAC (O05209) VCP-like ATPase 200 6e-51
YB56_METJA (Q58556) Cell division cycle protein 48 homolog MJ1156 199 2e-50
CC48_SOYBN (P54774) Cell division cycle protein 48 homolog (Valo... 199 2e-50
CC48_CAPAN (Q96372) Cell division cycle protein 48 homolog 197 4e-50
TERA_XENLA (P23787) Transitional endoplasmic reticulum ATPase (T... 197 7e-50
C48E_ARATH (Q9LZF6) Cell division control protein 48 homolog E (... 196 1e-49
>MEI1_CAEEL (P34808) Meiotic spindle formation protein mei-1
Length = 472
Score = 290 bits (742), Expect = 6e-78
Identities = 173/421 (41%), Positives = 245/421 (58%), Gaps = 38/421 (9%)
Query: 104 LDEYPTSSSG---PMDDPDVWRPPSRDTSRRPTRPGQMSTRKSDGAWARGATPRGGATGR 160
L E T SG P DPDVW PS P+ +T+K GA A PR +
Sbjct: 82 LHEAMTRQSGSPEPPADPDVWSKPSPPL---PSSSKFGATKKGVGA----AGPRPREISK 134
Query: 161 GAKAGATSKVNTGTRASTTG----KKSGVSGKAGKTDSVISDAEESKSKKSQYEGPDPEL 216
+ +T+ + T G +G S A D+ I A +
Sbjct: 135 STSSMSTNPADVKPANPTQGILPQNSAGDSFDASAYDAYIVQAVRGTMATN--------- 185
Query: 217 AEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPP 276
T + +D+ G+ + K++L EAV LPL +PE+FQG+R PWK +++ GPP
Sbjct: 186 ----------TENTMSLDDIIGMHDVKQVLHEAVTLPLLVPEFFQGLRSPWKAMVLAGPP 235
Query: 277 GTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEID 336
GTGKTL+A+A+A+E +TFF VSS L+SKWRG+SE++VR LF+LAR YAPS IFIDEID
Sbjct: 236 GTGKTLIARAIASESSSTFFTVSSTDLSSKWRGDSEKIVRLLFELARFYAPSIIFIDEID 295
Query: 337 SLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALR 396
+L RG SGEHE+SRRVKSE LVQ+DG + N+ SR+ V VLAATN PW++DEALR
Sbjct: 296 TLGGQRGNSGEHEASRRVKSEFLVQMDG----SQNKFDSRR-VFVLAATNIPWELDEALR 350
Query: 397 RRLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASL 456
RR EKRI+IPLP+ ++RK+LI +++ + ++N D++A RT+G+SG D+ ++CR A++
Sbjct: 351 RRFEKRIFIPLPDIDARKKLIEKSMEGTPKSDEINYDDLAARTEGFSGADVVSLCRTAAI 410
Query: 457 NGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKWFHEFG 516
N +RR R + + + + V DFE AL V S + + ++W FG
Sbjct: 411 NVLRRYDTKSLRGGELTAAMESLKAELVRNIDFEAALQAVSPSAGPDTMLKCKEWCDSFG 470
Query: 517 S 517
+
Sbjct: 471 A 471
>SKD1_SCHPO (Q09803) Suppressor protein of bem1/bed5 double mutants
Length = 432
Score = 268 bits (685), Expect = 3e-71
Identities = 150/363 (41%), Positives = 218/363 (59%), Gaps = 35/363 (9%)
Query: 181 KKSGVSGKAGKTDSVISDAEESKSKKSQYEGPDPELAEMLERDVLETSPGVRWEDVAGLT 240
K + +S K+ ++ + + S + + +L L +L P VRW+D+AGL
Sbjct: 77 KNNQISSKSRVSNGNV-EGSNSPTANEALDSDAKKLRSALTSAILVEKPNVRWDDIAGLE 135
Query: 241 EAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSS 300
AK L+E V+LP+ +P+ F R+PW G+L++GPPGTGK+ LAKAVATE G+TFF++SS
Sbjct: 136 NAKEALKETVLLPIKLPQLFSHGRKPWSGILLYGPPGTGKSYLAKAVATEAGSTFFSISS 195
Query: 301 ATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLV 360
+ L SKW GESER+VR LF++AR PS IFIDEIDSLC SR + GE ESSRR+K+E LV
Sbjct: 196 SDLVSKWMGESERLVRQLFEMAREQKPSIIFIDEIDSLCGSR-SEGESESSRRIKTEFLV 254
Query: 361 QVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRIN 420
Q++GV +E G V+VL ATN PW +D A+RRR EKRIYIPLPN +R + +N
Sbjct: 255 QMNGVGK---DESG----VLVLGATNIPWTLDSAIRRRFEKRIYIPLPNAHARARMFELN 307
Query: 421 L-KTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKN------ 473
+ K + E+A+ TDGYSG D++ V RDA + +RR E+ +
Sbjct: 308 VGKIPSELTSQDFKELAKMTDGYSGSDISIVVRDAIMEPVRRIHTATHFKEVYDNKSNRT 367
Query: 474 -------------------MSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKWFHE 514
++ +DI + + + DF A+ KV+ +++ DIE+H ++ +
Sbjct: 368 LVTPCSPGDPDAFESSWLEVNPEDIMEPKLTVRDFYSAVRKVKPTLNAGDIEKHTQFTKD 427
Query: 515 FGS 517
FG+
Sbjct: 428 FGA 430
>TBP6_YEAST (P40328) Probable 26S protease subunit YTA6 (TAT-binding
homolog 6)
Length = 754
Score = 263 bits (672), Expect = 8e-70
Identities = 158/437 (36%), Positives = 250/437 (57%), Gaps = 25/437 (5%)
Query: 86 RRPPSPPISTKSSFVFQPLDEYPTSSSGPMDDPDVWRPPSRDTSRRPTRPGQMSTRKSDG 145
R P+PP+ + + + PT + + P + R S+ + PT +++ +
Sbjct: 335 RYKPTPPLKKRYDY------KKPTVNRPIIKSPTLNRQNSKSSRNIPTNSKLKASKSNTN 388
Query: 146 AWARGATPRGGATGRGAKAGATSKVNTGTRASTTGKKSGVSGKAGKTDSVISDAEESKSK 205
+R R + S + KSG K S + +E K
Sbjct: 389 KVSR----RNEQNLEPSSPVLVSATAVPAESKPMRSKSGTPDKESSASSSLDSRKEDILK 444
Query: 206 KSQYEGPDPELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRR 265
Q G D E + ++L T V WED+AGL AK L+EAVV P P+ F+G+R
Sbjct: 445 SVQ--GVDRNACEQILNEILVTDEKVYWEDIAGLRNAKNSLKEAVVYPFLRPDLFKGLRE 502
Query: 266 PWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAY 325
P +G+L+FGPPGTGKT++AKAVATE +TFF+VS+++L SK+ GESE++VR LF +A+
Sbjct: 503 PVRGMLLFGPPGTGKTMIAKAVATESNSTFFSVSASSLLSKYLGESEKLVRALFYMAKKL 562
Query: 326 APSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSN-TATNEDGSRKI---VMV 381
+PS IFIDEIDS+ +R + E+ESSRR+K+ELL+Q +S+ TA +ED + + V+V
Sbjct: 563 SPSIIFIDEIDSMLTAR-SDNENESSRRIKTELLIQWSSLSSATAQSEDRNNTLDSRVLV 621
Query: 382 LAATNFPWDIDEALRRRLEKRIYIPLPNFESR-KELIRINLKTVEVAPDVNIDEVARRTD 440
L ATN PW ID+A RRR +++YIPLP++E+R L R+ K D++ + + T+
Sbjct: 622 LGATNLPWAIDDAARRRFSRKLYIPLPDYETRLYHLKRLMAKQKNSLQDLDYELITEMTE 681
Query: 441 GYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSV 500
G+SG DLT++ ++A++ +R D++ D I + + DF+ AL+ +++SV
Sbjct: 682 GFSGSDLTSLAKEAAMEPIR-----DLGDKLMFADFDKIR--GIEIKDFQNALLTIKKSV 734
Query: 501 SQADIERHEKWFHEFGS 517
S ++++E+W +FGS
Sbjct: 735 SSESLQKYEEWSSKFGS 751
>SAP1_YEAST (P39955) SAP1 protein
Length = 897
Score = 259 bits (661), Expect = 2e-68
Identities = 166/436 (38%), Positives = 241/436 (55%), Gaps = 68/436 (15%)
Query: 125 SRDTSRRPTRPGQMSTRKSDGAWARGATPRGGATGRGAKAGATSKVNTGTRASTTGKKSG 184
S+ TS P+RP S S GA N T S T +K
Sbjct: 486 SKKTSSHPSRPVSNSKPYS----------------HGASQNKKPSKNQTTSMSKTNRKIP 529
Query: 185 VSGKAGKT---DSVISDA-EESKSKKSQYEGPDPE---LAEMLERDVLETSPGV------ 231
K G D DA E + S Q E P+ + L E+LE +++++ GV
Sbjct: 530 AQKKIGSPKIEDVGTEDATEHATSLNEQREEPEIDKKVLREILEDEIIDSLQGVDRQAAK 589
Query: 232 -------------RWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGT 278
W+D+AGL AK L+EAVV P P+ F+G+R P +G+L+FGPPGT
Sbjct: 590 QIFAEIVVHGDEVHWDDIAGLESAKYSLKEAVVYPFLRPDLFRGLREPVRGMLLFGPPGT 649
Query: 279 GKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSL 338
GKT+LA+AVATE +TFF++S+++L SK+ GESE++VR LF +A+ +PS IF+DEIDS+
Sbjct: 650 GKTMLARAVATESHSTFFSISASSLTSKYLGESEKLVRALFAIAKKLSPSIIFVDEIDSI 709
Query: 339 CNSRGASGEHESSRRVKSELLVQVDGVS------------NTATNEDGSRKIVMVLAATN 386
SR E+ESSRR+K+E LVQ +S N+ TN D V+VLAATN
Sbjct: 710 MGSRNNENENESSRRIKNEFLVQWSSLSSAAAGSNKSNTNNSDTNGDEDDTRVLVLAATN 769
Query: 387 FPWDIDEALRRRLEKRIYIPLPNFESR----KELIRINLKTVEVAPDVNIDEVARRTDGY 442
PW IDEA RRR +R YIPLP ++R K+L+ T+ + + DE+ + T+GY
Sbjct: 770 LPWSIDEAARRRFVRRQYIPLPEDQTRHVQFKKLLSHQKHTL---TESDFDELVKITEGY 826
Query: 443 SGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQ 502
SG D+T++ +DA++ G R + K + + M + P+ + DF+ +LV ++ SVSQ
Sbjct: 827 SGSDITSLAKDAAM-GPLRDLGDKLLETEREMIR------PIGLVDFKNSLVYIKPSVSQ 879
Query: 503 ADIERHEKWFHEFGSA 518
+ ++EKW +FGS+
Sbjct: 880 DGLVKYEKWASQFGSS 895
>SPAS_MOUSE (Q9QYY8) Spastin
Length = 614
Score = 252 bits (643), Expect = 2e-66
Identities = 154/418 (36%), Positives = 238/418 (56%), Gaps = 35/418 (8%)
Query: 119 DVWRPPSRDTSRR---PTRPGQMSTRKSDGAWARGATPRGGATGRGAKAGATS------- 168
DV+ + T R + G + RK A + PR + AG +
Sbjct: 208 DVYNESTNLTCRNGHLQSESGAVPKRKDPLTHASNSLPRSKTVLKSGSAGLSGHHRAPSC 267
Query: 169 ---KVNTGTR-----ASTTGKKSGVSGKAGKTDSVISDAEESKSKKSQYEGPDPELAEML 220
+ +G R A+TT K + + K + + + K K+ + D LA ++
Sbjct: 268 SGLSMVSGARPGPGPAATTHKGTPKPNRTNKPSTPTTAVRKKKDLKN-FRNVDSNLANLI 326
Query: 221 ERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGK 280
++++ V+++D+AG AK+ L+E V+LP PE F G+R P +G+L+FGPPG GK
Sbjct: 327 MNEIVDNGTAVKFDDIAGQELAKQALQEIVILPSLRPELFTGLRAPARGLLLFGPPGNGK 386
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
T+LAKAVA E TFFN+S+A+L SK+ GE E++VR LF +AR PS IFIDE+DSL
Sbjct: 387 TMLAKAVAAESNATFFNISAASLTSKYVGEGEKLVRALFAVARELQPSIIFIDEVDSLLC 446
Query: 341 SRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLE 400
R GEH++SRR+K+E L++ DGV + + V+V+ ATN P ++DEA+ RR
Sbjct: 447 ER-REGEHDASRRLKTEFLIEFDGVQSAGDDR------VLVMGATNRPQELDEAVLRRFI 499
Query: 401 KRIYIPLPNFESRKELIRINLKTVEVAP--DVNIDEVARRTDGYSGDDLTNVCRDASLNG 458
KR+Y+ LPN E+R L++ NL + +P + ++AR TDGYSG DLT + +DA+L
Sbjct: 500 KRVYVSLPNEETRLLLLK-NLLCKQGSPLTQKELAQLARMTDGYSGSDLTALAKDAALGP 558
Query: 459 MRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKWFHEFG 516
+R +++KNMS ++ + + DF E+L K++RSVS +E + +W +FG
Sbjct: 559 IRE----LKPEQVKNMSASEMRN--IRLSDFTESLKKIKRSVSPQTLEAYIRWNKDFG 610
>SPAS_HUMAN (Q9UBP0) Spastin
Length = 616
Score = 251 bits (641), Expect = 3e-66
Identities = 140/343 (40%), Positives = 215/343 (61%), Gaps = 17/343 (4%)
Query: 176 ASTTGKKSGVSGKAGKTDSVISDAEESKSKKSQYEGPDPELAEMLERDVLETSPGVRWED 235
A TT K + + + K + + + K K+ + D LA ++ ++++ V+++D
Sbjct: 285 APTTHKGTPKTNRTNKPSTPTTATRKKKDLKN-FRNVDSNLANLIMNEIVDNGTAVKFDD 343
Query: 236 VAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTF 295
+AG AK+ L+E V+LP PE F G+R P +G+L+FGPPG GKT+LAKAVA E TF
Sbjct: 344 IAGQDLAKQALQEIVILPSLRPELFTGLRAPARGLLLFGPPGNGKTMLAKAVAAESNATF 403
Query: 296 FNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVK 355
FN+S+A+L SK+ GE E++VR LF +AR PS IFIDE+DSL R GEH++SRR+K
Sbjct: 404 FNISAASLTSKYVGEGEKLVRALFAVARELQPSIIFIDEVDSLLCER-REGEHDASRRLK 462
Query: 356 SELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKE 415
+E L++ DGV + + V+V+ ATN P ++DEA+ RR KR+Y+ LPN E+R
Sbjct: 463 TEFLIEFDGVQSAGDDR------VLVMGATNRPQELDEAVLRRFIKRVYVSLPNEETRLL 516
Query: 416 LIRINLKTVEVAP--DVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKN 473
L++ NL + +P + ++AR TDGYSG DLT + +DA+L +R +++KN
Sbjct: 517 LLK-NLLCKQGSPLTQKELAQLARMTDGYSGSDLTALAKDAALGPIRE----LKPEQVKN 571
Query: 474 MSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKWFHEFG 516
MS ++ + + DF E+L K++RSVS +E + +W +FG
Sbjct: 572 MSASEMRN--IRLSDFTESLKKIKRSVSPQTLEAYIRWNKDFG 612
>VPS4_YEAST (P52917) Vacuolar protein sorting-associated protein
VPS4 (END13 protein)
Length = 437
Score = 248 bits (633), Expect = 3e-65
Identities = 146/367 (39%), Positives = 213/367 (57%), Gaps = 40/367 (10%)
Query: 182 KSGVSGKAGKTDSVISDAEESKSKKSQYEGPD------PELAEMLERDVLETSPGVRWED 235
+S + A K+ S S + K SQ EG D +L L +L P V+WED
Sbjct: 75 ESEEANAAKKSPSAGSGSNGGNKKISQEEGEDNGGEDNKKLRGALSSAILSEKPNVKWED 134
Query: 236 VAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTF 295
VAGL AK L+EAV+LP+ P F+G R+P G+L++GPPGTGK+ LAKAVATE +TF
Sbjct: 135 VAGLEGAKEALKEAVILPVKFPHLFKGNRKPTSGILLYGPPGTGKSYLAKAVATEANSTF 194
Query: 296 FNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVK 355
F+VSS+ L SKW GESE++V+ LF +AR PS IFIDE+D+L +RG GE E+SRR+K
Sbjct: 195 FSVSSSDLVSKWMGESEKLVKQLFAMARENKPSIIFIDEVDALTGTRG-EGESEASRRIK 253
Query: 356 SELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKE 415
+ELLVQ++GV N + V+VL ATN PW +D A+RRR E+RIYIPLP+ +R
Sbjct: 254 TELLVQMNGVGNDSQG-------VLVLGATNIPWQLDSAIRRRFERRIYIPLPDLAARTT 306
Query: 416 LIRINL-KTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRR-------KIAGKT 467
+ IN+ T V + + T+GYSG D+ V +DA + +R+ K
Sbjct: 307 MFEINVGDTPCVLTKEDYRTLGAMTEGYSGSDIAVVVKDALMQPIRKIQSATHFKDVSTE 366
Query: 468 RDEIKNMS------------------KDDISKDPVAMCDFEEALVKVQRSVSQADIERHE 509
DE + ++ D++ + + + DF +A+ + +V++ D+ + E
Sbjct: 367 DDETRKLTPCSPGDDGAIEMSWTDIEADELKEPDLTIKDFLKAIKSTRPTVNEDDLLKQE 426
Query: 510 KWFHEFG 516
++ +FG
Sbjct: 427 QFTRDFG 433
>SKD1_MOUSE (P46467) SKD1 protein (Vacuolar sorting protein 4b)
Length = 444
Score = 248 bits (633), Expect = 3e-65
Identities = 144/353 (40%), Positives = 207/353 (57%), Gaps = 44/353 (12%)
Query: 200 EESKSKKSQYEGPDPE---LAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWM 256
E+ + E DPE L L+ ++ P V+W DVAGL AK L+EAV+LP+
Sbjct: 97 EKGNDSDGEAESDDPEKKKLQNQLQGAIVIERPNVKWSDVAGLEGAKEALKEAVILPIKF 156
Query: 257 PEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATEC-GTTFFNVSSATLASKWRGESERMV 315
P F G R PW+G+L+FGPPGTGK+ LAKAVATE +TFF++SS+ L SKW GESE++V
Sbjct: 157 PHLFTGKRTPWRGILLFGPPGTGKSYLAKAVATEANNSTFFSISSSDLVSKWLGESEKLV 216
Query: 316 RCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGS 375
+ LF LAR PS IFIDEIDSLC SR + E E++RR+K+E LVQ+ GV + DG
Sbjct: 217 KNLFQLARENKPSIIFIDEIDSLCGSR-SENESEAARRIKTEFLVQMQGV---GVDNDG- 271
Query: 376 RKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINL-KTVEVAPDVNIDE 434
++VL ATN PW +D A+RRR EKRIYIPLP +R + R++L T + + E
Sbjct: 272 ---ILVLGATNIPWVLDSAIRRRFEKRIYIPLPEAHARAAMFRLHLGSTQNSLTEADFQE 328
Query: 435 VARRTDGYSGDDLTNVCRDASLNGMR--------RKIAGKTRDEIK-------------- 472
+ R+TDGYSG D++ + RDA + +R +K+ G +R +
Sbjct: 329 LGRKTDGYSGADISIIVRDALMQPVRKVQSATHFKKVRGPSRADPNCIVNDLLTPCSPGD 388
Query: 473 ---------NMSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKWFHEFG 516
++ D + + V+M D +L + +V++ D+ + +K+ +FG
Sbjct: 389 PGAIEMTWMDVPGDKLLEPVVSMWDMLRSLSSTKPTVNEQDLLKLKKFTEDFG 441
>SKD1_HUMAN (O75351) SKD1 protein (Vacuolar sorting protein 4b)
Length = 444
Score = 248 bits (632), Expect = 4e-65
Identities = 145/353 (41%), Positives = 211/353 (59%), Gaps = 44/353 (12%)
Query: 200 EESKSKKSQYEGPDPE---LAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWM 256
E+ + E DPE L L+ ++ P V+W DVAGL AK L+EAV+LP+
Sbjct: 97 EKGNDSDGEGESDDPEKKKLQNQLQGAIVIERPNVKWSDVAGLEGAKEALKEAVILPIKF 156
Query: 257 PEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATEC-GTTFFNVSSATLASKWRGESERMV 315
P F G R PW+G+L+FGPPGTGK+ LAKAVATE +TFF++SS+ L SKW GESE++V
Sbjct: 157 PHLFTGKRTPWRGILLFGPPGTGKSYLAKAVATEANNSTFFSISSSDLVSKWLGESEKLV 216
Query: 316 RCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGS 375
+ LF LAR PS IFIDEIDSLC SR + E E++RR+K+E LVQ+ GV + DG
Sbjct: 217 KNLFQLARENKPSIIFIDEIDSLCGSR-SENESEAARRIKTEFLVQMQGV---GVDNDG- 271
Query: 376 RKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINLKTVEVA-PDVNIDE 434
++VL ATN PW +D A+RRR EKRIYIPLP +R + +++L T + + + + E
Sbjct: 272 ---ILVLGATNIPWVLDSAIRRRFEKRIYIPLPEPHARAAMFKLHLGTTQNSLTEADFRE 328
Query: 435 VARRTDGYSGDDLTNVCRDASLNGMR--------RKIAGKTRDEIKNMSKDDISK----D 482
+ R+TDGYSG D++ + RDA + +R +K+ G +R + ++ D ++ D
Sbjct: 329 LGRKTDGYSGADISIIVRDALMQPVRKVQSATHFKKVRGPSRADPNHLVDDLLTPCSPGD 388
Query: 483 P-------------------VAMCDFEEALVKVQRSVSQADIERHEKWFHEFG 516
P V+M D +L + +V++ D+ + +K+ +FG
Sbjct: 389 PGAIEMTWMDVPGDKLLEPVVSMSDMLRSLSNTKPTVNEHDLLKLKKFTEDFG 441
>YC97_ARCFU (O28972) Cell division cycle protein 48 homolog AF1297
Length = 733
Score = 238 bits (607), Expect = 3e-62
Identities = 140/298 (46%), Positives = 188/298 (62%), Gaps = 16/298 (5%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMFGPPGTGK 280
R+VL P V+WED+ GL AK+ L EAV PL PE F+ +P +G+L+FGPPGTGK
Sbjct: 443 REVLVEVPNVKWEDIGGLEHAKQELMEAVEWPLKYPEVFRAANIKPPRGILLFGPPGTGK 502
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKAVA E F +V L SKW GESE+ VR +F AR AP IF DEIDSL
Sbjct: 503 TLLAKAVANESNANFISVKGPELLSKWVGESEKHVREMFRKARQVAPCVIFFDEIDSLAP 562
Query: 341 SRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR--R 398
RG G+ + RV S+LL ++DG+ K V+V+AATN P ID AL R R
Sbjct: 563 RRGGIGDSHVTERVVSQLLTELDGLEEL--------KDVVVIAATNRPDMIDPALLRPGR 614
Query: 399 LEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNG 458
LE+ IYIP P+ ++R E+ +I+L+ +A DVNI+E+A +T+GYSG D+ VCR+A +
Sbjct: 615 LERHIYIPPPDKKARVEIFKIHLRGKPLADDVNIEELAEKTEGYSGADIEAVCREAGMLA 674
Query: 459 MRRKI-AGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKWFHEF 515
+R I G TR+E K +K K + FEEAL KV+ S+++ D+E++EK +F
Sbjct: 675 IRELIKPGMTREEAKEAAK----KLKITKKHFEEALKKVRPSLTKEDVEKYEKLIEDF 728
Score = 188 bits (477), Expect = 3e-47
Identities = 121/290 (41%), Positives = 176/290 (59%), Gaps = 17/290 (5%)
Query: 215 ELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLM 272
EL E +V P V +ED+ GL RL+ E + LPL PE FQ GI P KGVL+
Sbjct: 163 ELKEKPAEEVKRAVPDVTYEDIGGLKRELRLVREMIELPLKHPELFQRLGIEPP-KGVLL 221
Query: 273 FGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFI 332
+GPPGTGKTL+AKAVA E F +S + SK+ GESE+ +R +F+ A+ APS IFI
Sbjct: 222 YGPPGTGKTLIAKAVANEVDAHFIPISGPEIMSKYYGESEQRLREIFEEAKENAPSIIFI 281
Query: 333 DEIDSLCNSR-GASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDI 391
DEIDS+ R +GE E RRV ++LL +DG+ +R V+V+AATN P I
Sbjct: 282 DEIDSIAPKREEVTGEVE--RRVVAQLLALMDGLE--------ARGDVIVIAATNRPDAI 331
Query: 392 DEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTN 449
D ALRR R ++ I I +P+ E RKE++ I+ + + +A DV+++E+A T+G+ G DL
Sbjct: 332 DPALRRPGRFDREIEIGVPDKEGRKEILEIHTRKMPLAEDVDLEELAELTNGFVGADLEA 391
Query: 450 VCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRS 499
+C++A+++ +RR + + E + + + I V DF EAL ++ S
Sbjct: 392 LCKEAAMHALRR-VLPEIDIEAEEIPAEVIENLKVTREDFMEALKNIEPS 440
>CC48_SCHPO (Q9P3A7) Cell division cycle protein 48 homolog
Length = 815
Score = 211 bits (536), Expect = 5e-54
Identities = 130/305 (42%), Positives = 176/305 (57%), Gaps = 20/305 (6%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMFGPPGTGK 280
R+ + P VRWED+ GL E KR L E V +P+ E F P KGVL FGPPGTGK
Sbjct: 485 RETVVEVPNVRWEDIGGLEEVKRELRETVQMPVMYAEKFLRFGVTPSKGVLFFGPPGTGK 544
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKA+A EC F +V L S W GESE VR +FD ARA AP +F+DE+DS+
Sbjct: 545 TLLAKAIANECSANFISVKGPELLSMWFGESESNVRDIFDKARAAAPCVVFLDELDSIAK 604
Query: 341 SRGAS-GEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR-- 397
+RGAS G+ RV ++LL ++DGV+ S+K V V+ ATN P ID AL R
Sbjct: 605 ARGASAGDSGGGDRVVNQLLTEMDGVN--------SKKNVFVIGATNRPDQIDPALMRPG 656
Query: 398 RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLN 457
RL++ IY+PLP+ E+R +++ L+ VA DV++ VA+ T G+SG DL V + A
Sbjct: 657 RLDQLIYVPLPDEEARFSILQTQLRHTPVAEDVDLRAVAKATHGFSGADLEFVVQRAVKL 716
Query: 458 GMRRKIAG--KTRDEIKNMSKDDISKDPVAMCD------FEEALVKVQRSVSQADIERHE 509
++ I K +E DD+ D A EEA+ +RSVS A++ R+E
Sbjct: 717 AIKDSIEEDIKRENETGEAPADDVVMDEDASVSQVQRHHVEEAMKMARRSVSDAEVRRYE 776
Query: 510 KWFHE 514
+ H+
Sbjct: 777 AYAHQ 781
Score = 167 bits (424), Expect = 5e-41
Identities = 99/250 (39%), Positives = 153/250 (60%), Gaps = 13/250 (5%)
Query: 218 EMLERDVLETSPG-VRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMFGP 275
E + R+ E+S V ++D+ G + E V LPL P+ F+ I +P +G+LM+GP
Sbjct: 207 EPINREDEESSLAEVGYDDIGGCRRQMAQIRELVELPLRHPQLFKSIGIKPPRGILMYGP 266
Query: 276 PGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEI 335
PGTGKTL+A+AVA E G FF ++ + SK GESE +R F+ A +P+ IFIDEI
Sbjct: 267 PGTGKTLMARAVANETGAFFFLINGPEIMSKMAGESESNLRKAFEEAEKNSPAIIFIDEI 326
Query: 336 DSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEAL 395
DS+ R + E RRV S+LL +DG+ +R V+V+AATN P ID AL
Sbjct: 327 DSIAPKREKT-NGEVERRVVSQLLTLMDGMK--------ARSNVVVMAATNRPNSIDPAL 377
Query: 396 RR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRD 453
RR R ++ + + +P+ R E++RI+ K +++A DV+++++A T GY G DL ++C +
Sbjct: 378 RRFGRFDREVDVGIPDPTGRLEILRIHTKNMKLADDVDLEQIAAETHGYVGSDLASLCSE 437
Query: 454 ASLNGMRRKI 463
A++ +R K+
Sbjct: 438 AAMQQIREKM 447
>SAV_SULAC (Q07590) SAV protein
Length = 780
Score = 210 bits (534), Expect = 8e-54
Identities = 130/315 (41%), Positives = 194/315 (61%), Gaps = 25/315 (7%)
Query: 220 LERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYF--QGIRRPWKGVLMFGPPG 277
L R+V P V W D+ GL K+ L EAV PL PE F G+ P KG+L+FGPPG
Sbjct: 473 LLREVYVEVPKVNWNDIGGLDNVKQQLREAVEWPLRFPELFTKSGVTPP-KGILLFGPPG 531
Query: 278 TGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDS 337
TGKT+LAKAVATE G F V + SKW GESE+ +R +F AR AP+ IF DEIDS
Sbjct: 532 TGKTMLAKAVATESGANFIAVRGPEILSKWVGESEKAIREIFRKARQAAPTVIFFDEIDS 591
Query: 338 LCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR 397
+ RG S + + R+ ++LL ++DG+ N+ V+++AATN P +D AL R
Sbjct: 592 IAPIRGLSTDSGVTERIVNQLLAEMDGI--VPLNK------VVIIAATNRPDILDPALLR 643
Query: 398 --RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDAS 455
R ++ IY+P P+ +R E+++++ K V +A DV+++++A + +GY+G DL + R+A+
Sbjct: 644 PGRFDRLIYVPPPDKTARFEILKVHTKNVPLAEDVSLEDIAEKAEGYTGADLEALVREAT 703
Query: 456 LNGMRRKIA---GKTRDEIK-NMS------KDDISKD--PVAMCDFEEALVKVQRSVSQA 503
+N MR + ++RDE K NM K+ ++K V+ DFE+AL V+ S++QA
Sbjct: 704 INAMRSIYSMCDKQSRDECKGNMECYQKHIKECMNKTSFKVSKEDFEKALNVVKASLTQA 763
Query: 504 DIERHEKWFHEFGSA 518
DI+R+E++ E A
Sbjct: 764 DIQRYERFSKELKRA 778
Score = 176 bits (445), Expect = 2e-43
Identities = 115/292 (39%), Positives = 174/292 (59%), Gaps = 20/292 (6%)
Query: 218 EMLERDVLETS---PGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLM 272
E+ E V E+S P V WED+ L EAK+ + E V P+ PE FQ GI P KG+L+
Sbjct: 193 EIREEPVKESSLAYPKVSWEDIGDLEEAKQKIREIVEWPMRHPELFQRLGIDPP-KGILL 251
Query: 273 FGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFI 332
+GPPGTGKTLLA+A+ E G F V+ + SK+ GESE+ +R +F A APS IFI
Sbjct: 252 YGPPGTGKTLLARALRNEIGAYFITVNGPEIMSKFYGESEQRIREIFKEAEENAPSIIFI 311
Query: 333 DEIDSLCNSR-GASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDI 391
DEID++ R +GE E +RV ++LL +DG+ R V+V+ ATN P I
Sbjct: 312 DEIDAIAPKREDVTGEVE--KRVVAQLLTLMDGIK--------GRGRVIVIGATNRPDAI 361
Query: 392 DEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTN 449
D ALRR R ++ I I P+ + RK++++++ + + + DV++D++A T GY+G DL
Sbjct: 362 DPALRRPGRFDREIEIRPPDTKGRKDILQVHTRNMPITDDVDLDKLAEMTYGYTGADLAA 421
Query: 450 VCRDASLNGMRRKIAGKTRDEIKNMSKDDISKD-PVAMCDFEEALVKVQRSV 500
+ ++A++ +RR + K + + +I K+ V+M DF AL +Q S+
Sbjct: 422 LAKEAAIYALRRFVDEKKLNLDQPTIPAEIIKELKVSMNDFLNALKSIQPSL 473
>CDCH_HALSA (P46464) CdcH protein
Length = 742
Score = 202 bits (514), Expect = 2e-51
Identities = 120/315 (38%), Positives = 178/315 (56%), Gaps = 29/315 (9%)
Query: 205 KKSQYEGPDPELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYF--QG 262
K+ ++G E+ R+VL P + W+DV GLTEAK ++E+V PL PE F G
Sbjct: 433 KREDFKGALSEVEPSAMREVLVELPKITWDDVGGLTEAKNNVKESVEWPLNQPEKFTRMG 492
Query: 263 IRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLA 322
+ P GVL++GPPGTGKTL+AKAVA E F +V L SKW GESE+ +R F A
Sbjct: 493 VEPP-AGVLLYGPPGTGKTLMAKAVANETNANFISVRGPQLLSKWVGESEKAIRQTFRKA 551
Query: 323 RAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVL 382
R AP+ IF DE+DSL RG +G + S RV ++LL ++DG+ + VMV+
Sbjct: 552 RQVAPTVIFFDELDSLAPGRGQTGGNNVSERVVNQLLTELDGLE--------EMEEVMVI 603
Query: 383 AATNFPWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTD 440
AATN P ID AL R R ++ + + P E R+++++I+ + +A DV++ E+A R D
Sbjct: 604 AATNRPDIIDPALIRSGRFDRLVQVGQPGIEGREQILKIHTQDTPLAADVSLRELAERAD 663
Query: 441 GYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSV 500
GY G DL N+ R+A++ +R DD D V M F A+ V+ ++
Sbjct: 664 GYVGSDLANIAREAAIEALR----------------DDEDADDVGMAHFRAAMENVRPTI 707
Query: 501 SQADIERHEKWFHEF 515
+ +E +++ +F
Sbjct: 708 TDDLMEYYDQVEDQF 722
Score = 165 bits (417), Expect = 3e-40
Identities = 111/295 (37%), Positives = 174/295 (58%), Gaps = 17/295 (5%)
Query: 210 EGPDPELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPW 267
E D EL E T G+ +ED+ GL + + E V LP+ P+ FQ GI P
Sbjct: 165 EDTDVELREEPISGFERTGGGITYEDIGGLENEIQRVREMVELPMKHPQIFQKLGIEPP- 223
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
+GVL+ GPPGTGKTLLAKAVA E +FF+++ + SK+ GESE+ +R +F+ A+ +P
Sbjct: 224 QGVLLHGPPGTGKTLLAKAVANETSASFFSIAGPEIISKYYGESEQQLREIFEDAKDDSP 283
Query: 328 STIFIDEIDSLCNSR-GASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATN 386
S IFIDE+DS+ R +GE E RRV ++LL +DG+ R V+V+AATN
Sbjct: 284 SIIFIDELDSIAPKREDVTGEVE--RRVVAQLLTMMDGLE--------GRGQVIVIAATN 333
Query: 387 FPWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSG 444
+D ALRR R ++ I I +P+ R+E+++I+ + + ++ DVN+ +A T G+ G
Sbjct: 334 RVDAVDPALRRPGRFDREIEIGVPDEIGREEILKIHTRGMPLSDDVNLSTLADDTHGFVG 393
Query: 445 DDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRS 499
D+ ++ ++A++ +RR + DE +++ I + V DF+ AL +V+ S
Sbjct: 394 ADIESLSKEAAMRALRRYLPEIDLDE-EDIPPSLIDRMIVKREDFKGALSEVEPS 447
>CDCH_HALN1 (Q9HPF0) CdcH protein
Length = 742
Score = 202 bits (514), Expect = 2e-51
Identities = 120/315 (38%), Positives = 178/315 (56%), Gaps = 29/315 (9%)
Query: 205 KKSQYEGPDPELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYF--QG 262
K+ ++G E+ R+VL P + W+DV GLTEAK ++E+V PL PE F G
Sbjct: 433 KREDFKGALSEVEPSAMREVLVELPKITWDDVGGLTEAKNNVKESVEWPLNQPEKFTRMG 492
Query: 263 IRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLA 322
+ P GVL++GPPGTGKTL+AKAVA E F +V L SKW GESE+ +R F A
Sbjct: 493 VEPP-AGVLLYGPPGTGKTLMAKAVANETNANFISVRGPQLLSKWVGESEKAIRQTFRKA 551
Query: 323 RAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVL 382
R AP+ IF DE+DSL RG +G + S RV ++LL ++DG+ + VMV+
Sbjct: 552 RQVAPTVIFFDELDSLAPGRGQTGGNNVSERVVNQLLTELDGLE--------EMEEVMVI 603
Query: 383 AATNFPWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTD 440
AATN P ID AL R R ++ + + P E R+++++I+ + +A DV++ E+A R D
Sbjct: 604 AATNRPDIIDPALIRSGRFDRLVQVGQPGIEGREQILKIHTQDTPLAADVSLRELAERAD 663
Query: 441 GYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSV 500
GY G DL N+ R+A++ +R DD D V M F A+ V+ ++
Sbjct: 664 GYVGSDLANIAREAAIEALR----------------DDEDADDVGMAHFRAAMENVRPTI 707
Query: 501 SQADIERHEKWFHEF 515
+ +E +++ +F
Sbjct: 708 TDDLMEYYDQVEDQF 722
Score = 165 bits (417), Expect = 3e-40
Identities = 111/295 (37%), Positives = 174/295 (58%), Gaps = 17/295 (5%)
Query: 210 EGPDPELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPW 267
E D EL E T G+ +ED+ GL + + E V LP+ P+ FQ GI P
Sbjct: 165 EDTDVELREEPISGFERTGGGITYEDIGGLENEIQRVREMVELPMKHPQIFQKLGIEPP- 223
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
+GVL+ GPPGTGKTLLAKAVA E +FF+++ + SK+ GESE+ +R +F+ A+ +P
Sbjct: 224 QGVLLHGPPGTGKTLLAKAVANETSASFFSIAGPEIISKYYGESEQQLREIFEDAKDDSP 283
Query: 328 STIFIDEIDSLCNSR-GASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATN 386
S IFIDE+DS+ R +GE E RRV ++LL +DG+ R V+V+AATN
Sbjct: 284 SIIFIDELDSIAPKREDVTGEVE--RRVVAQLLTMMDGLE--------GRGQVIVIAATN 333
Query: 387 FPWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSG 444
+D ALRR R ++ I I +P+ R+E+++I+ + + ++ DVN+ +A T G+ G
Sbjct: 334 RVDAVDPALRRPGRFDREIEIGVPDEIGREEILKIHTRGMPLSDDVNLSTLADDTHGFVG 393
Query: 445 DDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRS 499
D+ ++ ++A++ +RR + DE +++ I + V DF+ AL +V+ S
Sbjct: 394 ADIESLSKEAAMRALRRYLPEIDLDE-EDIPPSLIDRMIVKREDFKGALSEVEPS 447
>VAT_THEAC (O05209) VCP-like ATPase
Length = 745
Score = 200 bits (509), Expect = 6e-51
Identities = 108/284 (38%), Positives = 171/284 (60%), Gaps = 16/284 (5%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMFGPPGTGK 280
R+V+ P V W+D+ GL + KR ++E V LPL P+ F+ + RP KG L++GPPG GK
Sbjct: 455 REVMVEVPNVHWDDIGGLEDVKREIKETVELPLLKPDVFKRLGIRPSKGFLLYGPPGVGK 514
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKAVATE F ++ + SKW GESE+ +R +F A+ AP+ +F+DEIDS+
Sbjct: 515 TLLAKAVATESNANFISIKGPEVLSKWVGESEKAIREIFKKAKQVAPAIVFLDEIDSIAP 574
Query: 341 SRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR--R 398
RG + + + R+ ++LL +DG+ V+V+ ATN P +D AL R R
Sbjct: 575 RRGTTSDSGVTERIVNQLLTSLDGIE--------VMNGVVVIGATNRPDIMDPALLRAGR 626
Query: 399 LEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNG 458
+K IYIP P+ E+R +++++ K + +APDV+++++A+RT+GY G DL N+CR+A +N
Sbjct: 627 FDKLIYIPPPDKEARLSILKVHTKNMPLAPDVDLNDIAQRTEGYVGADLENLCREAGMNA 686
Query: 459 MRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQ 502
R + D K+ + +E ++K R++S+
Sbjct: 687 YR-----ENPDATSVSQKNFLDALKTIRPSVDEEVIKFYRTLSE 725
Score = 158 bits (399), Expect = 4e-38
Identities = 108/294 (36%), Positives = 166/294 (55%), Gaps = 21/294 (7%)
Query: 215 ELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLM 272
E+ E +VLE + +ED+ GL+E + E + LPL PE F+ GI P KGV++
Sbjct: 171 EIREEPASEVLEEVSRISYEDIGGLSEQLGKIREMIELPLKHPELFERLGITPP-KGVIL 229
Query: 273 FGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFI 332
+GPPGTGKTL+A+AVA E G F +++ + SK+ G+SE+ +R +F A APS IFI
Sbjct: 230 YGPPGTGKTLIARAVANESGANFLSINGPEIMSKYYGQSEQKLREIFSKAEETAPSIIFI 289
Query: 333 DEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDID 392
DEIDS+ R + E RRV ++LL +DG+ R V+V+ ATN ID
Sbjct: 290 DEIDSIAPKR-EEVQGEVERRVVAQLLTLMDGMK--------ERGHVIVIGATNRIDAID 340
Query: 393 EALRR--RLEKRIYIPLPNFESRKELIRINLKTV-----EVAPDVNIDEVARRTDGYSGD 445
ALRR R ++ I I +P+ RKE++ I+ + + E + ++E+A T G+ G
Sbjct: 341 PALRRPGRFDREIEIGVPDRNGRKEILMIHTRNMPLGMSEEEKNKFLEEMADYTYGFVGA 400
Query: 446 DLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRS 499
DL + R++++N +RR + D K + + + K V DF+ AL ++ S
Sbjct: 401 DLAALVRESAMNALRRYLPEIDLD--KPIPTEILEKMVVTEDDFKNALKSIEPS 452
>YB56_METJA (Q58556) Cell division cycle protein 48 homolog MJ1156
Length = 903
Score = 199 bits (505), Expect = 2e-50
Identities = 111/254 (43%), Positives = 162/254 (63%), Gaps = 11/254 (4%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMFGPPGTGK 280
R+VL P V+WED+ GL E K+ L EAV PL E F+ I RP KGVL+FGPPGTGK
Sbjct: 440 REVLVEVPNVKWEDIGGLEEVKQELREAVEWPLKAKEVFEKIGVRPPKGVLLFGPPGTGK 499
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKAVA E G F +V + SKW GESE+ +R +F AR AP IF DEID++
Sbjct: 500 TLLAKAVANESGANFISVKGPEIFSKWVGESEKAIREIFRKARQSAPCIIFFDEIDAIAP 559
Query: 341 SRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR--R 398
RG + +V ++LL ++DG+ K V+V+AATN P ID AL R R
Sbjct: 560 KRGRDLSSAVTDKVVNQLLTELDGMEEP--------KDVVVIAATNRPDIIDPALLRPGR 611
Query: 399 LEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNG 458
L++ I +P+P+ ++R ++ +I+ +++ +A DVN++E+A++T+GY+G D+ +CR+A++
Sbjct: 612 LDRVILVPVPDEKARLDIFKIHTRSMNLAEDVNLEELAKKTEGYTGADIEALCREAAMLA 671
Query: 459 MRRKIAGKTRDEIK 472
+R I E+K
Sbjct: 672 VRESIGKPWDIEVK 685
Score = 182 bits (463), Expect = 1e-45
Identities = 118/291 (40%), Positives = 176/291 (59%), Gaps = 18/291 (6%)
Query: 215 ELAEMLERDVLETS-PGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVL 271
EL E ++ ET P V +ED+ GL E + + E + LP+ PE F+ GI P KGVL
Sbjct: 159 ELKEEPVSEIKETKVPDVTYEDIGGLKEEVKKVREMIELPMRHPELFEKLGIEPP-KGVL 217
Query: 272 MFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIF 331
+ GPPGTGKTLLAKAVA E G F+ ++ + SK+ GE+E +R +F+ A APS IF
Sbjct: 218 LVGPPGTGKTLLAKAVANEAGANFYVINGPEIMSKYVGETEENLRKIFEEAEENAPSIIF 277
Query: 332 IDEIDSLCNSRG-ASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWD 390
IDEID++ R A+GE E RR+ ++LL +DG+ R V+V+ ATN P
Sbjct: 278 IDEIDAIAPKRDEATGEVE--RRLVAQLLTLMDGLK--------GRGQVVVIGATNRPNA 327
Query: 391 IDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLT 448
+D ALRR R ++ I I +P+ E RKE+++I+ + + +A DV++D +A T G+ G DL
Sbjct: 328 LDPALRRPGRFDREIVIGVPDREGRKEILQIHTRNMPLAEDVDLDYLADVTHGFVGADLA 387
Query: 449 NVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRS 499
+C++A++ +RR + E + + K+ + V M DF+EAL V+ S
Sbjct: 388 ALCKEAAMRALRR-VLPSIDLEAEEIPKEVLDNLKVTMDDFKEALKDVEPS 437
Score = 31.2 bits (69), Expect = 6.8
Identities = 28/93 (30%), Positives = 47/93 (50%), Gaps = 22/93 (23%)
Query: 432 IDEVARRTDGY------SGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDISKDPVA 485
IDE+ + T S D+L N+ +D + +R K++ K ++K I K+
Sbjct: 813 IDEIIKTTQNIIQRFTTSLDELKNILKD--IESIRLKVSTK---DVK------IKKE--- 858
Query: 486 MCDFEEALVKVQRSVSQADIERHEKWFHEFGSA 518
F +AL K++ SVS+ D+ +EK E+G A
Sbjct: 859 --HFMKALEKIKPSVSKEDMRVYEKLAQEYGRA 889
>CC48_SOYBN (P54774) Cell division cycle protein 48 homolog (Valosin
containing protein homolog) (VCP)
Length = 807
Score = 199 bits (505), Expect = 2e-50
Identities = 118/301 (39%), Positives = 175/301 (57%), Gaps = 21/301 (6%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGI-RRPWKGVLMFGPPGTGK 280
R+ + P V WED+ GL KR L+E V P+ PE F+ P KGVL +GPPG GK
Sbjct: 469 RETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGK 528
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKA+A EC F +V L + W GESE VR +FD AR AP +F DE+DS+
Sbjct: 529 TLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDSIAT 588
Query: 341 SRGAS--GEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR- 397
RG+S ++ RV ++LL ++DG+S ++K V ++ ATN P ID AL R
Sbjct: 589 QRGSSVGDAGGAADRVLNQLLTEMDGMS--------AKKTVFIIGATNRPDIIDPALLRP 640
Query: 398 -RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASL 456
RL++ IYIPLP+ +SR ++ + L+ +A +V++ +AR T G+SG D+T +C+ A
Sbjct: 641 GRLDQLIYIPLPDEDSRHQIFKACLRKSPIAKNVDLRALARHTQGFSGADITEICQRACK 700
Query: 457 NGMRRKI------AGKTRDEIKNMSKDDISKD--PVAMCDFEEALVKVQRSVSQADIERH 508
+R I K+R+ + M +D + + + FEE++ +RSVS ADI ++
Sbjct: 701 YAIRENIEKDIERERKSRENPEAMDEDTVDDEVAEIKAAHFEESMKFARRSVSDADIRKY 760
Query: 509 E 509
+
Sbjct: 761 Q 761
Score = 170 bits (430), Expect = 9e-42
Identities = 105/258 (40%), Positives = 156/258 (59%), Gaps = 15/258 (5%)
Query: 212 PDPEL---AEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPW 267
PD E+ E L+R+ E V ++DV G+ + + E V LPL P+ F+ I +P
Sbjct: 183 PDTEIFCEGEPLKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPP 242
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
KG+L++GPPG+GKTL+A+AVA E G FF ++ + SK GESE +R F+ A AP
Sbjct: 243 KGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAP 302
Query: 328 STIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNF 387
S IFIDEIDS+ R + E RR+ S+LL +DG+ SR V+V+ ATN
Sbjct: 303 SIIFIDEIDSIAPKREKT-HGEVERRIVSQLLTLMDGLK--------SRAHVIVIGATNR 353
Query: 388 PWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGD 445
P ID ALRR R ++ I I +P+ R E++RI+ K ++++ DV+++ +A+ T GY G
Sbjct: 354 PNSIDPALRRFGRFDREIDIGVPDEVGRLEVLRIHTKNMKLSDDVDLERIAKDTHGYVGA 413
Query: 446 DLTNVCRDASLNGMRRKI 463
DL +C +A+L +R K+
Sbjct: 414 DLAALCTEAALQCIREKM 431
>CC48_CAPAN (Q96372) Cell division cycle protein 48 homolog
Length = 805
Score = 197 bits (502), Expect = 4e-50
Identities = 116/299 (38%), Positives = 174/299 (57%), Gaps = 19/299 (6%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGI-RRPWKGVLMFGPPGTGK 280
R+ + P V WED+ GL KR L+E V P+ PE F+ P KGVL +GPPG GK
Sbjct: 469 RETVVEVPNVSWEDIGGLENVKRELQETVQYPVEPPEKFEKFGMSPSKGVLFYGPPGCGK 528
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKA+A EC F +V L + W GESE VR +FD AR AP +F DE+DS+
Sbjct: 529 TLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDSIAT 588
Query: 341 SRGASGEHE--SSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR- 397
RG+S ++ RV ++LL ++DG++ ++K V ++ ATN P ID AL R
Sbjct: 589 QRGSSSGDAGGAADRVLNQLLTEMDGMN--------AKKTVFIIGATNRPDIIDPALLRP 640
Query: 398 -RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASL 456
RL++ IYIPLP+ +SR ++ + L+ ++ D+++ +A+ T G+SG D+T +C+ A
Sbjct: 641 GRLDQLIYIPLPDEDSRHQIFKACLRKSPLSKDIDLRALAKHTQGFSGADVTEICQRACK 700
Query: 457 NGMRRKI-----AGKTRDEIKNMSKDDISKDP-VAMCDFEEALVKVQRSVSQADIERHE 509
+R I K R E + +D+ + P + FEE++ +RSVS ADI +++
Sbjct: 701 YAIRENIEKDIEREKRRQENPDSMDEDVDEVPEIKPAHFEESMKYARRSVSDADIRKYQ 759
Score = 164 bits (416), Expect = 4e-40
Identities = 102/258 (39%), Positives = 155/258 (59%), Gaps = 15/258 (5%)
Query: 212 PDPEL---AEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPW 267
PD E+ E ++R+ E V ++DV G+ + + E V LPL P+ F+ I +P
Sbjct: 183 PDTEIFCEGEPVKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPP 242
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
KG+L++GPPG+GKTL+A+AVA E G FF ++ + SK GESE +R F+ A AP
Sbjct: 243 KGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAP 302
Query: 328 STIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNF 387
S IFIDEIDS+ R + E RR+ S+LL +DG+ SR V+V+ ATN
Sbjct: 303 SIIFIDEIDSIAPKREKT-HGEVERRIVSQLLTLMDGLK--------SRAHVIVMGATNR 353
Query: 388 PWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGD 445
P ID ALRR R ++ I I +P+ R E++ I+ K +++A +V+++ +++ T GY G
Sbjct: 354 PNSIDPALRRFGRFDREIDIGVPDEVGRLEVLGIHTKNMKLAEEVDLERISKDTHGYVGA 413
Query: 446 DLTNVCRDASLNGMRRKI 463
DL +C +A+L +R K+
Sbjct: 414 DLAALCTEAALQCIREKM 431
>TERA_XENLA (P23787) Transitional endoplasmic reticulum ATPase (TER
ATPase) (15S Mg(2+)-ATPase p97 subunit)
Length = 805
Score = 197 bits (500), Expect = 7e-50
Identities = 119/300 (39%), Positives = 175/300 (57%), Gaps = 20/300 (6%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGI-RRPWKGVLMFGPPGTGK 280
R+ + P V WED+ GL + KR L+E V P+ P+ F P KGVL +GPPG GK
Sbjct: 465 RETVVEVPQVTWEDIGGLEDVKRELQELVQYPVEHPDKFLKFGMTPSKGVLFYGPPGCGK 524
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKA+A EC F ++ L + W GESE VR +FD AR AP +F DE+DS+
Sbjct: 525 TLLAKAIANECQANFISIKGPELLTMWFGESEANVREIFDKARQAAPCVLFFDELDSIAK 584
Query: 341 SRGAS--GEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR- 397
+RG + ++ RV +++L ++DG+S +K V ++ ATN P ID A+ R
Sbjct: 585 ARGGNIGDGGGAADRVINQILTEMDGMS--------IKKNVFIIGATNRPDIIDPAILRP 636
Query: 398 -RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASL 456
RL++ IYIPLP+ +SR +++ NL+ VA DV++D +A+ T+G+SG DLT +C+ A
Sbjct: 637 GRLDQLIYIPLPDEKSRMAILKANLRKSPVAKDVDVDFLAKMTNGFSGADLTEICQRACK 696
Query: 457 NGMRRKIAG---KTRDEIKNMSKDDISKD----PVAMCDFEEALVKVQRSVSQADIERHE 509
+R I + RD N S ++ +D + FEEA+ +RSVS DI ++E
Sbjct: 697 LAIRESIENEIRRERDRQTNPSAMEVEEDDPVPEIRRDHFEEAMRLARRSVSDNDIRKYE 756
Score = 161 bits (407), Expect = 4e-39
Identities = 94/244 (38%), Positives = 149/244 (60%), Gaps = 12/244 (4%)
Query: 223 DVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMFGPPGTGKT 281
D E+ V ++D+ G + ++E V LPL P F+ I +P +G+L++GPPGTGKT
Sbjct: 193 DEEESLNEVGYDDIGGCRKQLAQIKEMVELPLRHPALFKAIGVKPPRGILLYGPPGTGKT 252
Query: 282 LLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNS 341
L+A+AVA E G FF ++ + SK GESE +R F+ A AP+ IFIDE+D++
Sbjct: 253 LIARAVANETGAFFFLINGPEIMSKLAGESESNLRKAFEEAEKNAPAIIFIDELDAIAPK 312
Query: 342 RGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR--RL 399
R + E RR+ S+LL +DG+ R V+V+AATN P ID ALRR R
Sbjct: 313 REKT-HGEVERRIVSQLLTLMDGLK--------QRAHVIVMAATNRPNSIDPALRRFGRF 363
Query: 400 EKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGM 459
++ + I +P+ R E+++I+ K ++++ DV++++VA T G+ G DL +C +A+L +
Sbjct: 364 DREVDIGIPDSTGRLEILQIHTKNMKLSDDVDLEQVANETHGHVGADLAALCSEAALQAI 423
Query: 460 RRKI 463
R+K+
Sbjct: 424 RKKM 427
>C48E_ARATH (Q9LZF6) Cell division control protein 48 homolog E
(AtCDC48e) (Transitional endoplasmic reticulum ATPase E)
Length = 810
Score = 196 bits (498), Expect = 1e-49
Identities = 118/303 (38%), Positives = 173/303 (56%), Gaps = 25/303 (8%)
Query: 222 RDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGI-RRPWKGVLMFGPPGTGK 280
R+ + P V WED+ GL KR L+E V P+ PE F+ P KGVL +GPPG GK
Sbjct: 468 RETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSPSKGVLFYGPPGCGK 527
Query: 281 TLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCN 340
TLLAKA+A EC F +V L + W GESE VR +FD AR AP +F DE+DS+
Sbjct: 528 TLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSAPCVLFFDELDSIAT 587
Query: 341 SRG--ASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR- 397
RG A ++ RV ++LL ++DG++ ++K V ++ ATN P ID AL R
Sbjct: 588 QRGNSAGDAGGAADRVLNQLLTEMDGMN--------AKKTVFIIGATNRPDIIDSALLRP 639
Query: 398 -RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASL 456
RL++ IYIPLP+ +SR + + L+ VA DV++ +A+ T G+SG D+T +C+ A
Sbjct: 640 GRLDQLIYIPLPDEDSRLNIFKACLRKSPVAKDVDVTALAKYTQGFSGADITEICQRACK 699
Query: 457 NGMRRKIAGKTRDEIK----------NMSKDDISKDPVAMCDFEEALVKVQRSVSQADIE 506
+R I +E + +M D++S+ + FEE++ +RSVS ADI
Sbjct: 700 YAIRENIEKDIENERRRSQNPEAMEEDMVDDEVSE--IRAAHFEESMKYARRSVSDADIR 757
Query: 507 RHE 509
+++
Sbjct: 758 KYQ 760
Score = 168 bits (426), Expect = 3e-41
Identities = 104/258 (40%), Positives = 156/258 (60%), Gaps = 15/258 (5%)
Query: 212 PDPEL---AEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPW 267
PD E+ E ++R+ E V ++DV G+ + + E V LPL P+ F+ I +P
Sbjct: 182 PDTEIFCEGEPVKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPP 241
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
KG+L++GPPG+GKTL+A+AVA E G FF ++ + SK GESE +R F+ A AP
Sbjct: 242 KGILLYGPPGSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAP 301
Query: 328 STIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNF 387
S IFIDEIDS+ R + E RR+ S+LL +DG+ SR V+V+ ATN
Sbjct: 302 SIIFIDEIDSIAPKREKT-NGEVERRIVSQLLTLMDGLK--------SRAHVIVMGATNR 352
Query: 388 PWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGD 445
P ID ALRR R ++ I I +P+ R E++RI+ K +++A DV+++ +++ T GY G
Sbjct: 353 PNSIDPALRRFGRFDREIDIGVPDEIGRLEVLRIHTKNMKLAEDVDLERISKDTHGYVGA 412
Query: 446 DLTNVCRDASLNGMRRKI 463
DL +C +A+L +R K+
Sbjct: 413 DLAALCTEAALQCIREKM 430
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.131 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 61,165,098
Number of Sequences: 164201
Number of extensions: 2649031
Number of successful extensions: 11537
Number of sequences better than 10.0: 812
Number of HSP's better than 10.0 without gapping: 451
Number of HSP's successfully gapped in prelim test: 364
Number of HSP's that attempted gapping in prelim test: 10064
Number of HSP's gapped (non-prelim): 1247
length of query: 518
length of database: 59,974,054
effective HSP length: 115
effective length of query: 403
effective length of database: 41,090,939
effective search space: 16559648417
effective search space used: 16559648417
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0226.8


