

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.6
(263 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
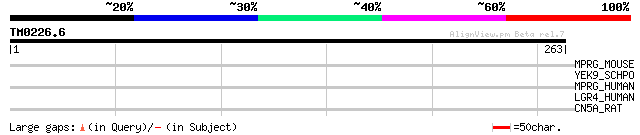
Score E
Sequences producing significant alignments: (bits) Value
MPRG_MOUSE (Q9DCU0) Membrane progestin receptor gamma (mPR gamma... 32 1.6
YEK9_SCHPO (O36021) Hypothetical protein C4F10.09c in chromosome I 31 3.5
MPRG_HUMAN (Q9NXK6) Membrane progestin receptor gamma (mPR gamma... 31 3.5
LGR4_HUMAN (Q9BXB1) Leucine-rich repeat-containing G protein-cou... 31 3.5
CN5A_RAT (O54735) cGMP-specific 3',5'-cyclic phosphodiesterase (... 30 6.0
>MPRG_MOUSE (Q9DCU0) Membrane progestin receptor gamma (mPR gamma)
(Progestin and adipoQ receptor family member V)
Length = 330
Score = 32.0 bits (71), Expect = 1.6
Identities = 16/48 (33%), Positives = 23/48 (47%)
Query: 132 RHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNN 179
RH+ + +G +N FSL Y A+ + FHE V L+V N
Sbjct: 112 RHICYFLDYGAVNLFSLGSAIAYSAYTFPDALVCSTFHECYVALAVLN 159
>YEK9_SCHPO (O36021) Hypothetical protein C4F10.09c in chromosome I
Length = 860
Score = 30.8 bits (68), Expect = 3.5
Identities = 18/52 (34%), Positives = 30/52 (57%), Gaps = 3/52 (5%)
Query: 30 IFRSAADSDAVWDRFLPSDSHSIVSQSLSRANASSKKRFYLDLSDHPVVIDN 81
IF+++A D + DR+ S S++ L+ SSK+ YL+L ++IDN
Sbjct: 417 IFQASASRDFISDRYYKSLYESLLDPRLT---TSSKQSLYLNLLYKSLIIDN 465
>MPRG_HUMAN (Q9NXK6) Membrane progestin receptor gamma (mPR gamma)
(Progestin and adipoQ receptor family member V)
Length = 330
Score = 30.8 bits (68), Expect = 3.5
Identities = 15/48 (31%), Positives = 23/48 (47%)
Query: 132 RHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNN 179
RH+ + +G +N FSL Y A+ + FH+ V L+V N
Sbjct: 112 RHICYFLDYGAVNLFSLGSAIAYSAYTFPDALMCTTFHDYYVALAVLN 159
>LGR4_HUMAN (Q9BXB1) Leucine-rich repeat-containing G
protein-coupled receptor 4 precursor (G protein-coupled
receptor 48)
Length = 951
Score = 30.8 bits (68), Expect = 3.5
Identities = 26/88 (29%), Positives = 43/88 (48%), Gaps = 9/88 (10%)
Query: 118 TTLPESRFPDVVVLRHLLWLEIHGM----INTFSLSPNTQYGAFLVFKIIDAPNFH---- 169
T++PE F +V LRH LWL+ + + ++ S P Q + KI P+F
Sbjct: 142 TSVPEDSFEGLVQLRH-LWLDDNSLTEVPVHPLSNLPTLQALTLALNKISSIPDFAFTNL 200
Query: 170 EVLVVLSVNNIGGHNITKIAYLNPDSLE 197
LVVL ++N +++ + D+LE
Sbjct: 201 SSLVVLHLHNNKIRGLSQHCFDGLDNLE 228
>CN5A_RAT (O54735) cGMP-specific 3',5'-cyclic phosphodiesterase (EC
3.1.4.17) (CGB-PDE) (cGMP-binding cGMP-specific
phosphodiesterase)
Length = 833
Score = 30.0 bits (66), Expect = 6.0
Identities = 34/127 (26%), Positives = 53/127 (40%), Gaps = 10/127 (7%)
Query: 45 LPSDSHSIVSQSLSRANASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARAL 104
LP H+ + + A S F D P+V+ + + + SGKK M
Sbjct: 41 LPQSPHADNTTPGAPARKISASEF--DRPLRPIVVKDSEGTVSFLSDSGKKEQMPLTSPR 98
Query: 105 FIAWDNGDHYRCWTTLPE--SRFPDVVVLRHLLWLEIHGMINT-----FSLSPNTQYGAF 157
F + D GD L + S DV L H ++L IHG+I+ F + ++ F
Sbjct: 99 FDS-DEGDQCSRLLELVKDISSHLDVTALCHKIFLHIHGLISADRYSLFLVCEDSSKDKF 157
Query: 158 LVFKIID 164
LV ++ D
Sbjct: 158 LVSRLFD 164
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,441,500
Number of Sequences: 164201
Number of extensions: 1297068
Number of successful extensions: 2526
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 2523
Number of HSP's gapped (non-prelim): 5
length of query: 263
length of database: 59,974,054
effective HSP length: 108
effective length of query: 155
effective length of database: 42,240,346
effective search space: 6547253630
effective search space used: 6547253630
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0226.6


