

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.2
(583 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
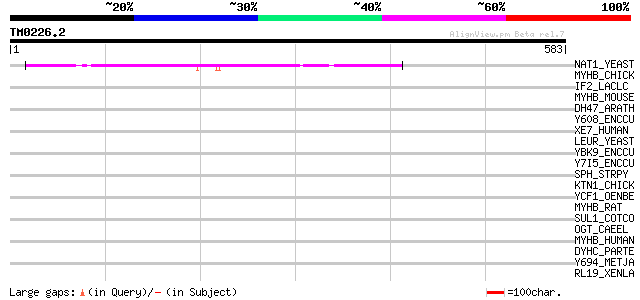
NAT1_YEAST (P12945) N-terminal acetyltransferase 1 (EC 2.3.1.88)... 184 4e-46
MYHB_CHICK (P10587) Myosin heavy chain, gizzard smooth muscle 41 0.010
IF2_LACLC (Q9X764) Translation initiation factor IF-2 39 0.037
MYHB_MOUSE (O08638) Myosin heavy chain, smooth muscle isoform (S... 38 0.064
DH47_ARATH (P31168) Dehydrin COR47 (Cold-induced COR47 protein) 38 0.084
Y608_ENCCU (Q8SVF4) Hypothetical protein ECU06_0080 37 0.14
XE7_HUMAN (Q02040) B-lymphocyte antigen precursor (B-lymphocyte ... 37 0.19
LEUR_YEAST (P08638) Regulatory protein LEU3 36 0.24
YBK9_ENCCU (Q8STZ1) Hypothetical protein ECU11_2090 36 0.32
Y7I5_ENCCU (Q8ST44) Hypothetical protein ECU07_1850/ECU10_0050 36 0.32
SPH_STRPY (P50470) Immunoglobulin G binding protein H precursor ... 35 0.41
KTN1_CHICK (Q90631) Kinectin 35 0.41
YCF1_OENBE (P31563) Hypothetical protein ycf1 (ORF 1005) (Fragment) 35 0.54
MYHB_RAT (Q63862) Myosin heavy chain, smooth muscle isoform (SMM... 35 0.54
SUL1_COTCO (Q90XB6) Extracellular sulfatase Sulf-1 precursor (EC... 35 0.71
OGT_CAEEL (O18158) UDP-N-acetylglucosamine--peptide N-acetylgluc... 35 0.71
MYHB_HUMAN (P35749) Myosin heavy chain, smooth muscle isoform (S... 35 0.71
DYHC_PARTE (Q27171) Dynein heavy chain, cytosolic (DYHC) 35 0.71
Y694_METJA (Q58105) Hypothetical protein MJ0694 34 0.92
RL19_XENLA (Q7ZYS1) 60S ribosomal protein L19 34 0.92
>NAT1_YEAST (P12945) N-terminal acetyltransferase 1 (EC 2.3.1.88)
(Amino-terminal, alpha-amino, acetyltransferase 1)
Length = 853
Score = 184 bits (468), Expect = 4e-46
Identities = 130/441 (29%), Positives = 210/441 (47%), Gaps = 60/441 (13%)
Query: 17 IPLDFLQG-DKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGK--ADILEQLILDLEHS 73
IPL FLQ ++ + Y+ P L +GVP+ FS++ LY + +LE+++LD
Sbjct: 313 IPLTFLQDKEELSKKLREYVLPQLERGVPATFSNVKPLYQRRKSKVSPLLEKIVLDY--- 369
Query: 74 IRTSGQYPGSMEKEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDLYSVK 133
SG P + P +WT + L+QH+ + A ID A++HTPT+++ Y +K
Sbjct: 370 --LSGLDP----TQDPIPFIWTNYYLSQHFLFLKDFPKAQEYIDAALDHTPTLVEFYILK 423
Query: 134 SRILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTKEGDQ 193
+RILKH G AA +E R +DL DR++N + VK L+A+ + A + A LFTK D
Sbjct: 424 ARILKHLGLMDTAAGILEEGRQLDLQDRFINCKTVKYFLRANNIDKAVEVASLFTKNDDS 483
Query: 194 HN---NLHDMQCMWYELASGESFFR--------QGDL----------------------- 219
N +LH ++ W+ + E+++R DL
Sbjct: 484 VNGIKDLHLVEASWFIVEQAEAYYRLYLDRKKKLDDLASLKKEVESDKSEQIANDIKENQ 543
Query: 220 -------GRALKKFLGVEKHYADINEDQFDFHSYCLRKMTLRTYLEMLKFQDQLHSHSYF 272
G ALK+F + K Y +DQ DFHSYC+RK T R YLEML++ L++ +
Sbjct: 544 WLVRKYKGLALKRFNAIPKFYKQFEDDQLDFHSYCMRKGTPRAYLEMLEWGKALYTKPMY 603
Query: 273 HKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNE 332
+A A + Y ++HD K ++ +E S + Q +K +A+ + A K +
Sbjct: 604 VRAMKEASKLYFQMHDDRLKRKSDSLDENSDEI--QNNGQNSSSQKKKAKKEAAAMNKRK 661
Query: 333 ELSASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSE-AIKYLKLLQKNSPDSLETHLLSF 391
E A KS + D D GEKL++ P+ + A ++ + ++L F
Sbjct: 662 ETEA----KSVAAYPSDQDNDVFGEKLIETSTPMEDFATEFYNNYSMQVREDERDYILDF 717
Query: 392 ELYTRKQKVLLTFQAVKQLLR 412
E R K+ L F ++ + +
Sbjct: 718 EFNYRIGKLALCFASLNKFAK 738
>MYHB_CHICK (P10587) Myosin heavy chain, gizzard smooth muscle
Length = 1978
Score = 40.8 bits (94), Expect = 0.010
Identities = 64/276 (23%), Positives = 118/276 (42%), Gaps = 42/276 (15%)
Query: 230 EKHYADINEDQFDFHSYCLRKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIK--LH 287
+K A++ E + C K L+ E L+ + +L++ + + A + ++ LH
Sbjct: 874 QKAEAELKELEQKHTQLCEEKNLLQ---EKLQAETELYAEAEEMRVRLAAKKQELEEILH 930
Query: 288 DSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHV 347
+ ++ EE+EE S+ L A+KKKM+Q+ E + ++ E ++L V+ GK +
Sbjct: 931 EM--EARIEEEEERSQQLQAEKKKMQQQMLDLEEQLEE-EEAARQKLQLEKVTADGK--I 985
Query: 348 KPVDPDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAV 407
K ++ D +L ++D ++ K KLL++ D L T+L E +
Sbjct: 986 KKMEDD-----ILIMEDQNNKLTKERKLLEERVSD-LTTNLAEEE------------EKA 1027
Query: 408 KQLLRLDAEHPDSHRCLVCQLHE-----------KTLFEANNSFLEKHKDSLMHRAAFAE 456
K L +L +H L +L + K E +S L + L + A +
Sbjct: 1028 KNLTKLKNKHESMISELEVRLKKEEKSRQELEKIKRKLEGESSDLHEQIAELQAQIAELK 1087
Query: 457 TLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIRE 492
A+ +E+ T+ +N AL IRE
Sbjct: 1088 AQLAKKEEELQAALARLEDETSQ---KNNALKKIRE 1120
>IF2_LACLC (Q9X764) Translation initiation factor IF-2
Length = 950
Score = 38.9 bits (89), Expect = 0.037
Identities = 30/104 (28%), Positives = 51/104 (48%), Gaps = 3/104 (2%)
Query: 248 LRKMTLRTYLEMLKFQDQLHSHSYFHKAAAG--AIRCYIKLHDSPPKSTAEEDEEMSKLL 305
L K+ LR E K + + H ++ KA AI IK + K+ E E + +++
Sbjct: 65 LLKLKLRLVPETAKSKQEDHPRTFAGKAVVEDPAILARIKEKEEAKKAAKTEAEPIEEVI 124
Query: 306 PAQKKKMRQKQRKAEARAKKGAEE-KNEELSASGVSKSGKRHVK 348
+K K+ + +K+E +A AEE K E++ A + + K VK
Sbjct: 125 TTEKPKVAEPVKKSEPKAAAKAEETKVEKVEAKANTVTPKAEVK 168
>MYHB_MOUSE (O08638) Myosin heavy chain, smooth muscle isoform (SMMHC)
Length = 1972
Score = 38.1 bits (87), Expect = 0.064
Identities = 60/248 (24%), Positives = 107/248 (42%), Gaps = 39/248 (15%)
Query: 258 EMLKFQDQLHSHSYFHKAAAGAIRCYIK--LHDSPPKSTAEEDEEMSKLLPAQKKKMRQK 315
E L+ + +L++ S + A + ++ LH+ ++ EE+E+ + L A++KKM Q+
Sbjct: 894 EQLQAETELYAESEEMRVRLAAKKQELEEILHEM--EARLEEEEDRRQQLQAERKKMAQQ 951
Query: 316 QRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLKL 375
E + ++ E ++L V+ K +K ++ D +L +DD S+ K KL
Sbjct: 952 MLDLEEQLEE-EEAARQKLQLEKVTAEAK--IKKLEDD-----ILVMDDQNSKLSKERKL 1003
Query: 376 LQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDSHRCLVCQLHEKTLFE 435
L++ D L T+L E + K L +L ++H L +L ++ E
Sbjct: 1004 LEERVSD-LTTNLAEEE------------EKAKNLTKLKSKHESMISELEVRLKKE---E 1047
Query: 436 ANNSFLEKHKDSLMHRAA-FAETLYILD----------PNRKSEAVKLIEESTNNIVPRN 484
+ LEK K L A+ F E + L ++ E + I +N
Sbjct: 1048 KSRQELEKLKRKLEGDASDFHEQIADLQAQIAELKMQLAKKEEELQAALARLDEEIAQKN 1107
Query: 485 GALGPIRE 492
AL IRE
Sbjct: 1108 NALKKIRE 1115
Score = 33.9 bits (76), Expect = 1.2
Identities = 31/125 (24%), Positives = 61/125 (48%), Gaps = 2/125 (1%)
Query: 293 STAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDP 352
+T++E+E+ +K L A ++++ AE RA+K A+ + EEL+ S R+ +
Sbjct: 1673 ATSKENEKKAKSLEADLMQLQEDLAAAE-RARKQADLEKEELAEELASSLSGRNTLQDEK 1731
Query: 353 DPHGEKLLQVDDPLSEAIKYLKLL-QKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLL 411
++ Q+++ L E ++ + + +L+ LS EL T + A +QL
Sbjct: 1732 RRLEARIAQLEEELEEEQGNMEAMSDRVRKATLQAEQLSNELATERSTAQKNESARQQLE 1791
Query: 412 RLDAE 416
R + E
Sbjct: 1792 RQNKE 1796
>DH47_ARATH (P31168) Dehydrin COR47 (Cold-induced COR47 protein)
Length = 265
Score = 37.7 bits (86), Expect = 0.084
Identities = 37/150 (24%), Positives = 66/150 (43%), Gaps = 11/150 (7%)
Query: 285 KLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGK 344
KLH S S++ DEE ++KK ++K+ KKG EK +E K+ +
Sbjct: 103 KLHRSNSSSSSSSDEE------GEEKKEKKKKIVEGEEDKKGLVEKIKEKLPGHHDKTAE 156
Query: 345 RHVKPVD---PDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVL 401
V PV P P E +++ D P E ++ +++ P + T ++
Sbjct: 157 DDV-PVSTTIPVPVSESVVEHDHPEEEKKGLVEKIKEKLPGHHDEKAEDSPAVT-STPLV 214
Query: 402 LTFQAVKQLLRLDAEHPDSHRCLVCQLHEK 431
+T V+ L EHP+ + ++ ++ EK
Sbjct: 215 VTEHPVEPTTELPVEHPEEKKGILEKIKEK 244
>Y608_ENCCU (Q8SVF4) Hypothetical protein ECU06_0080
Length = 580
Score = 37.0 bits (84), Expect = 0.14
Identities = 49/211 (23%), Positives = 85/211 (40%), Gaps = 13/211 (6%)
Query: 214 FRQGDLGRALKKFLGVEK-------HYADINED-QFDFHSYCLRKMTLRTYLEMLKFQDQ 265
F +G + +ALK LG E+ YA IN++ D H ++T ++ML ++
Sbjct: 231 FAEGSMRKALKAKLGEEEASRRGYLEYAIINDEILLDAHREHTGEVTKELVMQMLLGKNG 290
Query: 266 LHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEE---DEEMSKLLPAQKKKMRQKQRKAEAR 322
+ A ++ K + + + +E DEE +K +KKK +
Sbjct: 291 EEIDRRYIGKVANVVKERQKRREREMEKSMKELLRDEEKAKSKKGRKKKSVGVSETKKEE 350
Query: 323 AKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLKLLQKNSPD 382
++ E +EE+ S V G R K G K ++ + K + +++
Sbjct: 351 SETEEVEASEEMEISSVEVGGARR-KTGKKSKGGRKCFKIHRRVLRWRKSPEKIKEELDR 409
Query: 383 SLETHLLSFEL-YTRKQKVLLTFQAVKQLLR 412
E L Y ++QKVL V +LLR
Sbjct: 410 GREERWRGKSLEYIKEQKVLHDIAGVLELLR 440
>XE7_HUMAN (Q02040) B-lymphocyte antigen precursor (B-lymphocyte
surface antigen) (721P) (Protein XE7)
Length = 695
Score = 36.6 bits (83), Expect = 0.19
Identities = 64/260 (24%), Positives = 108/260 (40%), Gaps = 42/260 (16%)
Query: 188 TKEGDQHNNLH--DMQCMWYELA-SGESFFRQGDLGRALKKF---LGVEKHYADINEDQF 241
T G++ + +H + C W+ L SG + L + +KF V+ D ++
Sbjct: 140 TLPGERPDTIHLEGLPCKWFALKESGSEKPSEDVLVKVFEKFGEIRNVDIPMLDPYREEM 199
Query: 242 ---DFHSYCL-------RKMTLRTYLEMLKFQDQLHSHSYFHKAAAG-AIRCYIKLHDSP 290
+FH++ + R Y+ ++ L +K G A+ C IK+
Sbjct: 200 TGRNFHTFSFGGHLNFEAYVQYREYMGFIQAMSALRGMKLMYKGEDGKAVACNIKVSFDS 259
Query: 291 PK-----STAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNE------ELSASGV 339
K S + E KL ++++ QK+R+ EA ++ AEE+ + E
Sbjct: 260 TKHLSDASIKKRQLERQKLQELEQQREEQKRREKEAEERQRAEERKQKELEELERERKRE 319
Query: 340 SKSGKRHVKPVDPD--PHGEKL--LQVDD--PLSEAIKY----LKLLQKNSPDSLETHLL 389
K KR K D + + +KL LQ ++ L E IK L L Q+N L++ L
Sbjct: 320 EKLRKREQKQRDRELRRNQKKLEKLQAEEQKQLQEKIKLEERKLLLAQRN----LQSIRL 375
Query: 390 SFELYTRKQKVLLTFQAVKQ 409
EL +R + V L Q K+
Sbjct: 376 IAELLSRAKAVKLREQEQKE 395
>LEUR_YEAST (P08638) Regulatory protein LEU3
Length = 886
Score = 36.2 bits (82), Expect = 0.24
Identities = 41/161 (25%), Positives = 70/161 (43%), Gaps = 27/161 (16%)
Query: 296 EEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDP- 354
EE+EE+S +P + + R +K ++ E A + ++ P+DP+P
Sbjct: 693 EEEEELSSKVP-------ENMDSQQLRTRKFTNVRHPEKKARKIIET-----IPLDPNPI 740
Query: 355 ------HGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVK 408
G L + ++ I Y +L K SP H L + + +AV
Sbjct: 741 NAGSTSSGSSLTTPNSQVANTISYRGILNKMSPREQLNHA---NLDSSVSTDIKDTEAVN 797
Query: 409 QLLRL--DAEHPDSHRCL-VCQLHEKTL--FEANNSFLEKH 444
+ L + +AEHP + L + Q+ E TL +AN+S LE +
Sbjct: 798 EPLPIGRNAEHPANQPPLSITQMQENTLPATQANSSLLETY 838
>YBK9_ENCCU (Q8STZ1) Hypothetical protein ECU11_2090
Length = 634
Score = 35.8 bits (81), Expect = 0.32
Identities = 21/77 (27%), Positives = 40/77 (51%), Gaps = 3/77 (3%)
Query: 295 AEEDEEMSKLLPAQKKKMRQ--KQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDP 352
A+ +EE+ +++ ++++ R+ K R + R +GA E EE G K G +
Sbjct: 342 AKNEEELLRMVEREEREKREESKGRGKKKRGNRGAGESKEESKGRGKRKRGNKGAGESKE 401
Query: 353 DPHGEK-LLQVDDPLSE 368
+ GE+ ++ +DPL E
Sbjct: 402 EDRGEEGGVEAEDPLEE 418
>Y7I5_ENCCU (Q8ST44) Hypothetical protein ECU07_1850/ECU10_0050
Length = 634
Score = 35.8 bits (81), Expect = 0.32
Identities = 21/77 (27%), Positives = 40/77 (51%), Gaps = 3/77 (3%)
Query: 295 AEEDEEMSKLLPAQKKKMRQ--KQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDP 352
A+ +EE+ +++ ++++ R+ K R + R +GA E EE G K G +
Sbjct: 342 AKNEEELLRMVEREEREKREESKGRGKKKRGNRGAGESKEESKGRGKRKRGNKGAGESKE 401
Query: 353 DPHGEK-LLQVDDPLSE 368
+ GE+ ++ +DPL E
Sbjct: 402 EDRGEEGGVEAEDPLEE 418
>SPH_STRPY (P50470) Immunoglobulin G binding protein H precursor
(IgG binding protein H)
Length = 376
Score = 35.4 bits (80), Expect = 0.41
Identities = 49/215 (22%), Positives = 90/215 (41%), Gaps = 12/215 (5%)
Query: 283 YIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKS 342
Y+K D K +E +E + Q++ R+ QR+ E R ++ +++ + + +S++
Sbjct: 105 YLKKLDHEHKEHQKEQQEQEERQKNQEQLERKYQREVEKRYQEQLQKQQQLETEKQISEA 164
Query: 343 GKRHV-KPVDPDPHGEKLLQVDDPLSEA----IKYLKLLQKNSPDSLETHL-----LSFE 392
++ + + ++ +K L+ + EA +K K + S L L E
Sbjct: 165 SRKSLSRDLEASRAAKKDLEAEHQKLEAEHQKLKEDKQISDASRQGLSRDLEASRAAKKE 224
Query: 393 LYTRKQKVLLTFQAVKQLLRL-DAEHPDSHRCLVCQLHEKTLFEANNSFLEKHKDSLMHR 451
L QK+ Q +K+ ++ DA R L K EAN+ LE +L +
Sbjct: 225 LEANHQKLEAEHQKLKEDKQISDASRQGLSRDLEASRAAKKELEANHQKLEAEAKALKEQ 284
Query: 452 -AAFAETLYILDPNRKSEAVKLIEESTNNIVPRNG 485
A AE L L + S++ + N VP G
Sbjct: 285 LAKQAEELAKLRAGKASDSQTPDTKPGNKAVPGKG 319
>KTN1_CHICK (Q90631) Kinectin
Length = 1364
Score = 35.4 bits (80), Expect = 0.41
Identities = 35/135 (25%), Positives = 61/135 (44%), Gaps = 17/135 (12%)
Query: 286 LHDSPPKSTAEE---------DEEMSKLLP--AQKKKMRQKQRKAEARAKKGAEEKNEEL 334
+H+S +ST + DEE +P A + ++RK + + +K A++ +
Sbjct: 72 IHESDSESTPRDFKLSDALGTDEEQVAPVPLSATEASAGIRERKKKEKKQKAAQDDHVTK 131
Query: 335 SASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELY 394
+ G SGK+ V+P P ++ +P + K + QKN D +T S
Sbjct: 132 ESEGSKSSGKK----VEPVPVTKQPTPPSEPAAAKKKPGQKKQKN--DDQDTKTDSVASP 185
Query: 395 TRKQKVLLTFQAVKQ 409
+KQ+ +L Q VKQ
Sbjct: 186 AKKQEPVLLHQEVKQ 200
>YCF1_OENBE (P31563) Hypothetical protein ycf1 (ORF 1005) (Fragment)
Length = 1005
Score = 35.0 bits (79), Expect = 0.54
Identities = 18/57 (31%), Positives = 33/57 (57%)
Query: 292 KSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVK 348
KS E + K + +++KK++ KQ+K +++ KK ++NE S KS ++ VK
Sbjct: 774 KSKQNEIKSKQKKVKSKQKKVKSKQKKVKSKQKKVKSKQNEIKSKQNEIKSKQKKVK 830
>MYHB_RAT (Q63862) Myosin heavy chain, smooth muscle isoform (SMMHC)
(Fragments)
Length = 1327
Score = 35.0 bits (79), Expect = 0.54
Identities = 31/125 (24%), Positives = 62/125 (48%), Gaps = 2/125 (1%)
Query: 293 STAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDP 352
+T++E+E+ +K L A+ ++++ AE RA+K A+ + EEL+ S R+ +
Sbjct: 1028 ATSKENEKKAKSLEAELMQLQEDLAAAE-RARKQADLEKEELAEELASSLSGRNTLQDEK 1086
Query: 353 DPHGEKLLQVDDPLSEAIKYLKLL-QKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLL 411
++ Q+++ L E ++ + + +L+ LS EL T + A +QL
Sbjct: 1087 RRLEARIAQLEEELEEEQGNMEAMSDRVRKATLQAEQLSNELVTERSAAQKNESARQQLE 1146
Query: 412 RLDAE 416
R + E
Sbjct: 1147 RQNKE 1151
>SUL1_COTCO (Q90XB6) Extracellular sulfatase Sulf-1 precursor (EC
3.1.6.-) (QSulf1)
Length = 867
Score = 34.7 bits (78), Expect = 0.71
Identities = 22/59 (37%), Positives = 33/59 (55%), Gaps = 2/59 (3%)
Query: 270 SYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAE 328
SY++K + IK H P K A+E + SKL ++ + R+K+RK + R KKG E
Sbjct: 678 SYYNKEKGVKTQEKIKSHLHPFKEAAQEVD--SKLQLFKENRRRKKERKGKKRQKKGDE 734
>OGT_CAEEL (O18158) UDP-N-acetylglucosamine--peptide
N-acetylglucosaminyltransferase (EC 2.4.1.-) (O-GlcNAc)
(OGT)
Length = 1151
Score = 34.7 bits (78), Expect = 0.71
Identities = 34/113 (30%), Positives = 47/113 (41%), Gaps = 7/113 (6%)
Query: 99 LAQHYDRRGQYELAIAKIDEAIEHTPTVIDLYSVKSRILKHAGDFSAAAAFADEARCMD- 157
LA ++G+ AI EAI PT D YS LK GD SAA A + A ++
Sbjct: 471 LASILQQQGKLNDAILHYKEAIRIAPTFADAYSNMGNTLKEMGDSSAAIACYNRAIQINP 530
Query: 158 -LADRYVNSECVKRMLQADQVALAEKTAVLFTKEG--DQHNNL---HDMQCMW 204
AD + N + + A+ + L K D + NL H + C W
Sbjct: 531 AFADAHSNLASIHKDAGNMAEAIQSYSTALKLKPDFPDAYCNLAHCHQIICDW 583
>MYHB_HUMAN (P35749) Myosin heavy chain, smooth muscle isoform (SMMHC)
Length = 1972
Score = 34.7 bits (78), Expect = 0.71
Identities = 51/208 (24%), Positives = 89/208 (42%), Gaps = 35/208 (16%)
Query: 296 EEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPH 355
EE+E+ + L A++KKM Q+ E + ++ E ++L V+ K +K ++
Sbjct: 932 EEEEDRGQQLQAERKKMAQQMLDLEEQLEE-EEAARQKLQLEKVTAEAK--IKKLE---- 984
Query: 356 GEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDA 415
+++L +DD ++ K KLL++ D L T+L E + K L +L
Sbjct: 985 -DEILVMDDQNNKLSKERKLLEERISD-LTTNLAEEE------------EKAKNLTKLKN 1030
Query: 416 EHPDSHRCLVCQLHEKTLFEANNSFLEKHKDSLMHRAA-FAETLYILD----------PN 464
+H L +L ++ E + LEK K L A+ F E + L
Sbjct: 1031 KHESMISELEVRLKKE---EKSRQELEKLKRKLEGDASDFHEQIADLQAQIAELKMQLAK 1087
Query: 465 RKSEAVKLIEESTNNIVPRNGALGPIRE 492
++ E + + I +N AL IRE
Sbjct: 1088 KEEELQAALARLDDEIAQKNNALKKIRE 1115
Score = 33.1 bits (74), Expect = 2.1
Identities = 39/156 (25%), Positives = 73/156 (46%), Gaps = 14/156 (8%)
Query: 293 STAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDP 352
+TA+E+E+ +K L A ++++ AE RA+K A+ + EEL+ S R+ +
Sbjct: 1673 ATAKENEKKAKSLEADLMQLQEDLAAAE-RARKQADLEKEELAEELASSLSGRNALQDEK 1731
Query: 353 DPHGEKLLQVDDPLSEAIKYLKLL-QKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLL 411
++ Q+++ L E ++ + + + + LS EL T + A +QL
Sbjct: 1732 RRLEARIAQLEEELEEEQGNMEAMSDRVRKATQQAEQLSNELATERSTAQKNESARQQLE 1791
Query: 412 RLDAEHPDSHRCLVCQLHE-----KTLFEANNSFLE 442
R + E L +LHE K+ F++ + LE
Sbjct: 1792 RQNKE-------LRSKLHEMEGAVKSKFKSTIAALE 1820
>DYHC_PARTE (Q27171) Dynein heavy chain, cytosolic (DYHC)
Length = 4540
Score = 34.7 bits (78), Expect = 0.71
Identities = 33/130 (25%), Positives = 65/130 (49%), Gaps = 16/130 (12%)
Query: 251 MTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKK 310
+T R YL+ LK ++LH+ K+ + ++ + K T ++ EM K L +K
Sbjct: 3059 ITPRDYLDFLKHFEKLHNEK---KSQLEDQQLHLNVGLDKLKETEQQVLEMQKSLDQKKV 3115
Query: 311 KMRQKQRKAEAR------AKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQVDD 364
++ K+R+A + KK AE+K E+ ++ +S ++ K ++ + QV+
Sbjct: 3116 ELLTKERQAGEKLQTIIEEKKIAEKKKED--STRLSSDAEKKAKEME-----VRQSQVNK 3168
Query: 365 PLSEAIKYLK 374
L+EA+ L+
Sbjct: 3169 ELNEALPALE 3178
>Y694_METJA (Q58105) Hypothetical protein MJ0694
Length = 414
Score = 34.3 bits (77), Expect = 0.92
Identities = 16/52 (30%), Positives = 30/52 (56%)
Query: 299 EEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPV 350
EE+ + P KK ++++ KA+ + KKG +EK+++ K GK+ K +
Sbjct: 356 EEIRRKYPKPPKKKKKEKPKAKKKEKKGKKEKSKKKKDKKKDKKGKKERKVI 407
>RL19_XENLA (Q7ZYS1) 60S ribosomal protein L19
Length = 197
Score = 34.3 bits (77), Expect = 0.92
Identities = 36/137 (26%), Positives = 58/137 (42%), Gaps = 9/137 (6%)
Query: 215 RQGDLGRALKKFLGVEKHYADINEDQFDFHSYCLR----KMTLRTYLEMLKFQDQLHSHS 270
R+ L R + +G+ K N + S+ R + LR Y E K ++ HS
Sbjct: 64 RKNTLARRKGRHMGIGKRKGTANARMPEKLSWMRRMRILRRLLRRYRESKKIDRHMY-HS 122
Query: 271 YFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEK 330
+ K + L + K A D+ KLL Q + R K ++A R ++ + K
Sbjct: 123 LYLKVKGNVFKNKRILMEHIHKLKA--DKARKKLLADQAEARRSKTKEARKRREERLQAK 180
Query: 331 NEEL--SASGVSKSGKR 345
EE+ S S +SGK+
Sbjct: 181 KEEIIKSLSKEEESGKK 197
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 67,617,148
Number of Sequences: 164201
Number of extensions: 2880248
Number of successful extensions: 10914
Number of sequences better than 10.0: 91
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 10664
Number of HSP's gapped (non-prelim): 244
length of query: 583
length of database: 59,974,054
effective HSP length: 116
effective length of query: 467
effective length of database: 40,926,738
effective search space: 19112786646
effective search space used: 19112786646
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0226.2


