

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.15
(219 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
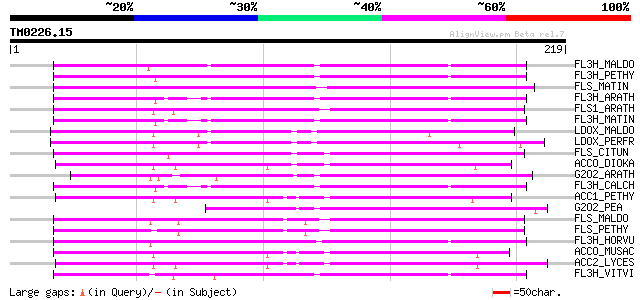
Score E
Sequences producing significant alignments: (bits) Value
FL3H_MALDO (Q06942) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 78 2e-14
FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 75 1e-13
FLS_MATIN (O04395) Flavonol synthase (EC 1.14.11.-) (FLS) (Fragm... 74 2e-13
FL3H_ARATH (Q9S818) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 74 2e-13
FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1) 74 3e-13
FL3H_MATIN (Q05965) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 74 3e-13
LDOX_MALDO (P51091) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 74 3e-13
LDOX_PERFR (O04274) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 73 5e-13
FLS_CITUN (Q9ZWQ9) Flavonol synthase (EC 1.14.11.-) (FLS) (CitFLS) 72 1e-12
ACCO_DIOKA (Q8S932) 1-aminocyclopropane-1-carboxylate oxidase (E... 72 1e-12
G2O2_ARATH (Q9XFR9) Gibberellin 2-beta-dioxygenase 2 (EC 1.14.11... 71 2e-12
FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 71 2e-12
ACC1_PETHY (Q08506) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 71 2e-12
G2O2_PEA (Q9XHM5) Gibberellin 2-beta-dioxygenase 2 (EC 1.14.11.1... 70 4e-12
FLS_MALDO (Q9XHG2) Flavonol synthase (EC 1.14.11.-) (FLS) 70 4e-12
FLS_PETHY (Q07512) Flavonol synthase (EC 1.14.11.-) (FLS) 69 7e-12
FL3H_HORVU (P28038) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 69 7e-12
ACCO_MUSAC (Q9FR99) 1-aminocyclopropane-1-carboxylate oxidase (E... 69 7e-12
ACC2_LYCES (P07920) 1-aminocyclopropane-1-carboxylate oxidase 2 ... 69 7e-12
FL3H_VITVI (P41090) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 69 9e-12
>FL3H_MALDO (Q06942) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 364
Score = 77.8 bits (190), Expect = 2e-14
Identities = 54/189 (28%), Positives = 89/189 (46%), Gaps = 6/189 (3%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGV--EKYLDEHLNSTNYFLKTMKYKG 75
WP+ ++R +S+ L L + ++ E++G+ E ++ + K
Sbjct: 147 WPDKPEAWREVTKKYSDELMGLACKLLGVLSEAMGLDTEALTKACVDMDQKVVVNFYPKC 206
Query: 76 PQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCF 135
PQ D +GL HTD +T+L Q QV GL+ DG+ WI+ +P +F+V + D
Sbjct: 207 PQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDDGKTWITVQP--VEGAFVVNLGDHG 263
Query: 136 HAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLEE 195
H SNGR + H+ ++ N +R S A F P I+ P + + E P+L P E
Sbjct: 264 HLLSNGRFKNADHQAVVNSNSSRLSIATFQNPAQEAIV-YPLSVREGEKPILEAPITYTE 322
Query: 196 FLKHSFNKE 204
K +K+
Sbjct: 323 MYKKKMSKD 331
>FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 369
Score = 75.1 bits (183), Expect = 1e-13
Identities = 51/188 (27%), Positives = 89/188 (47%), Gaps = 4/188 (2%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKY-LDEHLNSTNYFLKTMKYKGP 76
WP+ + +SE L EL + ++ E++G+EK L + + + Y
Sbjct: 149 WPDKPEGWIAVTQKYSEKLMELACKLLDVLSEAMGLEKEALTKACVDMDQKVVVNFYPKC 208
Query: 77 QTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFH 136
D +GL HTD +T+L Q QV GL+ +G+ WI+ +P +F+V + D H
Sbjct: 209 PEPDLTLGLKRHTDPGTITLLLQDQVGGLQATKDNGKTWITVQP--VEGAFVVNLGDHGH 266
Query: 137 AWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLEEF 196
SNGR + H+ ++ N +R S A F P I+ P ++ + E ++ +P E
Sbjct: 267 FLSNGRFKNADHQAVVNSNSSRLSIATFQNPAPEAIV-YPLKIREGEKSIMDEPITFAEM 325
Query: 197 LKHSFNKE 204
+ +K+
Sbjct: 326 YRRKMSKD 333
>FLS_MATIN (O04395) Flavonol synthase (EC 1.14.11.-) (FLS)
(Fragment)
Length = 291
Score = 74.3 bits (181), Expect = 2e-13
Identities = 46/190 (24%), Positives = 85/190 (44%), Gaps = 4/190 (2%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEHLNSTNYFLKTMKYKGPQ 77
WP P +R ++ H+ L + + + + E LG+ + + +N Y + Y
Sbjct: 105 WPKSPPDYREVNEEYARHVKTLSEKIMEWLSEGLGLGREAIKEVNGCWYVMNINHYPPYP 164
Query: 78 TSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHA 137
SD+ GL+ HTD LT++ +++ GL+V D + Y PS + IV + D
Sbjct: 165 HSDSFNGLEPHTDINGLTLIITNEIPGLQVFKDDHWIEVEYIPS----AIIVNIGDQIMM 220
Query: 138 WSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLEEFL 197
SNG+ + H+ + + R S + P ++ EL E+ P +KP ++++
Sbjct: 221 LSNGKYKNVLHKTTVDKEKTRMSWPVLVSPTYDMVVGPLPELTSEDDPPKFKPIAYKDYV 280
Query: 198 KHSFNKEKGK 207
+ K K
Sbjct: 281 HNKITFLKNK 290
>FL3H_ARATH (Q9S818) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Naringenin 3-dioxygenase) (Flavanone
3-hydroxylase) (FH3) (TRANSPARENT TESTA 6 protein)
Length = 358
Score = 74.3 bits (181), Expect = 2e-13
Identities = 54/195 (27%), Positives = 92/195 (46%), Gaps = 18/195 (9%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKY--------LDEHLNSTNYFLK 69
WP+ + +SE L L + +++ E++G+EK +D+ + NY+ K
Sbjct: 146 WPDKPEGWVKVTEEYSERLMSLACKLLEVLSEAMGLEKESLTNACVDMDQKI-VVNYYPK 204
Query: 70 TMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIV 129
PQ D +GL HTD +T+L Q QV GL+ +G+ WI+ +P +F+V
Sbjct: 205 C-----PQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDNGKTWITVQP--VEGAFVV 256
Query: 130 MMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYK 189
+ D H SNGR + H+ ++ N +R S A F P + P ++ + E +L +
Sbjct: 257 NLGDHGHFLSNGRFKNADHQAVVNSNSSRLSIATFQNPAPDATV-YPLKVREGEKAILEE 315
Query: 190 PFDLEEFLKHSFNKE 204
P E K ++
Sbjct: 316 PITFAEMYKRKMGRD 330
>FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1)
Length = 336
Score = 73.9 bits (180), Expect = 3e-13
Identities = 51/189 (26%), Positives = 87/189 (45%), Gaps = 7/189 (3%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEK-YLDEHLNS--TNYFLKTMKYK 74
WP P +R ++ H+ +L + + ++ + LG+++ L E L Y +K Y
Sbjct: 148 WPKNPPEYREVNEEYAVHVKKLSETLLGILSDGLGLKRDALKEGLGGEMAEYMMKINYYP 207
Query: 75 GPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDC 134
D +G+ HTD + +T+L ++V GL+V D Y PS+ IV + D
Sbjct: 208 PCPRPDLALGVPAHTDLSGITLLVPNEVPGLQVFKDDHWFDAEYIPSA----VIVHIGDQ 263
Query: 135 FHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLE 194
SNGR + HR + + R S +F P I+ EL +++P +KPF +
Sbjct: 264 ILRLSNGRYKNVLHRTTVDKEKTRMSWPVFLEPPREKIVGPLPELTGDDNPPKFKPFAFK 323
Query: 195 EFLKHSFNK 203
++ NK
Sbjct: 324 DYSYRKLNK 332
>FL3H_MATIN (Q05965) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 357
Score = 73.9 bits (180), Expect = 3e-13
Identities = 54/195 (27%), Positives = 91/195 (45%), Gaps = 18/195 (9%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKY--------LDEHLNSTNYFLK 69
WP+ + +SE L L + +++ E++G+EK +D+ + NY+ K
Sbjct: 145 WPDKPQGWAKVTEEYSEKLMGLACKLLEVLSEAMGLEKESLTNACVDMDQKI-VVNYYPK 203
Query: 70 TMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIV 129
PQ D +GL HTD +T+L Q QV GL+ DG WI+ +P +F+V
Sbjct: 204 C-----PQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDDGNTWITVQP--VEGAFVV 255
Query: 130 MMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYK 189
+ D H SNGR + H+ ++ N +R S A F P + P ++ + E ++ +
Sbjct: 256 NLGDHGHFLSNGRFKNADHQAVVNSNSSRLSIATFQNPAPEATV-YPLKVREGEKAIMEE 314
Query: 190 PFDLEEFLKHSFNKE 204
P E K ++
Sbjct: 315 PITFAEMYKRKMGRD 329
>LDOX_MALDO (P51091) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase) (Anthocyanidin synthase)
Length = 357
Score = 73.6 bits (179), Expect = 3e-13
Identities = 53/189 (28%), Positives = 91/189 (48%), Gaps = 11/189 (5%)
Query: 17 LWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEK-YLDEHLNSTNYFLKTMKY-- 73
+WP + A +++ L EL V K++ LG+++ L++ + L MK
Sbjct: 161 IWPQTPADYIEATAEYAKQLRELATKVLKVLSLGLGLDEGRLEKEVGGLEELLLQMKINY 220
Query: 74 --KGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMM 131
K PQ + +G++ HTD + LT + + V GL++ + KW++ K S ++ +
Sbjct: 221 YPKCPQP-ELALGVEAHTDVSALTFILHNMVPGLQLFYEG--KWVTAK--CVPNSIVMHI 275
Query: 132 NDCFHAWSNGRLNSPFHRVMMTGNEARYSAALF-SVPKGGYIIKAPEELVDEEHPLLYKP 190
D SNG+ S HR M+ + R S A+F PK I+K E V E+ P ++ P
Sbjct: 276 GDTLEILSNGKYKSILHRGMVNKEKVRISWAVFCEPPKEKIILKPLPETVSEDEPAMFPP 335
Query: 191 FDLEEFLKH 199
E ++H
Sbjct: 336 RTFAEHIQH 344
>LDOX_PERFR (O04274) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase)
Length = 362
Score = 73.2 bits (178), Expect = 5e-13
Identities = 57/202 (28%), Positives = 93/202 (45%), Gaps = 12/202 (5%)
Query: 17 LWPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEK-YLDEHLNSTNYFLKTMKY-- 73
+WP P + A +++ L L + ++ LG+EK L++ + + MK
Sbjct: 163 IWPTKPPDYIPATSEYAKQLRALATKILSVLSIGLGLEKGRLEKEVGGAEDLIVQMKINF 222
Query: 74 --KGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMM 131
K PQ + +G + HTD + LT + + V GL++ +D KW++ K S I+ +
Sbjct: 223 YPKCPQP-ELALGWEAHTDVSALTFILHNMVPGLQLFYED--KWVTAK--CVPNSIIMHI 277
Query: 132 NDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAP-EELVDEEHPLLYKP 190
D SNG+ S HR ++ + R S A+F P I+ P E V E P + P
Sbjct: 278 GDTLEILSNGKYKSILHRGLVNKEKVRISWAVFCEPPKEKIVLQPLPETVSEVEPPRFPP 337
Query: 191 FDLEEFLKHS-FNKEKGKRDDQ 211
+ LKH F K G D++
Sbjct: 338 RTFAQHLKHKLFRKTDGDLDEK 359
>FLS_CITUN (Q9ZWQ9) Flavonol synthase (EC 1.14.11.-) (FLS) (CitFLS)
Length = 335
Score = 71.6 bits (174), Expect = 1e-12
Identities = 49/189 (25%), Positives = 87/189 (45%), Gaps = 7/189 (3%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEHL---NSTNYFLKTMKYK 74
WP PS+R +++++ E+ + + LGVE + + + Y LK Y
Sbjct: 148 WPKNPPSYRAVNEEYAKYMREVVDKLFTYLSLGLGVEGGVLKEAAGGDDIEYMLKINYYP 207
Query: 75 GPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDC 134
D +G+ HTD + LT+L ++V GL+V D +WI K + ++ + D
Sbjct: 208 PCPRPDLALGVVAHTDLSALTVLVPNEVPGLQVFKDD--RWIDAKYIPNA--LVIHIGDQ 263
Query: 135 FHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLE 194
SNG+ + HR + ++ R S +F P ++ +LVD+E+P YK +
Sbjct: 264 IEILSNGKYKAVLHRTTVNKDKTRMSWPVFLEPPADTVVGPLPQLVDDENPPKYKAKKFK 323
Query: 195 EFLKHSFNK 203
++ NK
Sbjct: 324 DYSYCKLNK 332
>ACCO_DIOKA (Q8S932) 1-aminocyclopropane-1-carboxylate oxidase (EC
1.14.17.4) (ACC oxidase) (Ethylene-forming enzyme) (EFE)
Length = 318
Score = 71.6 bits (174), Expect = 1e-12
Identities = 54/187 (28%), Positives = 90/187 (47%), Gaps = 11/187 (5%)
Query: 19 PNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEK-YLDEHLNST---NYFLKTMKYK 74
P+ + +R + F+E L +L + + ++ E+LG+EK YL + T N+ K Y
Sbjct: 104 PDLDEEYRRVMKDFAERLEKLAEYLLDLLCENLGLEKGYLKKAFYGTKGPNFGTKVANYP 163
Query: 75 GPQTSDTKVGLDTHTDTAILTILFQS-QVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 133
+D GL HTD + +LFQ +V GL++L D +WI P S ++ + D
Sbjct: 164 PCPKADLIKGLRAHTDAGGIILLFQDDKVSGLQLLKDD--QWIDVPPMKHS--IVINLGD 219
Query: 134 CFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDE--EHPLLYKPF 191
+NG+ S HRV+ + R S A F P +I LV++ E +Y F
Sbjct: 220 QLEVITNGKYKSVLHRVVAQTDGTRMSIASFYNPGNDAVIYPAPALVEKEVEEKEVYPKF 279
Query: 192 DLEEFLK 198
++++K
Sbjct: 280 VFDDYMK 286
>G2O2_ARATH (Q9XFR9) Gibberellin 2-beta-dioxygenase 2 (EC
1.14.11.13) (Gibberellin 2-beta-hydroxylase 2)
(Gibberellin 2-oxidase 2) (GA 2-oxidase 2)
Length = 341
Score = 71.2 bits (173), Expect = 2e-12
Identities = 55/192 (28%), Positives = 86/192 (44%), Gaps = 16/192 (8%)
Query: 25 FRYALCSFSEHLSELDQIVRKMVLESLGVE------KYL-DEHLNSTNYFLKTMKYKGPQ 77
FR ++ + + + E+ V +MV E LG+E K L DE +S L+ Y +
Sbjct: 137 FRESVEEYMKEIKEVSYKVLEMVAEELGIEPRDTLSKMLRDEKSDSC---LRLNHYPAAE 193
Query: 78 TSD---TKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDC 134
KVG HTD I+++L + GL++ KDG W++ P + SF + + D
Sbjct: 194 EEAEKMVKVGFGEHTDPQIISVLRSNNTAGLQICVKDG-SWVAVPP--DHSSFFINVGDA 250
Query: 135 FHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLE 194
+NGR S HRV+ +R S F P I LV E+ LYK F
Sbjct: 251 LQVMTNGRFKSVKHRVLADTRRSRISMIYFGGPPLSQKIAPLPCLVPEQDDWLYKEFTWS 310
Query: 195 EFLKHSFNKEKG 206
++ ++ + G
Sbjct: 311 QYKSSAYKSKLG 322
>FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 356
Score = 70.9 bits (172), Expect = 2e-12
Identities = 54/195 (27%), Positives = 92/195 (46%), Gaps = 18/195 (9%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKY--------LDEHLNSTNYFLK 69
WP+ +R +S+ L L + +++ E++G+EK +D+ + NY+ K
Sbjct: 144 WPDKPNEWRAVTEEYSKVLMGLACKLLEVLSEAMGLEKEALTKACVDMDQKV-VVNYYPK 202
Query: 70 TMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIV 129
PQ D +GL HTD +T+L Q QV GL+ G WI+ KP +F+V
Sbjct: 203 C-----PQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDGGESWITVKP--VEGAFVV 254
Query: 130 MMNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYK 189
+ D H SNGR + H+ ++ + +R S A F P I+ P ++ + E ++ +
Sbjct: 255 NLGDHGHYLSNGRFKNADHQAVVNSSTSRLSIATFQNPAPEAIV-YPLKINEGEKSIMEE 313
Query: 190 PFDLEEFLKHSFNKE 204
P E K + +
Sbjct: 314 PMTFMEMYKKKMSTD 328
>ACC1_PETHY (Q08506) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE)
Length = 319
Score = 70.9 bits (172), Expect = 2e-12
Identities = 57/188 (30%), Positives = 92/188 (48%), Gaps = 12/188 (6%)
Query: 19 PNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEK-YLDEHLNST---NYFLKTMKYK 74
P+ + +R + F++ L +L + + ++ E+LG+EK YL + N+ K Y
Sbjct: 104 PDLDEEYREVMRDFAKRLEKLAEELLDLLCENLGLEKGYLKNAFYGSKGPNFGTKVSNYP 163
Query: 75 GPQTSDTKVGLDTHTDTAILTILFQS-QVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 133
D GL HTD + +LFQ +V GL++L KDG+ WI P S +V + D
Sbjct: 164 PCPKPDLIKGLRAHTDAGGIILLFQDDKVSGLQLL-KDGQ-WIDVPPMRHS--IVVNLGD 219
Query: 134 CFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVD---EEHPLLYKP 190
+NG+ S HRV+ + AR S A F P +I LV+ EE+ +Y
Sbjct: 220 QLEVITNGKYKSVMHRVIAQKDGARMSLASFYNPGSDAVIYPAPALVEKEAEENKQVYPK 279
Query: 191 FDLEEFLK 198
F ++++K
Sbjct: 280 FVFDDYMK 287
>G2O2_PEA (Q9XHM5) Gibberellin 2-beta-dioxygenase 2 (EC 1.14.11.13)
(Gibberellin 2-beta-hydroxylase 2) (Gibberellin
2-oxidase 2) (GA 2-oxidase 2)
Length = 345
Score = 70.1 bits (170), Expect = 4e-12
Identities = 43/136 (31%), Positives = 67/136 (48%), Gaps = 4/136 (2%)
Query: 78 TSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHA 137
+++ +G H+D ILTIL + V GL++ T G WI P + F VM+ D
Sbjct: 199 SNNNNIGFGEHSDPQILTILRSNNVGGLQISTHHGL-WIPVPP--DPSEFYVMVGDALQV 255
Query: 138 WSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLEEFL 197
+NGR S HRV+ + R S F+ P ++I ++V P LY+PF ++
Sbjct: 256 LTNGRFVSVRHRVLTNTTKPRMSMMYFAAPPLNWLISPLSKMVTAHSPCLYRPFTWAQYK 315
Query: 198 KHSFNKEKG-KRDDQF 212
+ ++ G R DQF
Sbjct: 316 QAAYALRLGDTRLDQF 331
>FLS_MALDO (Q9XHG2) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 337
Score = 70.1 bits (170), Expect = 4e-12
Identities = 51/190 (26%), Positives = 91/190 (47%), Gaps = 9/190 (4%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVE-KYLDEHLNSTN--YFLKTMKYK 74
WP PS+R A +++HL + + + +++ LG+E + L + N Y LK Y
Sbjct: 150 WPKNPPSYREANEEYAKHLHNVVEKLFRLLSLGLGLEGQELKKAAGGDNLEYLLKINYYP 209
Query: 75 GPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKW-ISYKPSSESQSFIVMMND 133
D +G+ HTD + +TIL + V GL+ KDGR + + Y P++ ++ + D
Sbjct: 210 PCPRPDLALGVVAHTDMSTVTILVPNDVQGLQAC-KDGRWYDVKYIPNA----LVIHIGD 264
Query: 134 CFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDL 193
SNG+ S HR + ++ R S +F P +++ +LV+ + YK
Sbjct: 265 QMEIMSNGKYTSVLHRTTVNKDKTRISWPVFLEPPADHVVGPHPQLVNAVNQPKYKTKKY 324
Query: 194 EEFLKHSFNK 203
+++ NK
Sbjct: 325 GDYVYCKINK 334
>FLS_PETHY (Q07512) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 348
Score = 69.3 bits (168), Expect = 7e-12
Identities = 52/192 (27%), Positives = 91/192 (47%), Gaps = 13/192 (6%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEHLNSTN-----YFLKTMK 72
WP PS+R A + + + E+ + K + LG+E + E + + Y LK
Sbjct: 161 WPKNPPSYREANEEYGKRMREVVDRIFKSLSLGLGLEGH--EMIEAAGGDEIVYLLKINY 218
Query: 73 YKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKW-ISYKPSSESQSFIVMM 131
Y D +G+ HTD + +TIL ++V GL+V KDG + + Y P++ IV +
Sbjct: 219 YPPCPRPDLALGVVAHTDMSYITILVPNEVQGLQVF-KDGHWYDVKYIPNA----LIVHI 273
Query: 132 NDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPF 191
D SNG+ S +HR + ++ R S +F P + + +L+ E +P +K
Sbjct: 274 GDQVEILSNGKYKSVYHRTTVNKDKTRMSWPVFLEPPSEHEVGPIPKLLSEANPPKFKTK 333
Query: 192 DLEEFLKHSFNK 203
++++ NK
Sbjct: 334 KYKDYVYCKLNK 345
>FL3H_HORVU (P28038) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 377
Score = 69.3 bits (168), Expect = 7e-12
Identities = 49/188 (26%), Positives = 85/188 (45%), Gaps = 4/188 (2%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVE-KYLDEHLNSTNYFLKTMKYKGP 76
WP + + +SE L L + ++ E++G+E + L + + + Y
Sbjct: 149 WPEKPAGWCAVVERYSERLMGLSCNLMGVLSEAMGLETEALAKACVDMDQKVVVNFYPRC 208
Query: 77 QTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFH 136
D +GL HTD +T+L Q V GL+ G+ WI+ +P S +F+V + D H
Sbjct: 209 PQPDLTLGLKRHTDPGTITLLLQDLVGGLQATRDGGKNWITVQPI--SGAFVVNLGDHGH 266
Query: 137 AWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLEEF 196
SNGR + H+ ++ G +R S A F P + P + + E P+L +P E
Sbjct: 267 FMSNGRFKNADHQAVVNGESSRLSIATFQNPAPDARV-WPLAVREGEEPILEEPITFTEM 325
Query: 197 LKHSFNKE 204
+ ++
Sbjct: 326 YRRKMERD 333
>ACCO_MUSAC (Q9FR99) 1-aminocyclopropane-1-carboxylate oxidase (EC
1.14.17.4) (ACC oxidase) (Ethylene-forming enzyme) (EFE)
Length = 306
Score = 69.3 bits (168), Expect = 7e-12
Identities = 56/189 (29%), Positives = 87/189 (45%), Gaps = 13/189 (6%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEK------YLDEHLNSTNYFLKTM 71
WP+ P F + + E + +L + + +++ E+LG EK + + + + K
Sbjct: 101 WPSNPPDFEETMKEYREEIRKLAEKMMEVMDENLGFEKGCIKKAFSGDGQHPPFFGTKVS 160
Query: 72 KYKGPQTSDTKVGLDTHTDTAILTILFQS-QVDGLEVLTKDGRKWISYKPSSESQSFIVM 130
Y D GL HTD + +LFQ QV GL++L KDGR WI +P +++ ++
Sbjct: 161 HYPPCPRLDLVKGLRAHTDAGGVILLFQDDQVGGLQML-KDGR-WIDVQPLADA--IVIN 216
Query: 131 MNDCFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEE--HPLLY 188
D SNGR S +HRV+ T + R S A F P I EE P LY
Sbjct: 217 TGDQIEVLSNGRYKSAWHRVLATSHGNRRSIASFYNPSLKATIAPAAGAATEEAAPPALY 276
Query: 189 KPFDLEEFL 197
F +++
Sbjct: 277 PKFLFGDYM 285
>ACC2_LYCES (P07920) 1-aminocyclopropane-1-carboxylate oxidase 2 (EC
1.14.17.4) (ACC oxidase 2) (Ethylene-forming enzyme)
(EFE) (Protein GTOMA)
Length = 316
Score = 69.3 bits (168), Expect = 7e-12
Identities = 59/203 (29%), Positives = 96/203 (47%), Gaps = 13/203 (6%)
Query: 19 PNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEK-YLDEHLNST---NYFLKTMKYK 74
P+ + +R + F + L +L + + ++ E+LG+EK YL + N+ K Y
Sbjct: 104 PDLDDVYREVMRDFRKRLEKLAEELLDLLCENLGLEKSYLKNTFYGSKGPNFGTKVSNYP 163
Query: 75 GPQTSDTKVGLDTHTDTAILTILFQS-QVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 133
D GL HTD + +LFQ +V GL++L KDGR WI P S +V + D
Sbjct: 164 PCPKPDLIKGLRAHTDAGGIILLFQDDKVSGLQLL-KDGR-WIDVPPMRHS--IVVNLGD 219
Query: 134 CFHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEE----HPLLYK 189
+NG+ S HRV+ + R S A F P +I LVD+E + +Y
Sbjct: 220 QLEVITNGKYKSVMHRVIAQKDGTRMSLASFYNPGNDALIYPAPALVDKEAEEHNKQVYP 279
Query: 190 PFDLEEFLKHSFNKEKGKRDDQF 212
F ++++K N + ++ +F
Sbjct: 280 KFMFDDYMKLYANLKFQAKEPRF 302
>FL3H_VITVI (P41090) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 364
Score = 68.9 bits (167), Expect = 9e-12
Identities = 52/190 (27%), Positives = 89/190 (46%), Gaps = 8/190 (4%)
Query: 18 WPNGNPSFRYALCSFSEHLSELDQIVRKMVLESLGVEKYLDEHLNS-TNYFLKTMKYKGP 76
WP+ +R +SE L L + +++ E++ ++K D N+ + K + P
Sbjct: 147 WPDKPEGWRSVTQEYSEKLMGLACKLLEVLSEAMDLDK--DALTNACVDMDQKVVVNFYP 204
Query: 77 QTS--DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDC 134
Q D +GL HTD +T+L Q QV GL+ G+ WI+ +P +F+V + D
Sbjct: 205 QCPQPDLTLGLKRHTDPGTITLLLQDQVGGLQATRDGGKTWITVQP--VEGAFVVNLGDH 262
Query: 135 FHAWSNGRLNSPFHRVMMTGNEARYSAALFSVPKGGYIIKAPEELVDEEHPLLYKPFDLE 194
H SNGR + H+ ++ N +R S A F P + P ++ + E +L P
Sbjct: 263 GHYLSNGRFKNADHQAVVNSNHSRLSIATFQNPAPEATV-YPLKIREGEKAVLEGPITFA 321
Query: 195 EFLKHSFNKE 204
E + +K+
Sbjct: 322 EMYRRKMSKD 331
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,470,890
Number of Sequences: 164201
Number of extensions: 1156128
Number of successful extensions: 2895
Number of sequences better than 10.0: 86
Number of HSP's better than 10.0 without gapping: 67
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 2742
Number of HSP's gapped (non-prelim): 86
length of query: 219
length of database: 59,974,054
effective HSP length: 106
effective length of query: 113
effective length of database: 42,568,748
effective search space: 4810268524
effective search space used: 4810268524
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0226.15


