

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.11
(231 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
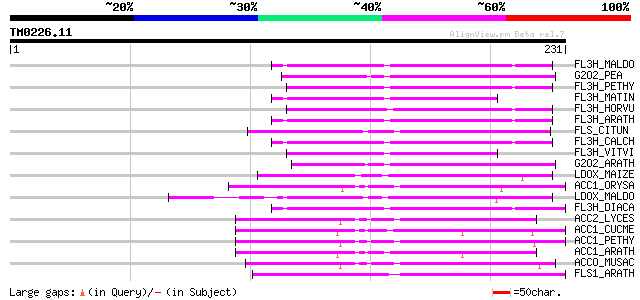
Score E
Sequences producing significant alignments: (bits) Value
FL3H_MALDO (Q06942) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 69 9e-12
G2O2_PEA (Q9XHM5) Gibberellin 2-beta-dioxygenase 2 (EC 1.14.11.1... 65 2e-10
FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 64 4e-10
FL3H_MATIN (Q05965) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 64 4e-10
FL3H_HORVU (P28038) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 63 5e-10
FL3H_ARATH (Q9S818) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 63 5e-10
FLS_CITUN (Q9ZWQ9) Flavonol synthase (EC 1.14.11.-) (FLS) (CitFLS) 63 7e-10
FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 62 9e-10
FL3H_VITVI (P41090) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 61 2e-09
G2O2_ARATH (Q9XFR9) Gibberellin 2-beta-dioxygenase 2 (EC 1.14.11... 59 7e-09
LDOX_MAIZE (P41213) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 59 1e-08
ACC1_ORYSA (Q40634) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 59 1e-08
LDOX_MALDO (P51091) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 59 1e-08
FL3H_DIACA (Q05964) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 59 1e-08
ACC2_LYCES (P07920) 1-aminocyclopropane-1-carboxylate oxidase 2 ... 59 1e-08
ACC1_CUCME (Q04644) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 59 1e-08
ACC1_PETHY (Q08506) 1-aminocyclopropane-1-carboxylate oxidase 1 ... 58 2e-08
ACC1_ARATH (Q06588) 1-aminocyclopropane-1-carboxylate oxidase (E... 58 2e-08
ACCO_MUSAC (Q9FR99) 1-aminocyclopropane-1-carboxylate oxidase (E... 57 3e-08
FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1) 57 4e-08
>FL3H_MALDO (Q06942) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 364
Score = 68.9 bits (167), Expect = 9e-12
Identities = 42/117 (35%), Positives = 61/117 (51%), Gaps = 4/117 (3%)
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 169
K PQ D +GL HTD +T+L Q QV GL+ DG+ WI+ +P +F+V + D
Sbjct: 205 KCPQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDDGKTWITVQP--VEGAFVVNLGD 261
Query: 170 CFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
H SNGR + H+ ++ N +R S A F P I+ P + + E P+L P
Sbjct: 262 HGHLLSNGRFKNADHQAVVNSNSSRLSIATFQNPAQEAIV-YPLSVREGEKPILEAP 317
>G2O2_PEA (Q9XHM5) Gibberellin 2-beta-dioxygenase 2 (EC 1.14.11.13)
(Gibberellin 2-beta-hydroxylase 2) (Gibberellin
2-oxidase 2) (GA 2-oxidase 2)
Length = 345
Score = 64.7 bits (156), Expect = 2e-10
Identities = 38/114 (33%), Positives = 55/114 (47%), Gaps = 3/114 (2%)
Query: 114 TSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHA 173
+++ +G H+D ILTIL + V GL++ T G WI P + F VM+ D
Sbjct: 199 SNNNNIGFGEHSDPQILTILRSNNVGGLQISTHHGL-WIPVPP--DPSEFYVMVGDALQV 255
Query: 174 WSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
+NGR S HRV+ + R S F P +I ++V P LY+PF
Sbjct: 256 LTNGRFVSVRHRVLTNTTKPRMSMMYFAAPPLNWLISPLSKMVTAHSPCLYRPF 309
>FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 369
Score = 63.5 bits (153), Expect = 4e-10
Identities = 36/111 (32%), Positives = 59/111 (52%), Gaps = 3/111 (2%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
D +GL HTD +T+L Q QV GL+ +G+ WI+ +P +F+V + D H S
Sbjct: 212 DLTLGLKRHTDPGTITLLLQDQVGGLQATKDNGKTWITVQP--VEGAFVVNLGDHGHFLS 269
Query: 176 NGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
NGR + H+ ++ N +R S A F P I+ P ++ + E ++ +P
Sbjct: 270 NGRFKNADHQAVVNSNSSRLSIATFQNPAPEAIV-YPLKIREGEKSIMDEP 319
>FL3H_MATIN (Q05965) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 357
Score = 63.5 bits (153), Expect = 4e-10
Identities = 36/94 (38%), Positives = 50/94 (52%), Gaps = 3/94 (3%)
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 169
K PQ D +GL HTD +T+L Q QV GL+ DG WI+ +P +F+V + D
Sbjct: 203 KCPQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDDGNTWITVQP--VEGAFVVNLGD 259
Query: 170 CFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVP 203
H SNGR + H+ ++ N +R S A F P
Sbjct: 260 HGHFLSNGRFKNADHQAVVNSNSSRLSIATFQNP 293
>FL3H_HORVU (P28038) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 377
Score = 63.2 bits (152), Expect = 5e-10
Identities = 37/111 (33%), Positives = 57/111 (51%), Gaps = 3/111 (2%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
D +GL HTD +T+L Q V GL+ G+ WI+ +P S +F+V + D H S
Sbjct: 212 DLTLGLKRHTDPGTITLLLQDLVGGLQATRDGGKNWITVQPI--SGAFVVNLGDHGHFMS 269
Query: 176 NGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
NGR + H+ ++ G +R S A F P + P + + E P+L +P
Sbjct: 270 NGRFKNADHQAVVNGESSRLSIATFQNPAPDARV-WPLAVREGEEPILEEP 319
>FL3H_ARATH (Q9S818) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Naringenin 3-dioxygenase) (Flavanone
3-hydroxylase) (FH3) (TRANSPARENT TESTA 6 protein)
Length = 358
Score = 63.2 bits (152), Expect = 5e-10
Identities = 39/117 (33%), Positives = 61/117 (51%), Gaps = 4/117 (3%)
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 169
K PQ D +GL HTD +T+L Q QV GL+ +G+ WI+ +P +F+V + D
Sbjct: 204 KCPQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDNGKTWITVQP--VEGAFVVNLGD 260
Query: 170 CFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
H SNGR + H+ ++ N +R S A F P + P ++ + E +L +P
Sbjct: 261 HGHFLSNGRFKNADHQAVVNSNSSRLSIATFQNPAPDATV-YPLKVREGEKAILEEP 316
>FLS_CITUN (Q9ZWQ9) Flavonol synthase (EC 1.14.11.-) (FLS) (CitFLS)
Length = 335
Score = 62.8 bits (151), Expect = 7e-10
Identities = 38/126 (30%), Positives = 61/126 (48%), Gaps = 4/126 (3%)
Query: 100 ITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSE 159
I Y LK Y D +G+ HTD + LT+L ++V GL+V D +WI K
Sbjct: 197 IEYMLKINYYPPCPRPDLALGVVAHTDLSALTVLVPNEVPGLQVFKDD--RWIDAKYIPN 254
Query: 160 SQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEE 219
+ ++ + D SNG+ + HR + ++ R S +F P ++ +LVD+E
Sbjct: 255 A--LVIHIGDQIEILSNGKYKAVLHRTTVNKDKTRMSWPVFLEPPADTVVGPLPQLVDDE 312
Query: 220 HPLLYK 225
+P YK
Sbjct: 313 NPPKYK 318
>FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 356
Score = 62.4 bits (150), Expect = 9e-10
Identities = 39/117 (33%), Positives = 60/117 (50%), Gaps = 4/117 (3%)
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 169
K PQ D +GL HTD +T+L Q QV GL+ G WI+ KP +F+V + D
Sbjct: 202 KCPQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDGGESWITVKP--VEGAFVVNLGD 258
Query: 170 CFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
H SNGR + H+ ++ + +R S A F P I+ P ++ + E ++ +P
Sbjct: 259 HGHYLSNGRFKNADHQAVVNSSTSRLSIATFQNPAPEAIV-YPLKINEGEKSIMEEP 314
>FL3H_VITVI (P41090) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 364
Score = 61.2 bits (147), Expect = 2e-09
Identities = 32/88 (36%), Positives = 47/88 (53%), Gaps = 2/88 (2%)
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
D +GL HTD +T+L Q QV GL+ G+ WI+ +P +F+V + D H S
Sbjct: 210 DLTLGLKRHTDPGTITLLLQDQVGGLQATRDGGKTWITVQP--VEGAFVVNLGDHGHYLS 267
Query: 176 NGRLHSPFHRVMMTGNEARYSAALFFVP 203
NGR + H+ ++ N +R S A F P
Sbjct: 268 NGRFKNADHQAVVNSNHSRLSIATFQNP 295
>G2O2_ARATH (Q9XFR9) Gibberellin 2-beta-dioxygenase 2 (EC
1.14.11.13) (Gibberellin 2-beta-hydroxylase 2)
(Gibberellin 2-oxidase 2) (GA 2-oxidase 2)
Length = 341
Score = 59.3 bits (142), Expect = 7e-09
Identities = 37/110 (33%), Positives = 52/110 (46%), Gaps = 3/110 (2%)
Query: 118 KVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNG 177
KVG HTD I+++L + GL++ KDG W++ P + SF + + D +NG
Sbjct: 201 KVGFGEHTDPQIISVLRSNNTAGLQICVKDG-SWVAVPP--DHSSFFINVGDALQVMTNG 257
Query: 178 RLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
R S HRV+ +R S F P I LV E+ LYK F
Sbjct: 258 RFKSVKHRVLADTRRSRISMIYFGGPPLSQKIAPLPCLVPEQDDWLYKEF 307
>LDOX_MAIZE (P41213) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase)
Length = 395
Score = 58.9 bits (141), Expect = 1e-08
Identities = 39/124 (31%), Positives = 59/124 (47%), Gaps = 5/124 (4%)
Query: 104 LKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSF 163
LK Y + VG++ HTD + L+ + + V GL+VL G +W++ + E +
Sbjct: 234 LKINYYPRCPQPELAVGVEAHTDVSALSFILHNGVPGLQVL--HGARWVTAR--HEPGTI 289
Query: 164 IVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAP-EELVDEEHPL 222
IV + D SNGR S HR ++ R S +F P ++ P ELV E HP
Sbjct: 290 IVHVGDALEILSNGRYTSVLHRGLVNREAVRISWVVFCEPPPDSVLLHPLPELVTEGHPA 349
Query: 223 LYKP 226
+ P
Sbjct: 350 RFTP 353
>ACC1_ORYSA (Q40634) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE)
Length = 322
Score = 58.9 bits (141), Expect = 1e-08
Identities = 48/142 (33%), Positives = 66/142 (45%), Gaps = 6/142 (4%)
Query: 92 SFRYASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRK 150
+FR + T+ K Y D GL HTD + +LFQ V GL++L KDG +
Sbjct: 151 AFRGPAGAPTFGTKVSSYPPCPRPDLVKGLRAHTDRGGIILLFQDDSVGGLQLL-KDG-E 208
Query: 151 WISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVP-KGGCII 209
W+ P S +V + D +NGR S HRV+ + R S A F+ P G I
Sbjct: 209 WVDVPPMRHS--IVVNLGDQLEVITNGRYKSVMHRVVAQTDGNRMSIASFYNPGSDGVIS 266
Query: 210 KAPEELVDEEHPLLYKPFDRED 231
AP + +EE + Y F ED
Sbjct: 267 PAPALVKEEEAVVAYPKFVFED 288
>LDOX_MALDO (P51091) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase) (Anthocyanidin synthase)
Length = 357
Score = 58.5 bits (140), Expect = 1e-08
Identities = 44/161 (27%), Positives = 76/161 (46%), Gaps = 21/161 (13%)
Query: 67 MGIDDANVYEKVESLTKILWPNGNPSFRYASQPITYFLKTMKYKGPQTSDTKVGLDTHTD 126
+G+D+ + ++V L ++L I Y+ K PQ + +G++ HTD
Sbjct: 195 LGLDEGRLEKEVGGLEELL----------LQMKINYYPKC-----PQP-ELALGVEAHTD 238
Query: 127 TAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRV 186
+ LT + + V GL++ + KW++ K S ++ + D SNG+ S HR
Sbjct: 239 VSALTFILHNMVPGLQLFYEG--KWVTAK--CVPNSIVMHIGDTLEILSNGKYKSILHRG 294
Query: 187 MMTGNEARYSAALFF-VPKGGCIIKAPEELVDEEHPLLYKP 226
M+ + R S A+F PK I+K E V E+ P ++ P
Sbjct: 295 MVNKEKVRISWAVFCEPPKEKIILKPLPETVSEDEPAMFPP 335
>FL3H_DIACA (Q05964) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 365
Score = 58.5 bits (140), Expect = 1e-08
Identities = 38/122 (31%), Positives = 60/122 (49%), Gaps = 4/122 (3%)
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 169
K PQ D +GL HTD +T+L Q QV GL+ G+ WI+ +P +F+V + D
Sbjct: 205 KCPQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDGGKTWITVQP--VPGAFVVNLGD 261
Query: 170 CFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDR 229
H SNGR + H+ ++ +R S A F P + P + + E+ ++ +P
Sbjct: 262 HGHFLSNGRFKNADHQAVVNSECSRLSIATFQNPSPDATV-YPLAIREGENSIMEEPITF 320
Query: 230 ED 231
D
Sbjct: 321 AD 322
>ACC2_LYCES (P07920) 1-aminocyclopropane-1-carboxylate oxidase 2 (EC
1.14.17.4) (ACC oxidase 2) (Ethylene-forming enzyme)
(EFE) (Protein GTOMA)
Length = 316
Score = 58.5 bits (140), Expect = 1e-08
Identities = 43/126 (34%), Positives = 59/126 (46%), Gaps = 5/126 (3%)
Query: 95 YASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQS-QVDGLEVLTKDGRKWIS 153
Y S+ + K Y D GL HTD + +LFQ +V GL++L KDGR WI
Sbjct: 148 YGSKGPNFGTKVSNYPPCPKPDLIKGLRAHTDAGGIILLFQDDKVSGLQLL-KDGR-WID 205
Query: 154 YKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPE 213
P S +V + D +NG+ S HRV+ + R S A F+ P +I
Sbjct: 206 VPPMRHS--IVVNLGDQLEVITNGKYKSVMHRVIAQKDGTRMSLASFYNPGNDALIYPAP 263
Query: 214 ELVDEE 219
LVD+E
Sbjct: 264 ALVDKE 269
>ACC1_CUCME (Q04644) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE) (PMEL1)
Length = 318
Score = 58.5 bits (140), Expect = 1e-08
Identities = 49/143 (34%), Positives = 64/143 (44%), Gaps = 10/143 (6%)
Query: 95 YASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQ-SQVDGLEVLTKDGRKWIS 153
Y S+ T+ K Y D GL HTD + +LFQ +V GL++L KDG WI
Sbjct: 148 YGSKGPTFGTKVSNYPPCPKPDLIKGLRAHTDAGGIILLFQDDKVSGLQLL-KDG-NWID 205
Query: 154 YKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVM-MTGNEARYSAALFFVPKGGCIIKAP 212
P + +V + D +NGR S HRV+ T R S A F+ P +I
Sbjct: 206 VPPM--RHAIVVNLGDQLEVITNGRYKSVMHRVLTQTSGTGRMSIASFYNPGSDAVIYPA 263
Query: 213 EELV----DEEHPLLYKPFDRED 231
LV DEE +Y F ED
Sbjct: 264 PALVEKDQDEEKKEVYPKFVFED 286
>ACC1_PETHY (Q08506) 1-aminocyclopropane-1-carboxylate oxidase 1 (EC
1.14.17.4) (ACC oxidase 1) (Ethylene-forming enzyme)
(EFE)
Length = 319
Score = 58.2 bits (139), Expect = 2e-08
Identities = 46/141 (32%), Positives = 66/141 (46%), Gaps = 8/141 (5%)
Query: 95 YASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQS-QVDGLEVLTKDGRKWIS 153
Y S+ + K Y D GL HTD + +LFQ +V GL++L KDG+ WI
Sbjct: 148 YGSKGPNFGTKVSNYPPCPKPDLIKGLRAHTDAGGIILLFQDDKVSGLQLL-KDGQ-WID 205
Query: 154 YKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPE 213
P S +V + D +NG+ S HRV+ + AR S A F+ P +I
Sbjct: 206 VPPMRHS--IVVNLGDQLEVITNGKYKSVMHRVIAQKDGARMSLASFYNPGSDAVIYPAP 263
Query: 214 ELVD---EEHPLLYKPFDRED 231
LV+ EE+ +Y F +D
Sbjct: 264 ALVEKEAEENKQVYPKFVFDD 284
>ACC1_ARATH (Q06588) 1-aminocyclopropane-1-carboxylate oxidase (EC
1.14.17.4) (ACC oxidase) (Ethylene-forming enzyme) (EFE)
Length = 323
Score = 58.2 bits (139), Expect = 2e-08
Identities = 43/127 (33%), Positives = 61/127 (47%), Gaps = 6/127 (4%)
Query: 95 YASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQ-SQVDGLEVLTKDGRKWIS 153
Y S+ T+ K Y D GL HTD + +LFQ +V GL++L KDG +W+
Sbjct: 148 YGSKRPTFGTKVSNYPPCPNPDLVKGLRAHTDAGGIILLFQDDKVSGLQLL-KDG-EWVD 205
Query: 154 YKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVM-MTGNEARYSAALFFVPKGGCIIKAP 212
P S +V + D +NG+ S HRV+ T E R S A F+ P +I
Sbjct: 206 VPP--VKHSIVVNLGDQLEVITNGKYKSVEHRVLSQTDGEGRMSIASFYNPGSDSVIFPA 263
Query: 213 EELVDEE 219
EL+ +E
Sbjct: 264 PELIGKE 270
>ACCO_MUSAC (Q9FR99) 1-aminocyclopropane-1-carboxylate oxidase (EC
1.14.17.4) (ACC oxidase) (Ethylene-forming enzyme) (EFE)
Length = 306
Score = 57.4 bits (137), Expect = 3e-08
Identities = 46/132 (34%), Positives = 61/132 (45%), Gaps = 7/132 (5%)
Query: 99 PITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQS-QVDGLEVLTKDGRKWISYKPS 157
P + K Y D GL HTD + +LFQ QV GL++L KDGR WI +P
Sbjct: 152 PPFFGTKVSHYPPCPRLDLVKGLRAHTDAGGVILLFQDDQVGGLQML-KDGR-WIDVQPL 209
Query: 158 SESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVD 217
+++ ++ D SNGR S +HRV+ T + R S A F+ P I
Sbjct: 210 ADA--IVINTGDQIEVLSNGRYKSAWHRVLATSHGNRRSIASFYNPSLKATIAPAAGAAT 267
Query: 218 EE--HPLLYKPF 227
EE P LY F
Sbjct: 268 EEAAPPALYPKF 279
>FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1)
Length = 336
Score = 57.0 bits (136), Expect = 4e-08
Identities = 39/130 (30%), Positives = 60/130 (46%), Gaps = 4/130 (3%)
Query: 102 YFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQ 161
Y +K Y D +G+ HTD + +T+L ++V GL+V D Y PS+
Sbjct: 199 YMMKINYYPPCPRPDLALGVPAHTDLSGITLLVPNEVPGLQVFKDDHWFDAEYIPSA--- 255
Query: 162 SFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHP 221
IV + D SNGR + HR + + R S +F P I+ EL +++P
Sbjct: 256 -VIVHIGDQILRLSNGRYKNVLHRTTVDKEKTRMSWPVFLEPPREKIVGPLPELTGDDNP 314
Query: 222 LLYKPFDRED 231
+KPF +D
Sbjct: 315 PKFKPFAFKD 324
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,470,005
Number of Sequences: 164201
Number of extensions: 1226853
Number of successful extensions: 2707
Number of sequences better than 10.0: 76
Number of HSP's better than 10.0 without gapping: 62
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 2589
Number of HSP's gapped (non-prelim): 79
length of query: 231
length of database: 59,974,054
effective HSP length: 107
effective length of query: 124
effective length of database: 42,404,547
effective search space: 5258163828
effective search space used: 5258163828
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Lotus: description of TM0226.11


