

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.8
(423 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
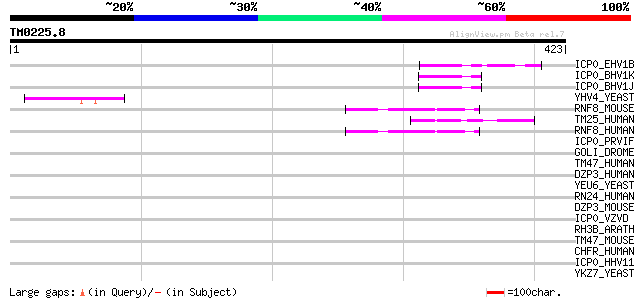
Score E
Sequences producing significant alignments: (bits) Value
ICP0_EHV1B (P28990) Trans-acting transcriptional protein ICP0 54 6e-07
ICP0_BHV1K (P29836) Trans-acting transcriptional protein ICP0 (P... 51 5e-06
ICP0_BHV1J (P29128) Trans-acting transcriptional protein ICP0 (P... 51 5e-06
YHV4_YEAST (P38850) Hypothetical 123.0 kDa protein in SPO16-REC1... 45 4e-04
RNF8_MOUSE (Q8VC56) Ubiquitin ligase protein RNF8 (EC 6.3.2.-) (... 45 4e-04
TM25_HUMAN (Q14258) Tripartite motif-containing protein 25 (Zinc... 44 6e-04
RNF8_HUMAN (O76064) Ubiquitin ligase protein RNF8 (EC 6.3.2.-) (... 44 6e-04
ICP0_PRVIF (P29129) Trans-acting transcriptional protein ICP0 (E... 44 0.001
GOLI_DROME (Q06003) Goliath protein (Protein g1) 44 0.001
TM47_HUMAN (Q96LD4) Tripartite motif-containing protein 47 (RING... 43 0.001
DZP3_HUMAN (Q86Y13) Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) ... 43 0.001
YEU6_YEAST (P40072) Hypothetical 30.8 kDa protein in SPR6-RPL23B... 43 0.002
RN24_HUMAN (Q9Y225) RING finger protein 24 43 0.002
DZP3_MOUSE (Q7TPV2) Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) ... 43 0.002
ICP0_VZVD (P09309) Trans-acting transcriptional protein ICP0 42 0.002
RH3B_ARATH (Q9ZT49) RING-H2 zinc finger protein RHA3b 42 0.003
TM47_MOUSE (Q8C0E3) Tripartite motif-containing protein 47 42 0.004
CHFR_HUMAN (Q96EP1) Ubiquitin ligase protein CHFR (EC 6.3.2.-) (... 42 0.004
ICP0_HHV11 (P08393) Trans-acting transcriptional protein ICP0 (I... 41 0.005
YKZ7_YEAST (P36113) Hypothetical 63.6 kDa protein in YPT52-GCN3 ... 41 0.007
>ICP0_EHV1B (P28990) Trans-acting transcriptional protein ICP0
Length = 532
Score = 54.3 bits (129), Expect = 6e-07
Identities = 33/93 (35%), Positives = 39/93 (41%), Gaps = 16/93 (17%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHAAT 372
C IC ED S+ LPC H FC CI +W Q TCPLCK + V H
Sbjct: 8 CPICLEDPSNYSMALPCLHAFCYVCITRWIRQN------PTCPLCKVP---VESVVHTIE 58
Query: 373 ADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
+D E V D D E ++SFE
Sbjct: 59 SDSEFKETKVSVD-------FDYDSEEDEDSFE 84
>ICP0_BHV1K (P29836) Trans-acting transcriptional protein ICP0 (P135
protein) (IER 2.9/ER2.6)
Length = 676
Score = 51.2 bits (121), Expect = 5e-06
Identities = 22/48 (45%), Positives = 26/48 (53%), Gaps = 6/48 (12%)
Query: 312 SCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKA 359
SC IC + + LPC H FCL CI++W R TCPLCKA
Sbjct: 12 SCCICLDAITGAARALPCLHAFCLACIRRWLEGR------PTCPLCKA 53
>ICP0_BHV1J (P29128) Trans-acting transcriptional protein ICP0 (P135
protein) (IER 2.9/ER2.6)
Length = 676
Score = 51.2 bits (121), Expect = 5e-06
Identities = 22/48 (45%), Positives = 26/48 (53%), Gaps = 6/48 (12%)
Query: 312 SCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKA 359
SC IC + + LPC H FCL CI++W R TCPLCKA
Sbjct: 12 SCCICLDAITGAARALPCLHAFCLACIRRWLEGR------PTCPLCKA 53
>YHV4_YEAST (P38850) Hypothetical 123.0 kDa protein in SPO16-REC104
intergenic region
Length = 1070
Score = 45.1 bits (105), Expect = 4e-04
Identities = 27/86 (31%), Positives = 43/86 (49%), Gaps = 10/86 (11%)
Query: 12 KIVATVTGYHGLERFNLIKLINYAGGSYIGSMSKSITHLKFE---GKKYDIARRLS---- 64
++ T Y G +RF + +L+ GG +++ THL + GKK+ +A++ S
Sbjct: 379 ELTVAYTNYFGSQRFYIQRLVEILGGLSTPELTRKNTHLITKSTIGKKFKVAKKWSLDPQ 438
Query: 65 --IPVVNHRWIEDCIREGKRL-PIDS 87
I V NH W+E C +L P DS
Sbjct: 439 NAIIVTNHMWLEQCYMNNSKLNPKDS 464
>RNF8_MOUSE (Q8VC56) Ubiquitin ligase protein RNF8 (EC 6.3.2.-)
(RING finger protein 8) (AIP37) (ActA-interacting
protein 37) (LaXp180)
Length = 488
Score = 45.1 bits (105), Expect = 4e-04
Identities = 28/102 (27%), Positives = 45/102 (43%), Gaps = 14/102 (13%)
Query: 257 MNGSSDGIEQITDSNQIEDPQTVEQTSIDLCSSDGKFIDGDQVDNVAGSPISNEWSCVIC 316
+N S E+I + E QT E+ D ++V + + NE C+IC
Sbjct: 357 LNCSKKDFEKIIQAKNKELEQTKEE-------KDKVQAQKEEVLSHMNDLLENELQCIIC 409
Query: 317 FEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
E F + L C H+FC CI +W +++ CP+C+
Sbjct: 410 SEYFIEAV-TLNCAHSFCSFCINEWMKRKVE------CPICR 444
>TM25_HUMAN (Q14258) Tripartite motif-containing protein 25 (Zinc
finger protein 147) (Estrogen responsive finger protein)
(Efp)
Length = 630
Score = 44.3 bits (103), Expect = 6e-04
Identities = 29/96 (30%), Positives = 43/96 (44%), Gaps = 16/96 (16%)
Query: 306 PISNEWSCVICFEDFSSTLGVLPCEHTFCLTCI-QKWAHQRISMGKISTCPLCKASFAWY 364
P++ E SC IC E F + PC H FC +C+ + WA Q G CP C+A +
Sbjct: 6 PLAEELSCSICLEPFKEPV-TTPCGHNFCGSCLNETWAVQ----GSPYLCPQCRAVY--- 57
Query: 365 KIVEHAATADKELNSQTVPCDNLASVTFISMDQELP 400
A +L+ TV C+ + + +E P
Sbjct: 58 -------QARPQLHKNTVLCNVVEQFLQADLAREPP 86
>RNF8_HUMAN (O76064) Ubiquitin ligase protein RNF8 (EC 6.3.2.-)
(RING finger protein 8)
Length = 485
Score = 44.3 bits (103), Expect = 6e-04
Identities = 28/102 (27%), Positives = 44/102 (42%), Gaps = 14/102 (13%)
Query: 257 MNGSSDGIEQITDSNQIEDPQTVEQTSIDLCSSDGKFIDGDQVDNVAGSPISNEWSCVIC 316
+N S E I + E QT E+ + ++V + + NE C+IC
Sbjct: 354 LNRSKKDFEAIIQAKNKELEQTKEE-------KEKMQAQKEEVLSHMNDVLENELQCIIC 406
Query: 317 FEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
E F + L C H+FC CI +W ++I CP+C+
Sbjct: 407 SEYFIEAV-TLNCAHSFCSYCINEWMKRKIE------CPICR 441
>ICP0_PRVIF (P29129) Trans-acting transcriptional protein ICP0
(Early protein 0) (EP0)
Length = 410
Score = 43.5 bits (101), Expect = 0.001
Identities = 19/47 (40%), Positives = 24/47 (50%), Gaps = 6/47 (12%)
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKA 359
C IC + ++ LPC H FCL CIQ+W + CPLC A
Sbjct: 46 CPICLDVAATEAQTLPCMHKFCLDCIQRWTLTS------TACPLCNA 86
>GOLI_DROME (Q06003) Goliath protein (Protein g1)
Length = 286
Score = 43.5 bits (101), Expect = 0.001
Identities = 29/95 (30%), Positives = 40/95 (41%), Gaps = 28/95 (29%)
Query: 277 QTVEQTSIDLCS-----------SDGKFIDGDQVDNVAGSPISNEWSCVICFEDF--SST 323
Q +Q S +LCS GKF D +D+ C IC E + + T
Sbjct: 90 QAKDQQSRNLCSVTKKAIMKIPTKTGKFSDEKDLDSDC---------CAICIEAYKPTDT 140
Query: 324 LGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
+ +LPC+H F CI W + TCP+CK
Sbjct: 141 IRILPCKHEFHKNCIDPWLIEH------RTCPMCK 169
>TM47_HUMAN (Q96LD4) Tripartite motif-containing protein 47 (RING
finger protein 100) (Gene overexpressed in astrocytoma
protein)
Length = 638
Score = 43.1 bits (100), Expect = 0.001
Identities = 23/59 (38%), Positives = 29/59 (48%), Gaps = 6/59 (10%)
Query: 308 SNEWSCVICFEDFSSTLGVLPCEHTFCLTCI-QKWAHQRIS----MGKISTCPLCKASF 361
S +SC IC E + LPC H FCL C+ W H+ S G + CPLC+ F
Sbjct: 4 SGPFSCPICLEPLREPV-TLPCGHNFCLACLGALWPHRGASGAGGPGGAARCPLCQEPF 61
>DZP3_HUMAN (Q86Y13) Ubiquitin ligase protein DZIP3 (EC 6.3.2.-)
(DAZ-interacting protein 3) (RNA-binding ubiquitin ligase
of 138 kDa) (hRUL138)
Length = 1208
Score = 43.1 bits (100), Expect = 0.001
Identities = 22/50 (44%), Positives = 26/50 (52%), Gaps = 7/50 (14%)
Query: 310 EWSCVICFEDFS-STLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
E CVIC E+ S L VLPC H F CI+ W Q+ TCP C+
Sbjct: 1145 EEPCVICHENLSPENLSVLPCAHKFHAQCIRPWLMQQ------GTCPTCR 1188
>YEU6_YEAST (P40072) Hypothetical 30.8 kDa protein in SPR6-RPL23B
intergenic region
Length = 274
Score = 42.7 bits (99), Expect = 0.002
Identities = 36/153 (23%), Positives = 64/153 (41%), Gaps = 10/153 (6%)
Query: 216 DRDRLHTEAAATSSLSGGVDVNDLQENSSGPDARLCNQNRAMNGSSDGIEQITDSNQIED 275
DR+ +HTE SS + N+++E + + + D T
Sbjct: 100 DRETMHTEEPEASSGNNITLTNNVEELHTMDVLSQTANTPSASPMLDAAPPTTKPGTNSK 159
Query: 276 PQTVEQTS--IDLCSSDGKFI---DGDQVDNVAGSP----ISNEWSCVICFEDFSSTLGV 326
QTV+ T+ IDL + + + + D D + +P + ++ C ICFE + L
Sbjct: 160 EQTVDLTADAIDLDAEEQQVLQISDDDFQEETKEAPKEYGAAKDYRCPICFEPPETALMT 219
Query: 327 LPCEHTFCLTCIQKWAHQRISMGKISTCPLCKA 359
L C H FC C+ + + + + C LC++
Sbjct: 220 L-CGHVFCCPCLFQMVNSSRTCRQFGHCALCRS 251
>RN24_HUMAN (Q9Y225) RING finger protein 24
Length = 148
Score = 42.7 bits (99), Expect = 0.002
Identities = 19/47 (40%), Positives = 23/47 (48%), Gaps = 8/47 (17%)
Query: 313 CVICFEDFS--STLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLC 357
C +C EDF LG+ PC+H F C+ KW R CPLC
Sbjct: 78 CAVCLEDFKPRDELGICPCKHAFHRKCLIKWLEVR------KVCPLC 118
>DZP3_MOUSE (Q7TPV2) Ubiquitin ligase protein DZIP3 (EC 6.3.2.-)
(DAZ-interacting protein 3 homolog)
Length = 1204
Score = 42.7 bits (99), Expect = 0.002
Identities = 22/50 (44%), Positives = 26/50 (52%), Gaps = 7/50 (14%)
Query: 310 EWSCVICFEDFS-STLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
E CVIC E+ S L VLPC H F CI+ W Q+ TCP C+
Sbjct: 1141 EEPCVICHENLSPENLSVLPCAHKFHSQCIRPWLMQQ------GTCPTCR 1184
>ICP0_VZVD (P09309) Trans-acting transcriptional protein ICP0
Length = 467
Score = 42.4 bits (98), Expect = 0.002
Identities = 21/78 (26%), Positives = 36/78 (45%), Gaps = 8/78 (10%)
Query: 303 AGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK--AS 360
+G+ +++ +C IC S +PC H FC CI+ W + CPLC+
Sbjct: 9 SGTSDASDNTCTICMSTVSDLGKTMPCLHDFCFVCIRAWTSTSVQ------CPLCRCPVQ 62
Query: 361 FAWYKIVEHAATADKELN 378
+KIV + + E++
Sbjct: 63 SILHKIVSDTSYKEYEVH 80
>RH3B_ARATH (Q9ZT49) RING-H2 zinc finger protein RHA3b
Length = 200
Score = 42.0 bits (97), Expect = 0.003
Identities = 32/100 (32%), Positives = 43/100 (43%), Gaps = 26/100 (26%)
Query: 287 CSSDGKFIDGDQVDNVAGSPISNEWSCVICFEDFSS--TLGVLP-CEHTFCLTCIQKWAH 343
CSS G DGD + C IC +FS + +LP C H F + CI KW
Sbjct: 101 CSSVG---DGD-----------SSTECAICITEFSEGEEIRILPLCSHAFHVACIDKWLT 146
Query: 344 QRISMGKISTCPLCKASFAWYK---IVEHAATADKELNSQ 380
R S+CP C+ K HA+TA+ ++ Q
Sbjct: 147 SR------SSCPSCRRILVPVKCDRCGHHASTAETQVKDQ 180
>TM47_MOUSE (Q8C0E3) Tripartite motif-containing protein 47
Length = 641
Score = 41.6 bits (96), Expect = 0.004
Identities = 22/59 (37%), Positives = 28/59 (47%), Gaps = 6/59 (10%)
Query: 308 SNEWSCVICFEDFSSTLGVLPCEHTFCLTCI-QKWAHQRI----SMGKISTCPLCKASF 361
S +SC IC E + LPC H FCL C+ W H+ G + CPLC+ F
Sbjct: 4 SGPFSCPICLEPLREPV-TLPCGHNFCLACLGALWPHRSAGGTGGSGGPARCPLCQEPF 61
>CHFR_HUMAN (Q96EP1) Ubiquitin ligase protein CHFR (EC 6.3.2.-)
(Checkpoint with forkhead and RING finger domains
protein)
Length = 664
Score = 41.6 bits (96), Expect = 0.004
Identities = 21/64 (32%), Positives = 29/64 (44%), Gaps = 8/64 (12%)
Query: 297 DQVDNVAGSPISNE--WSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTC 354
+ V AG P E +C+IC + + + PC HTFC C W M + S C
Sbjct: 286 EDVRAAAGKPDKMEETLTCIICQDLLHDCVSLQPCMHTFCAACYSGW------MERSSLC 339
Query: 355 PLCK 358
P C+
Sbjct: 340 PTCR 343
>ICP0_HHV11 (P08393) Trans-acting transcriptional protein ICP0
(Immediate-early protein IE110) (VMW110) (Alpha-0
protein)
Length = 775
Score = 41.2 bits (95), Expect = 0.005
Identities = 17/53 (32%), Positives = 26/53 (48%), Gaps = 8/53 (15%)
Query: 313 CVICFEDFSSTL--GVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAW 363
C +C ++ + L PC H FC+ C++ W R +TCPLC A +
Sbjct: 116 CAVCTDEIAPHLRCDTFPCMHRFCIPCMKTWMQLR------NTCPLCNAKLVY 162
>YKZ7_YEAST (P36113) Hypothetical 63.6 kDa protein in YPT52-GCN3
intergenic region
Length = 551
Score = 40.8 bits (94), Expect = 0.007
Identities = 16/63 (25%), Positives = 29/63 (45%)
Query: 295 DGDQVDNVAGSPISNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTC 354
D D + N+++C+IC + + L C H +C+ C + + ++ G I TC
Sbjct: 161 DNDYNSHFREVEFKNDFTCIICCDKKDTETFALECGHEYCINCYRHYIKDKLHEGNIITC 220
Query: 355 PLC 357
C
Sbjct: 221 MDC 223
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,882,215
Number of Sequences: 164201
Number of extensions: 2138428
Number of successful extensions: 5409
Number of sequences better than 10.0: 205
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 176
Number of HSP's that attempted gapping in prelim test: 5304
Number of HSP's gapped (non-prelim): 258
length of query: 423
length of database: 59,974,054
effective HSP length: 113
effective length of query: 310
effective length of database: 41,419,341
effective search space: 12839995710
effective search space used: 12839995710
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0225.8


