

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.5
(295 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
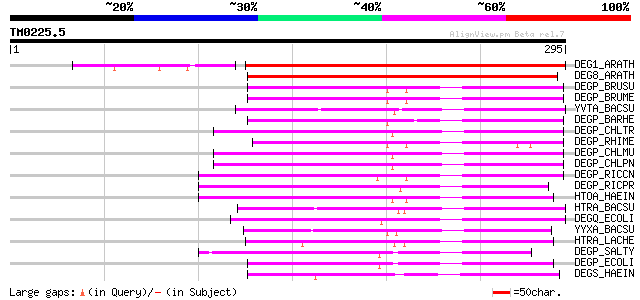
Score E
Sequences producing significant alignments: (bits) Value
DEG1_ARATH (O22609) Protease Do-like 1, chloroplast precursor (E... 307 2e-83
DEG8_ARATH (Q9LU10) Protease Do-like 8, chloroplast precursor (E... 162 7e-40
DEGP_BRUSU (Q44597) Probable serine protease do-like precursor (... 119 1e-26
DEGP_BRUME (Q8YG32) Probable serine protease do-like precursor (... 119 1e-26
YVTA_BACSU (Q9R9I1) Probable serine protease yvtA (EC 3.4.21.-) 117 3e-26
DEGP_BARHE (P54925) Probable periplasmic serine protease DO-like... 115 1e-25
DEGP_CHLTR (P18584) Probable serine protease do-like precursor (... 109 7e-24
DEGP_RHIME (Q52894) Probable serine protease do-like precursor (... 109 9e-24
DEGP_CHLMU (Q9PL97) Probable serine protease do-like precursor (... 108 2e-23
DEGP_CHLPN (Q9Z6T0) Probable serine protease do-like precursor (... 107 3e-23
DEGP_RICCN (Q92JA1) Probable serine protease do-like precursor (... 102 1e-21
DEGP_RICPR (O05942) Probable serine protease do-like precursor (... 101 2e-21
HTOA_HAEIN (P45129) Probable periplasmic serine protease do/hhoA... 100 3e-21
HTRA_BACSU (O34358) Probable serine protease do-like htrA (EC 3.... 100 4e-21
DEGQ_ECOLI (P39099) Protease degQ precursor (EC 3.4.21.-) 100 4e-21
YYXA_BACSU (P39668) Hypothetical serine protease yyxA (EC 3.4.21.-) 100 6e-21
HTRA_LACHE (Q9Z4H7) Serine protease do-like htrA (EC 3.4.21.-) 100 6e-21
DEGP_SALTY (P26982) Protease do precursor (EC 3.4.21.-) 98 3e-20
DEGP_ECOLI (P09376) Protease do precursor (EC 3.4.21.-) 98 3e-20
DEGS_HAEIN (P44947) Protease degS precursor (EC 3.4.21.-) 94 5e-19
>DEG1_ARATH (O22609) Protease Do-like 1, chloroplast precursor (EC
3.4.21.-)
Length = 439
Score = 307 bits (787), Expect = 2e-83
Identities = 153/170 (90%), Positives = 163/170 (95%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSG LIGINTAIYSPSGASSGVGFSIP+DTV GIVDQLV+F
Sbjct: 270 VIQTDAAINPGNSGGPLLDSSGTLIGINTAIYSPSGASSGVGFSIPVDTVGGIVDQLVRF 329
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAP +GPAGKAGLQSTKRD YGRL+LGDIIT
Sbjct: 330 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPPSGPAGKAGLQSTKRDGYGRLVLGDIIT 389
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
SVNGTKV++GSDLYRILDQCKVGD+VTVEVLRGDHKEKI V LEPKPDES
Sbjct: 390 SVNGTKVSNGSDLYRILDQCKVGDEVTVEVLRGDHKEKISVTLEPKPDES 439
Score = 73.2 bits (178), Expect = 7e-13
Identities = 51/116 (43%), Positives = 64/116 (54%), Gaps = 31/116 (26%)
Query: 34 TPKTCFNSILILCTSIALSFT------NADSAYAFVVTPPRKLQSDELATV--------- 78
TP + +LCTS+ALSF+ +SA AFVV+ P+KLQ+DELATV
Sbjct: 72 TPFSAVKPFFLLCTSVALSFSLFAASPAVESASAFVVSTPKKLQTDELATVRLFQENTPS 131
Query: 79 -VYITNLAVKMRLRWM-------------CWRFLKEGNIVTNYHVIPGASDLSLDL 120
VYITNLAV+ + W K+G+IVTNYHVI GASDL + L
Sbjct: 132 VVYITNLAVRQDAFTLDVLEVPQGSGSGFVWD--KQGHIVTNYHVIRGASDLRVTL 185
>DEG8_ARATH (Q9LU10) Protease Do-like 8, chloroplast precursor (EC
3.4.21.-)
Length = 448
Score = 162 bits (411), Expect = 7e-40
Identities = 86/166 (51%), Positives = 114/166 (67%), Gaps = 1/166 (0%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ +DAAINPGNSGGPLLDS GNLIGINTAI++ +G S+GVGF+IP TV IV QL++F
Sbjct: 281 IQTDAAINPGNSGGPLLDSKGNLIGINTAIFTQTGTSAGVGFAIPSSTVLKIVPQLIQFS 340
Query: 187 KVTRPILGIKFAPDQSVEQLGV-SGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
KV R + I+ APD QL V +G LVL P A KAGL T R G ++LGDII
Sbjct: 341 KVLRAGINIELAPDPVANQLNVRNGALVLQVPGKSLAEKAGLHPTSRGFAGNIVLGDIIV 400
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPK 291
+V+ V + ++L +ILD+ VGDKVT+++ RG+ ++ + LE K
Sbjct: 401 AVDDKPVKNKAELMKILDEYSVGDKVTLKIKRGNEDLELKISLEEK 446
>DEGP_BRUSU (Q44597) Probable serine protease do-like precursor (EC
3.4.21.-)
Length = 513
Score = 119 bits (297), Expect = 1e-26
Identities = 69/173 (39%), Positives = 97/173 (55%), Gaps = 16/173 (9%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ DAA+N GNSGGP D SG +IGINTAI+SPSG S G+ F+IP T +VDQL+K G
Sbjct: 246 IQIDAAVNKGNSGGPAFDLSGEVIGINTAIFSPSGGSVGIAFAIPSSTAKQVVDQLIKKG 305
Query: 187 KVTRPILGIKFAP--DQSVEQLGVS---GVLVLDAPANGPAGKAGLQSTKRDSYGRLILG 241
V R +G++ P LG++ G +V +GPA KAG+++ G
Sbjct: 306 SVERGWIGVQIQPVTKDIAASLGLAEEKGAIVASPQDDGPAAKAGIKA-----------G 354
Query: 242 DIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDE 294
D+IT+VNG V DL R + G+K + V R + E+I V + P++
Sbjct: 355 DVITAVNGETVQDPRDLARKVANIAPGEKAALTVWRKNKAEEINVTIAAMPND 407
>DEGP_BRUME (Q8YG32) Probable serine protease do-like precursor (EC
3.4.21.-)
Length = 513
Score = 119 bits (297), Expect = 1e-26
Identities = 69/173 (39%), Positives = 97/173 (55%), Gaps = 16/173 (9%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ DAA+N GNSGGP D SG +IGINTAI+SPSG S G+ F+IP T +VDQL+K G
Sbjct: 246 IQIDAAVNKGNSGGPAFDLSGEVIGINTAIFSPSGGSVGIAFAIPSSTAKQVVDQLIKKG 305
Query: 187 KVTRPILGIKFAP--DQSVEQLGVS---GVLVLDAPANGPAGKAGLQSTKRDSYGRLILG 241
V R +G++ P LG++ G +V +GPA KAG+++ G
Sbjct: 306 SVERGWIGVQIQPVTKDIAASLGLAEEKGAIVASPQDDGPAAKAGIKA-----------G 354
Query: 242 DIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDE 294
D+IT+VNG V DL R + G+K + V R + E+I V + P++
Sbjct: 355 DVITAVNGETVQDPRDLARKVANIAPGEKAALTVWRKNKAEEINVTIAAMPND 407
>YVTA_BACSU (Q9R9I1) Probable serine protease yvtA (EC 3.4.21.-)
Length = 458
Score = 117 bits (294), Expect = 3e-26
Identities = 76/189 (40%), Positives = 107/189 (56%), Gaps = 27/189 (14%)
Query: 121 TTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVD 180
T + ++ +DAAINPGNSGGPL+++SG +IGIN+ S SG S +GF+IP + V IVD
Sbjct: 281 TVEMNVLQTDAAINPGNSGGPLINASGQVIGINSLKVSESGVES-LGFAIPSNDVEPIVD 339
Query: 181 QLVKFGKVTRPILGIKFAPDQSV-------------EQLGVSGVLVLDAPANGPAGKAGL 227
QL++ GKV RP LG++ V +QLG GV V + AN PA KAG+
Sbjct: 340 QLLQNGKVDRPFLGVQMIDMSQVPETYQENTLGLFGDQLG-KGVYVKEVQANSPAEKAGI 398
Query: 228 QSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRIL-DQCKVGDKVTVEVLRGDHKEKIPV 286
+S D+I +NG V S +D+ +IL KVGDK T++VLR + +
Sbjct: 399 KSE-----------DVIVKLNGKDVESSADIRQILYKDLKVGDKTTIQVLRKGKTKTLNA 447
Query: 287 ILEPKPDES 295
L + + S
Sbjct: 448 TLTKQTESS 456
>DEGP_BARHE (P54925) Probable periplasmic serine protease DO-like
precursor (EC 3.4.21.-) (Antigen htrA)
Length = 503
Score = 115 bits (289), Expect = 1e-25
Identities = 65/173 (37%), Positives = 96/173 (54%), Gaps = 17/173 (9%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ DAA+N GNSGGP D +G ++G+NTAI+SPSG + G+ F+IP T +V QL++ G
Sbjct: 236 IQIDAAVNRGNSGGPTFDLNGKVVGVNTAIFSPSGGNVGIAFAIPAATAKQVVQQLIEKG 295
Query: 187 KVTRPILGIKFAP-----DQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILG 241
V R LG++ P S+ G L+ D P GPA KAG+++ G
Sbjct: 296 LVQRGWLGVQIQPVTKEISDSIGLKEAKGALITD-PLKGPAAKAGIKA-----------G 343
Query: 242 DIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDE 294
D+I SVNG K+ DL + + G+ VT+ V + +E I V L+ P++
Sbjct: 344 DVIISVNGEKINDVRDLAKRIANMSPGETVTLGVWKSGKEENIKVKLDSMPED 396
>DEGP_CHLTR (P18584) Probable serine protease do-like precursor (EC
3.4.21.-) (59 kDa immunogenic protein) (SK59)
Length = 497
Score = 109 bits (273), Expect = 7e-24
Identities = 70/191 (36%), Positives = 97/191 (50%), Gaps = 16/191 (8%)
Query: 109 VIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGF 168
VI L + + +DAAINPGNSGGPLL+ +G +IG+NTAI S SG G+GF
Sbjct: 218 VISAKGRNQLHIVDFEDFIQTDAAINPGNSGGPLLNINGQVIGVNTAIVSGSGGYIGIGF 277
Query: 169 SIPLDTVNGIVDQLVKFGKVTRPILGIKFAPDQS-----VEQLGVSGVLVLDAPANGPAG 223
+IP ++DQL+ G+VTR LG+ P S + V G LV D PA
Sbjct: 278 AIPSLMAKRVIDQLISDGQVTRGFLGVTLQPIDSELATCYKLEKVYGALVTDVVKGSPAE 337
Query: 224 KAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEK 283
KAGL+ D+I + NG +V S S L + G +V ++++R +
Sbjct: 338 KAGLRQE-----------DVIVAYNGKEVESLSALRNAISLMMPGTRVVLKIVREGKTIE 386
Query: 284 IPVILEPKPDE 294
IPV + P E
Sbjct: 387 IPVTVTQIPTE 397
Score = 32.0 bits (71), Expect = 1.9
Identities = 22/70 (31%), Positives = 34/70 (48%), Gaps = 12/70 (17%)
Query: 210 GVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGD 269
G+LV+ A PA AG+ G +I +VN +V S +L ++L K G+
Sbjct: 429 GILVVAVEAGSPAASAGVAP-----------GQLILAVNRQRVASVEELNQVLKNSK-GE 476
Query: 270 KVTVEVLRGD 279
V + V +GD
Sbjct: 477 NVLLMVSQGD 486
>DEGP_RHIME (Q52894) Probable serine protease do-like precursor (EC
3.4.21.-)
Length = 504
Score = 109 bits (272), Expect = 9e-24
Identities = 69/185 (37%), Positives = 98/185 (52%), Gaps = 31/185 (16%)
Query: 130 DAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFGKVT 189
DAA+N GNSGGP + SG ++GINTAI+SPSG + G+ F+IP +VD L+K G V+
Sbjct: 236 DAAVNRGNSGGPTFNLSGEVVGINTAIFSPSGGNVGIAFAIPASVAKDVVDSLIKDGTVS 295
Query: 190 RPILGIKFAP--DQSVEQLGVS---GVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDII 244
R LG++ P E LG+S G LV++ A P KAG+++ GD++
Sbjct: 296 RGWLGVQIQPVTKDIAESLGLSEANGALVVEPQAGSPGEKAGIKN-----------GDVV 344
Query: 245 TSVNGTKVTSGSDLYRILDQCKVG-------------DKVTVEV--LRGDHKEKIPVILE 289
T++NG V DL R + + G + V +E+ L D KE P E
Sbjct: 345 TALNGEPVKDPRDLARRVAALRPGSTAEVTLWRSGKSETVNLEIGTLPSDAKEPAPATGE 404
Query: 290 PKPDE 294
+PDE
Sbjct: 405 AQPDE 409
>DEGP_CHLMU (Q9PL97) Probable serine protease do-like precursor (EC
3.4.21.-)
Length = 497
Score = 108 bits (269), Expect = 2e-23
Identities = 70/191 (36%), Positives = 96/191 (49%), Gaps = 16/191 (8%)
Query: 109 VIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGF 168
VI L + + +DAAINPGNSGGPLL+ G +IG+NTAI S SG G+GF
Sbjct: 218 VISAKGRNQLHIVDFEDFIQTDAAINPGNSGGPLLNIDGQVIGVNTAIVSGSGGYIGIGF 277
Query: 169 SIPLDTVNGIVDQLVKFGKVTRPILGIKFAPDQS-----VEQLGVSGVLVLDAPANGPAG 223
+IP ++DQL+ G+VTR LG+ P S + V G L+ D PA
Sbjct: 278 AIPSLMAKRVIDQLISDGQVTRGFLGVTLQPIDSELAACYKLEKVYGALITDVVKGSPAE 337
Query: 224 KAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEK 283
KAGL+ D+I + NG +V S S L + G +V ++V+R +
Sbjct: 338 KAGLRQE-----------DVIVAYNGKEVESLSALRNAISLMMPGTRVVLKVVREGKFIE 386
Query: 284 IPVILEPKPDE 294
IPV + P E
Sbjct: 387 IPVTVTQIPAE 397
>DEGP_CHLPN (Q9Z6T0) Probable serine protease do-like precursor (EC
3.4.21.-)
Length = 488
Score = 107 bits (268), Expect = 3e-23
Identities = 69/191 (36%), Positives = 96/191 (50%), Gaps = 16/191 (8%)
Query: 109 VIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGF 168
VI L + + +DAAINPGNSGGPLL+ G +IG+NTAI S SG G+GF
Sbjct: 209 VISAKGRNQLHIADFEDFIQTDAAINPGNSGGPLLNIDGQVIGVNTAIVSGSGGYIGIGF 268
Query: 169 SIPLDTVNGIVDQLVKFGKVTRPILGIKFAPDQS-----VEQLGVSGVLVLDAPANGPAG 223
+IP N I+DQL++ G+VTR LG+ P + + V G LV D PA
Sbjct: 269 AIPSLMANRIIDQLIRDGQVTRGFLGVTLQPIDAELAACYKLEKVYGALVTDVVKGSPAD 328
Query: 224 KAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEK 283
KAGL+ D+I + NG +V S S + ++ ++V+R +
Sbjct: 329 KAGLKQE-----------DVIIAYNGKEVDSLSMFRNAVSLMNPDTRIVLKVVREGKVIE 377
Query: 284 IPVILEPKPDE 294
IPV + P E
Sbjct: 378 IPVTVSQAPKE 388
>DEGP_RICCN (Q92JA1) Probable serine protease do-like precursor (EC
3.4.21.-)
Length = 508
Score = 102 bits (253), Expect = 1e-21
Identities = 66/200 (33%), Positives = 106/200 (53%), Gaps = 17/200 (8%)
Query: 101 GNIVTNYHVIPGASDLSLDLTTLLQ-LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSP 159
G VT+ + D+ +D ++ + +DAAIN GNSGGP+ + +IG+NTAI+SP
Sbjct: 204 GGTVTSGIISSKGRDIDIDTDNIVDNFIQTDAAINNGNSGGPMFNLDQKVIGVNTAIFSP 263
Query: 160 SGASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILG--IKFAPDQSVEQLGVS---GVLVL 214
G + G+GF+IP +T I+++L K GKV+R LG I+ + E LG+ GVLV
Sbjct: 264 LGTNIGIGFAIPSNTAKPIIERLKKDGKVSRGRLGVTIQDLTEDISEGLGLKNTRGVLVA 323
Query: 215 DAPANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVE 274
+GP KAG+++ GDII V + L I+ + +V V+
Sbjct: 324 KVQEDGPGDKAGIKT-----------GDIIIEFADIPVKNTKKLRVIIADAPIDQEVKVK 372
Query: 275 VLRGDHKEKIPVILEPKPDE 294
+LR + ++P+ + +E
Sbjct: 373 ILRDKKELELPIKITSDNEE 392
>DEGP_RICPR (O05942) Probable serine protease do-like precursor (EC
3.4.21.-)
Length = 513
Score = 101 bits (252), Expect = 2e-21
Identities = 62/192 (32%), Positives = 102/192 (52%), Gaps = 17/192 (8%)
Query: 101 GNIVTNYHVIPGASDLSLDLTTLLQ-LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSP 159
G VT+ + D+ +D ++ + +DAAIN GNSGGP+ + +IG+NTAI+SP
Sbjct: 209 GGTVTSGIISSKGRDIDVDTDNIVDNFIQTDAAINNGNSGGPMFNLDQKVIGVNTAIFSP 268
Query: 160 SGASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILGIKFAP-DQSVEQL----GVSGVLVL 214
G + G+GF+IP +T I+++L K GKV+R LG+ + + ++ G +GVLV
Sbjct: 269 LGTNIGIGFAIPSNTAKPIIERLKKDGKVSRGRLGVTIQDLTEEISEVLGFKGTNGVLVS 328
Query: 215 DAPANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVE 274
NGP KAG++ GDII V + L I+ + +V ++
Sbjct: 329 KVQENGPGYKAGIKK-----------GDIIIKFGDRLVKNTKKLRVIIADTPINQEVKLK 377
Query: 275 VLRGDHKEKIPV 286
+LR + ++P+
Sbjct: 378 ILRDAQELELPI 389
>HTOA_HAEIN (P45129) Probable periplasmic serine protease
do/hhoA-like precursor (EC 3.4.21.-)
Length = 466
Score = 100 bits (250), Expect = 3e-21
Identities = 65/194 (33%), Positives = 104/194 (53%), Gaps = 16/194 (8%)
Query: 101 GNIVTNYHVIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPS 160
G VT+ V D T + +DAA+N GNSGG L++ +G LIGINTAI SPS
Sbjct: 189 GQTVTSGIVSALGRSTGSDSGTYENYIQTDAAVNRGNSGGALVNLNGELIGINTAIISPS 248
Query: 161 GASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILGIKFAPDQS--VEQLGVS---GVLVLD 215
G ++G+ F+IP + + +V Q+++FG+V R +LGIK + + VS G V +
Sbjct: 249 GGNAGIAFAIPSNQASNLVQQILEFGQVRRGLLGIKGGELNADLAKAFNVSAQQGAFVSE 308
Query: 216 APANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEV 275
A KAGL++ GDIIT++NG K++S +++ + G ++++
Sbjct: 309 VLPKSAAEKAGLKA-----------GDIITAMNGQKISSFAEIRAKIATTGAGKEISLTY 357
Query: 276 LRGDHKEKIPVILE 289
LR + + L+
Sbjct: 358 LRDGKSHDVKMKLQ 371
>HTRA_BACSU (O34358) Probable serine protease do-like htrA (EC
3.4.21.-)
Length = 449
Score = 100 bits (249), Expect = 4e-21
Identities = 62/187 (33%), Positives = 102/187 (54%), Gaps = 25/187 (13%)
Query: 122 TLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQ 181
T + ++ +DAAINPGNSGGPLL++ G ++GIN+ S G+GF+IP + V I ++
Sbjct: 274 TSINVIQTDAAINPGNSGGPLLNTDGKIVGINSMKISEDDV-EGIGFAIPSNDVKPIAEE 332
Query: 182 LVKFGKVTRPILGIKFAPDQSVEQ------LGV------SGVLVLDAPANGPAGKAGLQS 229
L+ G++ RP +G+ + V Q LG+ GV + + + PA KAGL++
Sbjct: 333 LLSKGQIERPYIGVSMLDLEQVPQNYQEGTLGLFGSQLNKGVYIREVASGSPAEKAGLKA 392
Query: 230 TKRDSYGRLILGDIITSVNGTKVTSGSDLYRIL-DQCKVGDKVTVEVLRGDHKEKIPVIL 288
DII + G ++ +GS+L IL K+GD V V++LR + + L
Sbjct: 393 E-----------DIIIGLKGKEIDTGSELRNILYKDAKIGDTVEVKILRNGKEMTKKIKL 441
Query: 289 EPKPDES 295
+ K +++
Sbjct: 442 DQKEEKT 448
>DEGQ_ECOLI (P39099) Protease degQ precursor (EC 3.4.21.-)
Length = 455
Score = 100 bits (249), Expect = 4e-21
Identities = 66/183 (36%), Positives = 97/183 (52%), Gaps = 16/183 (8%)
Query: 118 LDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNG 177
L+L L + +DA+IN GNSGG LL+ +G LIGINTAI +P G S G+GF+IP +
Sbjct: 194 LNLEGLENFIQTDASINRGNSGGALLNLNGELIGINTAILAPGGGSVGIGFAIPSNMART 253
Query: 178 IVDQLVKFGKVTRPILGIK----FAPDQSVEQLGVS-GVLVLDAPANGPAGKAGLQSTKR 232
+ QL+ FG++ R +LGIK A L V G V + + KAG+++
Sbjct: 254 LAQQLIDFGEIKRGLLGIKGTEMSADIAKAFNLDVQRGAFVSEVLPGSGSAKAGVKA--- 310
Query: 233 DSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKP 292
GDIITS+NG + S ++L + + G KV + +LR ++ V L+
Sbjct: 311 --------GDIITSLNGKPLNSFAELRSRIATTEPGTKVKLGLLRNGKPLEVEVTLDTST 362
Query: 293 DES 295
S
Sbjct: 363 SSS 365
>YYXA_BACSU (P39668) Hypothetical serine protease yyxA (EC 3.4.21.-)
Length = 400
Score = 100 bits (248), Expect = 6e-21
Identities = 65/175 (37%), Positives = 98/175 (55%), Gaps = 23/175 (13%)
Query: 125 QLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVK 184
+++ +DAAINPGNSGG LL+ G +IGIN+ + S A G+G SIP V +++ L +
Sbjct: 230 EVLQTDAAINPGNSGGALLNMDGKVIGINSMKIAES-AVEGIGLSIPSKLVIPVIEDLER 288
Query: 185 FGKVTRPILGIKFAP---------DQSVE--QLGVSGVLVLDAPANGPAGKAGLQSTKRD 233
+GKV RP LGI+ D++++ + +G +V+ A PAGKAGL+
Sbjct: 289 YGKVKRPFLGIEMKSLSDIASYHWDETLKLPKNVTNGAVVMGVDAFSPAGKAGLKEL--- 345
Query: 234 SYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVIL 288
D+IT +G KV DL + L Q KVGD+V V+ RG ++ + + L
Sbjct: 346 --------DVITEFDGYKVNDIVDLRKRLYQKKVGDRVKVKFYRGGKEKSVDIKL 392
>HTRA_LACHE (Q9Z4H7) Serine protease do-like htrA (EC 3.4.21.-)
Length = 413
Score = 100 bits (248), Expect = 6e-21
Identities = 68/177 (38%), Positives = 98/177 (54%), Gaps = 24/177 (13%)
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINT---AIYSPSGASSGVGFSIPLDTVNGIVDQL 182
++ +DAAINPGNSGG L++S+G +IGIN+ A S + G+ F+IP + V IV++L
Sbjct: 246 VIQTDAAINPGNSGGALVNSAGQVIGINSMKLAQSSDGTSVEGMAFAIPSNEVVTIVNEL 305
Query: 183 VKFGKVTRPILGIKFAPDQSV-----EQLGV-----SGVLVLDAPANGPAGKAGLQSTKR 232
VK GK+TRP LG++ Q + +L + +G+ + NG A AG++S
Sbjct: 306 VKKGKITRPQLGVRVIALQGIPEGYRSRLKIKSNLKNGIYIAFVSRNGSAANAGIKS--- 362
Query: 233 DSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILE 289
GD+IT V+G KV + L+ IL KVGD V V V R + V LE
Sbjct: 363 --------GDVITKVDGKKVEDVASLHSILYSHKVGDTVNVTVNRNGKDVDMKVKLE 411
>DEGP_SALTY (P26982) Protease do precursor (EC 3.4.21.-)
Length = 475
Score = 97.8 bits (242), Expect = 3e-20
Identities = 63/184 (34%), Positives = 98/184 (53%), Gaps = 21/184 (11%)
Query: 101 GNIVTNYHVIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPS 160
G VT+ ++ L++ + +DAAIN GNSGG L++ +G LIGINTAI +P
Sbjct: 201 GETVTS-GIVSALGRSGLNVENYENFIQTDAAINRGNSGGALVNLNGELIGINTAILAPD 259
Query: 161 GASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILGI-------KFAPDQSVEQLGVSGVLV 213
G + G+GF+IP + V + Q+V++G+V R LGI + A V+ G V
Sbjct: 260 GGNIGIGFAIPSNMVKNLTSQMVEYGQVKRGELGIMGTELNSELAKAMKVD--AQRGAFV 317
Query: 214 LDAPANGPAGKAGLQSTKRDSYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTV 273
N A KAG+++ GD+ITS+NG ++S + L + VG K+++
Sbjct: 318 SQVMPNSSAAKAGIKA-----------GDVITSLNGKPISSFAALRAQVGTMPVGSKISL 366
Query: 274 EVLR 277
+LR
Sbjct: 367 GLLR 370
>DEGP_ECOLI (P09376) Protease do precursor (EC 3.4.21.-)
Length = 474
Score = 97.8 bits (242), Expect = 3e-20
Identities = 61/170 (35%), Positives = 94/170 (54%), Gaps = 20/170 (11%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFG 186
+ +DAAIN GNSGG L++ +G LIGINTAI +P G + G+GF+IP + V + Q+V++G
Sbjct: 225 IQTDAAINRGNSGGALVNLNGELIGINTAILAPDGGNIGIGFAIPSNMVKNLTSQMVEYG 284
Query: 187 KVTRPILGI-------KFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLI 239
+V R LGI + A V+ G V N A KAG+++
Sbjct: 285 QVKRGELGIMGTELNSELAKAMKVD--AQRGAFVSQVLPNSSAAKAGIKA---------- 332
Query: 240 LGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILE 289
GD+ITS+NG ++S + L + VG K+T+ +LR + + + L+
Sbjct: 333 -GDVITSLNGKPISSFAALRAQVGTMPVGSKLTLGLLRDGKQVNVNLELQ 381
>DEGS_HAEIN (P44947) Protease degS precursor (EC 3.4.21.-)
Length = 340
Score = 93.6 bits (231), Expect = 5e-19
Identities = 53/168 (31%), Positives = 93/168 (54%), Gaps = 17/168 (10%)
Query: 127 VSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSG--ASSGVGFSIPLDTVNGIVDQLVK 184
+ +DA+IN GNSGG L++S+G L+GI+T + + G+ F+IP+D N ++ ++++
Sbjct: 186 IQTDASINRGNSGGALINSAGELVGISTLSIGKTANEIAEGLNFAIPIDIANDVLRKIMR 245
Query: 185 FGKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDII 244
G+V R G++ S E+ G+++ D N PA K+G+Q +GD+I
Sbjct: 246 DGRVIRGYFGVQSDISSSSEE----GIVITDVSPNSPAAKSGIQ-----------VGDVI 290
Query: 245 TSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKP 292
+N + S ++ +I+ K KV V +LR +IPV++E P
Sbjct: 291 LKLNNQEGISAREMMQIIANTKPNSKVLVTILRLGKILQIPVVIEEFP 338
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,706,166
Number of Sequences: 164201
Number of extensions: 1528466
Number of successful extensions: 4796
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 44
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 4648
Number of HSP's gapped (non-prelim): 110
length of query: 295
length of database: 59,974,054
effective HSP length: 109
effective length of query: 186
effective length of database: 42,076,145
effective search space: 7826162970
effective search space used: 7826162970
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0225.5


