

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0219.2
(365 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
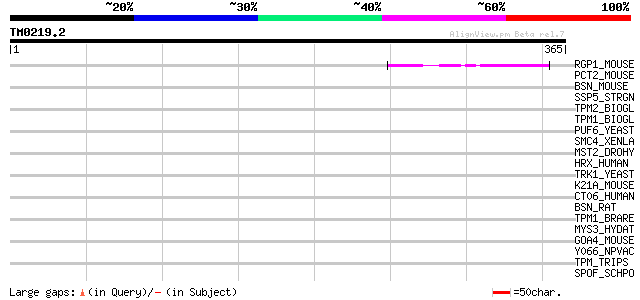
Score E
Sequences producing significant alignments: (bits) Value
RGP1_MOUSE (P46061) Ran GTPase-activating protein 1 44 8e-04
PCT2_MOUSE (P48725) Pericentrin 2 41 0.005
BSN_MOUSE (O88737) Bassoon protein 41 0.005
SSP5_STRGN (P16952) Agglutinin receptor precursor (SSP-5) 40 0.009
TPM2_BIOGL (P43689) Tropomyosin 2 (TMII) 40 0.012
TPM1_BIOGL (P42636) Tropomyosin 1 (TMI) (Bg 39) 39 0.021
PUF6_YEAST (Q04373) PUF6 protein 39 0.021
SMC4_XENLA (P50532) Structural maintenance of chromosome 4 (Chro... 39 0.027
MST2_DROHY (Q08696) Axoneme-associated protein mst101(2) 39 0.027
HRX_HUMAN (Q03164) Zinc finger protein HRX (ALL-1) (Trithorax-li... 39 0.027
TRK1_YEAST (P12685) Potassium transport protein, high-affinity 38 0.035
K21A_MOUSE (Q9QXL2) Kinesin family member 21A 38 0.046
CT06_HUMAN (Q9H501) Hypothetical protein C20orf6 38 0.046
BSN_RAT (O88778) Bassoon protein 38 0.046
TPM1_BRARE (P13104) Tropomyosin 1 alpha chain (Alpha-tropomyosin) 37 0.060
MYS3_HYDAT (P39922) Myosin heavy chain, clone 203 (Fragment) 37 0.060
GOA4_MOUSE (Q91VW5) Golgi autoantigen, golgin subfamily A member... 37 0.078
Y066_NPVAC (P41467) Hypothetical 94.0 kDa protein in POL-LEF3 in... 37 0.10
TPM_TRIPS (Q8WR63) Tropomyosin 37 0.10
SPOF_SCHPO (Q10411) Sporulation-specific protein 15 37 0.10
>RGP1_MOUSE (P46061) Ran GTPase-activating protein 1
Length = 589
Score = 43.5 bits (101), Expect = 8e-04
Identities = 29/107 (27%), Positives = 53/107 (49%), Gaps = 14/107 (13%)
Query: 249 QAKCQKLEKKYERLNASILGRASLLFAQGFLAAKEQISVVEPGFDLSRIGWLKEIKDGQV 308
+A K E + LN + LG EQ+ V F+++++ L + D +
Sbjct: 315 EAVADKAELEKLDLNGNALGEEGC----------EQLQEVMDSFNMAKV--LASLSDDE- 361
Query: 309 VGDDDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQ 355
G+D+ + + D+E E EED ED +E+ + ++ E+E PQ G+ +
Sbjct: 362 -GEDEDEEEEGEEDDEEEEDEEDEEDDDEEEEEQEEEEEPPQRGSGE 407
>PCT2_MOUSE (P48725) Pericentrin 2
Length = 1920
Score = 40.8 bits (94), Expect = 0.005
Identities = 29/104 (27%), Positives = 54/104 (51%), Gaps = 6/104 (5%)
Query: 177 ELIATKNRYEKKAADYKTAYE---QAKADAETANKNLKAAEERC-AKLTDDLAASDLLLQ 232
E+ +++++K+ ++ K E QAK +AE + KNL+A + KL +DL + Q
Sbjct: 245 EMEGLQSQFQKELSEQKVELEKIFQAKHEAEVSLKNLEAQHQAAIKKLQEDLQSEH--CQ 302
Query: 233 KTKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASLLFAQ 276
+ L++ +K A + + + L+ YE L A LL++Q
Sbjct: 303 YLQDLEQKFREKEKAKELELETLQASYEDLKAQSQEEIRLLWSQ 346
>BSN_MOUSE (O88737) Bassoon protein
Length = 3941
Score = 40.8 bits (94), Expect = 0.005
Identities = 34/130 (26%), Positives = 52/130 (39%), Gaps = 18/130 (13%)
Query: 8 ANPIKMK--EYLAQSAAAAKKRAAETEQKKKNEGTSGSDNVRDPKRQKTSSAAGVKPLH- 64
A P+K K + L Q + + +A+ K + T S Q T ++ KPL
Sbjct: 548 AAPLKQKGPQGLGQPSGSLPAKASPQATKASPQATKASPQATKASPQTTKASPQAKPLRA 607
Query: 65 ------------QSTLDP-KGRPTEKKKGHDNVPPHQPDSGALINR--PSTPFIQAGPSS 109
+ T P K P K VPP P + + + R P+TP ++ P +
Sbjct: 608 TEPSKTSSSAQEKKTATPAKAEPVPKPPPETTVPPGTPKAKSGVKRTDPATPVVKPVPEA 667
Query: 110 AIGGEALPPL 119
GGEA P+
Sbjct: 668 PKGGEAEEPV 677
>SSP5_STRGN (P16952) Agglutinin receptor precursor (SSP-5)
Length = 1500
Score = 40.0 bits (92), Expect = 0.009
Identities = 35/136 (25%), Positives = 60/136 (43%), Gaps = 10/136 (7%)
Query: 134 NRTFDNRIHKDVSGQGPPNIASVAIHHALSAASIVAGMAQCVKELIATKNRYEKKAADYK 193
N+ D++ KD+ + + +A + A A +AQ K+L K E DY+
Sbjct: 170 NKAADDKYQKDLKSH-QEEVEKINTANATAKAEYEAKLAQYQKDLATVKKANEDSQQDYQ 228
Query: 194 ---TAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKHTAVQA 250
+AY+ A + AN K A E+ K ++ A ++ L K E I ++ +A
Sbjct: 229 NKLSAYQTELARVQKANAEAKEAYEKAVK--ENTAKNEAL----KVENEAIKQRNETAKA 282
Query: 251 KCQKLEKKYERLNASI 266
+ K+YE A+I
Sbjct: 283 TYEAAMKQYEADLAAI 298
Score = 30.8 bits (68), Expect = 5.6
Identities = 31/129 (24%), Positives = 51/129 (39%), Gaps = 17/129 (13%)
Query: 152 NIASVAIHHALSAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLK 211
++A+V +A + A A +A EL R +K AD K YE+A D + N +K
Sbjct: 376 DLAAVKQANATNEADYQAKLAAYQTELA----RVQKANADAKATYEKAVEDNKAKNAAIK 431
Query: 212 AAEERCAKLTDDLAASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRAS 271
A E + Q+ K K +A K +K++ A++ S
Sbjct: 432 AENEE-------------IKQRNAVAKTDYEAKLAKYEADLAKYKKEFAAYTAALAEAES 478
Query: 272 LLFAQGFLA 280
G+L+
Sbjct: 479 KKKQDGYLS 487
>TPM2_BIOGL (P43689) Tropomyosin 2 (TMII)
Length = 284
Score = 39.7 bits (91), Expect = 0.012
Identities = 29/109 (26%), Positives = 56/109 (50%), Gaps = 10/109 (9%)
Query: 171 MAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKN-------LKAAEERCAKLTDD 223
M Q V+++ TKN+ E++ + + + + D +TAN+ L+A+E+ A+L D
Sbjct: 25 MEQKVRDVEETKNKLEEEFNNLQKKFSNLQNDFDTANEGLTEAQTKLEASEKHVAELESD 84
Query: 224 LAASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASL 272
A L ++ + L+E + +Q+ +KLE+ + + S GR L
Sbjct: 85 TAG---LNRRIQLLEEDLERSEERLQSATEKLEEASKAADESERGRKVL 130
>TPM1_BIOGL (P42636) Tropomyosin 1 (TMI) (Bg 39)
Length = 284
Score = 38.9 bits (89), Expect = 0.021
Identities = 28/109 (25%), Positives = 56/109 (50%), Gaps = 10/109 (9%)
Query: 171 MAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKN-------LKAAEERCAKLTDD 223
M Q ++++ TKN+ E++ + + + + D +TAN+ L+A+E+ A+L D
Sbjct: 25 MEQKLRDVEETKNKLEEEFNNLQNKFSNLQNDFDTANEGLTEAQTKLEASEKHVAELESD 84
Query: 224 LAASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASL 272
A L ++ + L+E + +Q+ +KLE+ + + S GR L
Sbjct: 85 TAG---LNRRIQLLEEDLERSEERLQSATEKLEEASKAADESERGRKVL 130
>PUF6_YEAST (Q04373) PUF6 protein
Length = 656
Score = 38.9 bits (89), Expect = 0.021
Identities = 25/70 (35%), Positives = 33/70 (46%), Gaps = 5/70 (7%)
Query: 300 LKEIKDGQVVGDDDISLDLLPQFDDESEPEEDG---EDGNEQHKNEDHEKE--DPQAGTS 354
L + +D DD LD L D E+E E D D +E+H+NE+ EKE D G
Sbjct: 41 LSKKEDAVSSSSDDDDLDDLSTSDSEAEEEADELDISDDSEEHENENEEKEGKDKSEGGE 100
Query: 355 QGNNANNENL 364
GN+ L
Sbjct: 101 NGNHTEQRKL 110
>SMC4_XENLA (P50532) Structural maintenance of chromosome 4
(Chromosome-associated protein C) (Chromosome assembly
protein XCAP-C)
Length = 1290
Score = 38.5 bits (88), Expect = 0.027
Identities = 18/74 (24%), Positives = 42/74 (56%)
Query: 183 NRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETIN 242
++ K+ + +A +A+ +TA++NLK +EE A+ ++ A+D +++ + +
Sbjct: 901 DKVTKEIDECASAITKAQVSIKTADRNLKKSEEAVARTEKEIVANDKSIEELTEDLKKLE 960
Query: 243 DKHTAVQAKCQKLE 256
+K T V +C++ E
Sbjct: 961 EKATTVMNECKEAE 974
>MST2_DROHY (Q08696) Axoneme-associated protein mst101(2)
Length = 1391
Score = 38.5 bits (88), Expect = 0.027
Identities = 26/83 (31%), Positives = 45/83 (53%), Gaps = 5/83 (6%)
Query: 180 ATKNRYEKKAADYKTAYEQAKADAETANKNLKAAE-ERCAKLTDDLAASDLLLQKTKSLK 238
A K + EK A + K A E+ K + E A K +AAE ++C +L ++ + +K K +
Sbjct: 394 AEKKKCEKAAKERKEAAEKKKCE-EAAKKEKEAAERKKCEELAKNIKKA---AEKKKCKE 449
Query: 239 ETINDKHTAVQAKCQKLEKKYER 261
+K A + KC++L KK ++
Sbjct: 450 AAKKEKEAAERKKCEELAKKIKK 472
Score = 33.5 bits (75), Expect = 0.87
Identities = 28/80 (35%), Positives = 39/80 (48%), Gaps = 5/80 (6%)
Query: 180 ATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKE 239
A K + EK A K A E+ K AE A K +AAE+ K ++ A + + K +E
Sbjct: 666 AEKKKCEKAAKKRKEAAEKKKC-AEAAKKEKEAAEK---KKCEEAAKKEKEAAERKKCEE 721
Query: 240 TIND-KHTAVQAKCQKLEKK 258
K A + KC+KL KK
Sbjct: 722 LAKKIKKAAEKKKCKKLAKK 741
Score = 33.5 bits (75), Expect = 0.87
Identities = 26/86 (30%), Positives = 38/86 (43%), Gaps = 9/86 (10%)
Query: 173 QCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQ 232
+ +KE + +KKAA+ K E AK + E A K K E+ K ++ +
Sbjct: 759 KALKEKKKCRELAKKKAAEKKKCKEAAKKEKEAAEK--KKCEKTAKKRKEE-------AE 809
Query: 233 KTKSLKETINDKHTAVQAKCQKLEKK 258
K K K K A + KC+K KK
Sbjct: 810 KKKCEKTAKKRKEAAEKKKCEKAAKK 835
Score = 32.7 bits (73), Expect = 1.5
Identities = 26/85 (30%), Positives = 40/85 (46%), Gaps = 8/85 (9%)
Query: 182 KNRYEKKA----ADYKTAYEQAKADAETANKNLKAAE-ERCAKLTDDLAASDLLLQKTKS 236
K EKKA A K ++ K E ANK KAAE ++C K + + +K K
Sbjct: 359 KEEDEKKACKELAKKKKEADEKKKCEEAANKEKKAAEKKKCEKAAKERKEA---AEKKKC 415
Query: 237 LKETINDKHTAVQAKCQKLEKKYER 261
+ +K A + KC++L K ++
Sbjct: 416 EEAAKKEKEAAERKKCEELAKNIKK 440
Score = 32.3 bits (72), Expect = 1.9
Identities = 25/82 (30%), Positives = 39/82 (47%), Gaps = 7/82 (8%)
Query: 187 KKAADYKTAYEQAKADAETANKNL--KAAEER-----CAKLTDDLAASDLLLQKTKSLKE 239
KKAA+ K + AK + ETA K KAA++R K + +K K +
Sbjct: 647 KKAAEKKKCKKLAKKEKETAEKKKCEKAAKKRKEAAEKKKCAEAAKKEKEAAEKKKCEEA 706
Query: 240 TINDKHTAVQAKCQKLEKKYER 261
+K A + KC++L KK ++
Sbjct: 707 AKKEKEAAERKKCEELAKKIKK 728
Score = 32.0 bits (71), Expect = 2.5
Identities = 22/91 (24%), Positives = 39/91 (42%), Gaps = 2/91 (2%)
Query: 173 QCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDL--L 230
+C K K EKK + + A+ + K+ K +E K + AA +
Sbjct: 526 KCEKAAKKRKEAAEKKKCEKAAKKRKEAAEKKKCEKSAKKRKEAAEKKKCEKAAKERKEA 585
Query: 231 LQKTKSLKETINDKHTAVQAKCQKLEKKYER 261
+K K + +K A + KC++L KK ++
Sbjct: 586 AEKKKCEEAAKKEKEVAERKKCEELAKKIKK 616
Score = 32.0 bits (71), Expect = 2.5
Identities = 28/86 (32%), Positives = 42/86 (48%), Gaps = 7/86 (8%)
Query: 180 ATKNRYEKKAADYKTAYEQAKADAETANKNLKAAE----ERCAKLTDDLAASDLLLQKTK 235
ATK + E++A K A E+ + + E A K +AAE E AK + A ++ K
Sbjct: 995 ATKKKCEERAKKQKEAAEKKQCE-ERAKKLKEAAEQKQCEERAKKLKEAAEKKQCEERAK 1053
Query: 236 SLKETINDKHTAVQAKCQKLEKKYER 261
LKE K +AK KL++ E+
Sbjct: 1054 KLKEAAEQKQCEERAK--KLKEAAEK 1077
>HRX_HUMAN (Q03164) Zinc finger protein HRX (ALL-1) (Trithorax-like
protein)
Length = 3969
Score = 38.5 bits (88), Expect = 0.027
Identities = 27/81 (33%), Positives = 41/81 (50%), Gaps = 3/81 (3%)
Query: 14 KEYLAQSAAAAKKRA--AETEQKKKNEGTSGSDNVRDPKRQKTSSAA-GVKPLHQSTLDP 70
K YL + A A KK+ ++T +KK ++ +S NV D ++ T SA P S+ P
Sbjct: 1203 KAYLQKQAKAVKKKEKKSKTSEKKDSKESSVVKNVVDSSQKPTPSAREDPAPKKSSSEPP 1262
Query: 71 KGRPTEKKKGHDNVPPHQPDS 91
+P E+K NV P+S
Sbjct: 1263 PRKPVEEKSEEGNVSAPGPES 1283
>TRK1_YEAST (P12685) Potassium transport protein, high-affinity
Length = 1235
Score = 38.1 bits (87), Expect = 0.035
Identities = 14/38 (36%), Positives = 24/38 (62%)
Query: 324 DESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGNNANN 361
++++ E+DG D ++ +NE HE + Q +S NN NN
Sbjct: 1013 EDTDTEDDGNDEDDDEENESHEGQSSQRSSSNNNNNNN 1050
>K21A_MOUSE (Q9QXL2) Kinesin family member 21A
Length = 1672
Score = 37.7 bits (86), Expect = 0.046
Identities = 45/190 (23%), Positives = 82/190 (42%), Gaps = 5/190 (2%)
Query: 164 AASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDD 223
A ++A + +E+ + Y K+ D + +++A E KNL A R +
Sbjct: 461 ANQVLARAGEGNEEISNMIHSYIKEIEDLRAKLLESEAVNENLRKNLTRATARSPYFSAS 520
Query: 224 LAASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNA-SILGRASLLFA-QGFLAA 281
A S +L K E I+ ++ +K +KK +RL GR A + A
Sbjct: 521 SAFSPTILSSDKETIEIIDLAKKDLEKLKRKEKKKKKRLQKLEESGREERSVAGKDDNAD 580
Query: 282 KEQISVVEPGFDLSRIGWLKEIKDGQVVGDDDISLDLLPQFDDESEPEEDGEDGNEQHKN 341
+Q E G L ++++ Q V D + + + D+E E + +GE+ +++ +
Sbjct: 581 TDQEKKEEKGVSEKENNEL-DVEENQEVSDHEDEEE--EEEDEEEEDDIEGEESSDESDS 637
Query: 342 EDHEKEDPQA 351
E EK + QA
Sbjct: 638 ESDEKANYQA 647
>CT06_HUMAN (Q9H501) Hypothetical protein C20orf6
Length = 851
Score = 37.7 bits (86), Expect = 0.046
Identities = 40/166 (24%), Positives = 70/166 (42%), Gaps = 21/166 (12%)
Query: 200 KADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKY 259
K E KN K + TD L +K ++L ++ + + +C K ++
Sbjct: 156 KDSKEFTQKNKKEKKNIVQHTTDSS-----LEEKQRTLDSGTSEIVKSPRIECSKTRREM 210
Query: 260 ERLNASILGRASLLF---AQGFLAAKEQISVVEPGFDLSRIGWLKEIKD-----GQVVGD 311
+ + I+ R S + G + K+ + E +S IG +E ++ G+ GD
Sbjct: 211 QSVVQLIMTRDSDGYENSTDGEMCDKDALE--EDSESVSEIGSDEESENEITSVGRASGD 268
Query: 312 DDISLDLLPQFDDESEPEEDGEDGNEQHKNEDHEKEDPQAGTSQGN 357
DD S D DE E E++ ED +E +++D P +GN
Sbjct: 269 DDGSED------DEEEDEDEEEDEDEDSEDDDKSDSGPDLARGKGN 308
Score = 31.2 bits (69), Expect = 4.3
Identities = 53/220 (24%), Positives = 94/220 (42%), Gaps = 29/220 (13%)
Query: 161 ALSAASIVAGMAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANK-NLKAAEERCA- 218
ALS I Q KE I +KN EKK K ++ ++ + N +K + C
Sbjct: 88 ALSQKKIKKKKTQTKKE-IDSKNLVEKKKETKKANHKGSENKTDLDNSIGIKKMKTSCKF 146
Query: 219 KLTDDLAA---SDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASILGRASLLFA 275
K+ +++ S QK K K+ I +HT + LE+K L++
Sbjct: 147 KIDSNISPKKDSKEFTQKNKKEKKNIV-QHTTDSS----LEEKQRTLDSGTSEIVKSPRI 201
Query: 276 QGFLAAKEQISVVE-------PGFDLSRIGWLKEIKDGQVVGDDDISLDLLPQFDDESEP 328
+ +E SVV+ G++ S G E+ D + +D S+ + D+ESE
Sbjct: 202 ECSKTRREMQSVVQLIMTRDSDGYENSTDG---EMCDKDALEEDSESVSEIGS-DEESEN 257
Query: 329 E-------EDGEDGNEQHKNEDHEKEDPQAGTSQGNNANN 361
E +DG+E + ED ++E+ + S+ ++ ++
Sbjct: 258 EITSVGRASGDDDGSEDDEEEDEDEEEDEDEDSEDDDKSD 297
>BSN_RAT (O88778) Bassoon protein
Length = 3937
Score = 37.7 bits (86), Expect = 0.046
Identities = 26/116 (22%), Positives = 49/116 (41%), Gaps = 4/116 (3%)
Query: 8 ANPIKMK--EYLAQSAAAAKKRAAETEQKKKNEGTSGSDNVRDPKRQKTSSAAGVKPLHQ 65
A+P+K K + Q + + +A+ K + S + + + S + P +
Sbjct: 546 ASPLKQKGPQGPGQPSGSLPPKASPQAAKASPQAAKASPQAKPLRASEPSKTSSSAPEKK 605
Query: 66 STLDPKGRPTEKKKGHDNVPPHQPDSGALINR--PSTPFIQAGPSSAIGGEALPPL 119
+ + K P K VPP P + + + R P+TP ++ P + GEA P+
Sbjct: 606 TGIPVKAEPVPKPPPETAVPPGTPKAKSGVKRTDPATPVVKPVPEAPKSGEAEEPV 661
>TPM1_BRARE (P13104) Tropomyosin 1 alpha chain (Alpha-tropomyosin)
Length = 284
Score = 37.4 bits (85), Expect = 0.060
Identities = 26/73 (35%), Positives = 35/73 (47%)
Query: 186 EKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKH 245
+KK K E A AE A + KAAEER +L DDL A L+ T+ + ++
Sbjct: 5 KKKMQMLKLDKENALDRAEQAETDKKAAEERSKQLEDDLVALQKKLKATEDELDKYSEAL 64
Query: 246 TAVQAKCQKLEKK 258
Q K + EKK
Sbjct: 65 KDAQEKLELAEKK 77
>MYS3_HYDAT (P39922) Myosin heavy chain, clone 203 (Fragment)
Length = 539
Score = 37.4 bits (85), Expect = 0.060
Identities = 27/124 (21%), Positives = 56/124 (44%), Gaps = 6/124 (4%)
Query: 139 NRIHKDVSGQGPPNIASVAIHHALSAASIVA------GMAQCVKELIATKNRYEKKAADY 192
NR+ K++ N A+++ A + A+I M + +L K+ + +
Sbjct: 375 NRLRKEIEALNIANDAAISAIKAKTNATIAEIQEENEAMKKAKAKLEKEKSALNNELNET 434
Query: 193 KTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKHTAVQAKC 252
K + +Q K ++KN + EE+ +L LA D L +++S +N + A+ ++
Sbjct: 435 KNSLDQIKKQKTNSDKNSRMLEEQINELNSKLAQVDELHSQSESKNSKVNSELLALNSQL 494
Query: 253 QKLE 256
+ E
Sbjct: 495 SESE 498
>GOA4_MOUSE (Q91VW5) Golgi autoantigen, golgin subfamily A member 4
(tGolgin-1)
Length = 2238
Score = 37.0 bits (84), Expect = 0.078
Identities = 36/135 (26%), Positives = 59/135 (43%), Gaps = 6/135 (4%)
Query: 156 VAIHHALSAASIVAGMAQC-VKELIATKNRYEKKAADYKTAYEQAKADAETANKNLKAAE 214
+++ L+ ++A +Q V EL EK+ K+ +E +K E + NLK+
Sbjct: 1166 LSLQEELAELKLLADKSQLRVSELTGQVQAAEKELQSCKSLHELSKKSLEDKSLNLKSLL 1225
Query: 215 ERCAKLTDDLA--ASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNASIL---GR 269
E A D LL KT L T DK A+ A+ + ++ + ++L G+
Sbjct: 1226 EELASQLDSRCERTKALLEAKTNELVCTSRDKADAILARLSQCQRHTATVGEALLRRMGQ 1285
Query: 270 ASLLFAQGFLAAKEQ 284
S L AQ +EQ
Sbjct: 1286 VSELEAQLTQLTEEQ 1300
>Y066_NPVAC (P41467) Hypothetical 94.0 kDa protein in POL-LEF3
intergenic region
Length = 808
Score = 36.6 bits (83), Expect = 0.10
Identities = 41/165 (24%), Positives = 74/165 (44%), Gaps = 19/165 (11%)
Query: 116 LPPLLNLSDPRFNGLDFMNRTFDNRIHKD-----VSGQGPPNIASVAIHHALSAASIVAG 170
L + LSD + + + R NRI K+ V + SVA+ A+SA +V
Sbjct: 608 LQSVSELSDLKSQIISIVPRNIVNRILKENYKVKVENVNAELLESVAVTSAVSA--LVQQ 665
Query: 171 MAQCVKELIATKNRYEKKAADYKTAYEQAKADAETANK--------NLKAAEERCAKLT- 221
+ K+ + + +E K D + EQ + D E+ ++ N +ER L
Sbjct: 666 YERSEKQNVKLRQEFEIKLNDLQRLLEQNQTDFESISEFISRDPAFNRNLNDERFQNLRQ 725
Query: 222 --DDLAASDLLLQKTKSLKETINDKHTAVQAKCQKLEKKYERLNA 264
D++++ L+ TK +KE + AV+++ KL + + LN+
Sbjct: 726 QYDEMSSKYSALETTK-IKEMESIADQAVKSEMSKLNTQLDELNS 769
>TPM_TRIPS (Q8WR63) Tropomyosin
Length = 284
Score = 36.6 bits (83), Expect = 0.10
Identities = 23/104 (22%), Positives = 49/104 (47%), Gaps = 2/104 (1%)
Query: 186 EKKAADYKTAYEQAKADAETANKNLKAAEERCAKLTDDLAASDLLLQKTKSLKETINDKH 245
+KK K + A A+ A + + +ER KL ++L + + + ++ + ++
Sbjct: 5 KKKMQAMKIEKDNAMDRADAAEEKARQQQERVEKLEEELRDTQKKMMQVENELDKAQEEL 64
Query: 246 TAVQAKCQKLEKKYERLNASI--LGRASLLFAQGFLAAKEQISV 287
T A+ ++ EKK + A + L R L + F A+E++ +
Sbjct: 65 TGANAQLEEKEKKVQEAEAEVAALNRRIQLLEEDFERAEERLKI 108
>SPOF_SCHPO (Q10411) Sporulation-specific protein 15
Length = 1957
Score = 36.6 bits (83), Expect = 0.10
Identities = 22/92 (23%), Positives = 46/92 (49%), Gaps = 16/92 (17%)
Query: 176 KELIATKNRYEKKAADYKTAYEQAK------ADAETANKNLKAAEERCAKLTDDLAASDL 229
++L+ + +KK DY+ E A+ + +NK+L+ +ER K L
Sbjct: 208 RQLLTVTEKLDKKEKDYEKIKEDVSSIKASLAEEQASNKSLRGEQERLEK---------L 258
Query: 230 LLQKTKSLKETINDKHTAVQAKCQKLEKKYER 261
L+ K++ T+ +++A+C+ L++K E+
Sbjct: 259 LVSSNKTV-STLRQTENSLRAECKTLQEKLEK 289
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.308 0.127 0.350
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,344,249
Number of Sequences: 164201
Number of extensions: 1980146
Number of successful extensions: 11107
Number of sequences better than 10.0: 348
Number of HSP's better than 10.0 without gapping: 152
Number of HSP's successfully gapped in prelim test: 199
Number of HSP's that attempted gapping in prelim test: 9644
Number of HSP's gapped (non-prelim): 1147
length of query: 365
length of database: 59,974,054
effective HSP length: 112
effective length of query: 253
effective length of database: 41,583,542
effective search space: 10520636126
effective search space used: 10520636126
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0219.2


